Preprocessing_Countdata
AGC, AY
15 3 2021
Last updated: 2022-02-21
Checks: 7 0
Knit directory: CePTER_RNASeq/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20210315) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 56805fa. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/.Rhistory
Ignored: analysis/MDSplots/D62_mdsplots.RData
Ignored: data/.Rhistory
Ignored: data/Countmatrix.RData
Ignored: oldcode/
Ignored: output/D62_GOresWGCNA.xlsx
Ignored: output/D62_ResTabs_KO.RData
Ignored: output/D62_ResTabs_KO_WGCNA.RData
Ignored: output/D62_WGCNA_adj_TOM.RData
Ignored: output/D62_dds_matrix.RData
Ignored: output/GOres_Comparisons.xlsx
Ignored: output/GOres_D244DIFFRAPA.xlsx
Ignored: output/GOres_D244DIFFnoRAPA.xlsx
Ignored: output/GOres_D244noDIFFRAPA.xlsx
Ignored: output/GOres_D244noDIFFnoRAPA.xlsx
Ignored: output/GOres_D62DIFFRAPA.xlsx
Ignored: output/GOres_D62DIFFnoRAPA.xlsx
Ignored: output/GOres_D62noDIFFRAPA.xlsx
Ignored: output/GOres_D62noDIFFnoRAPA.xlsx
Ignored: output/GOres_DIFFRAPA.xlsx
Ignored: output/GOres_DIFFnoRAPA.xlsx
Ignored: output/GOres_ReNDIFFRAPA.xlsx
Ignored: output/GOres_ReNDIFFnoRAPA.xlsx
Ignored: output/GOres_ReNnoDIFFRAPA.xlsx
Ignored: output/GOres_ReNnoDIFFnoRAPA.xlsx
Ignored: output/GOres_noDIFFRAPA.xlsx
Ignored: output/GOres_noDIFFnoRAPA.xlsx
Ignored: output/ResTabs_KO.RData
Ignored: output/ResTabs_KO_WGCNA.RData
Ignored: output/Restab_D244_DIFF_RAPA.xlsx
Ignored: output/Restab_D244_DIFF_noRAPA.xlsx
Ignored: output/Restab_D244_noDIFF_RAPA.xlsx
Ignored: output/Restab_D244_noDIFF_noRAPA.xlsx
Ignored: output/Restab_D62_DIFF_RAPA.xlsx
Ignored: output/Restab_D62_DIFF_noRAPA.xlsx
Ignored: output/Restab_D62_noDIFF_RAPA.xlsx
Ignored: output/Restab_D62_noDIFF_noRAPA.xlsx
Ignored: output/Restab_ReN_DIFF_RAPA.xlsx
Ignored: output/Restab_ReN_DIFF_noRAPA.xlsx
Ignored: output/Restab_ReN_noDIFF_RAPA.xlsx
Ignored: output/Restab_ReN_noDIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_All_DIFF_RAPA.xlsx
Ignored: output/Restab_Repl_All_noDIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_D62D244_DIFF_RAPA.xlsx
Ignored: output/Restab_Repl_D62D244_DIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_D62D244_noDIFF_RAPA.xlsx
Ignored: output/Restab_Repl_D62D244_noDIFF_noRAPA.xlsx
Ignored: output/WGCNA_adj_TOM.RData
Untracked files:
Untracked: Wrapper.sh
Untracked: data/HumanM1 singlecellseq/
Untracked: data/genelists_to_test/
Untracked: mattson/
Untracked: wflow_helper.R
Unstaged changes:
Modified: output/dds_matrix.RData
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with wflow_publish() to start tracking its development.
Preprocessing
#Load Count Matrix
if (file.exists(secret)){
source(secret)
} else {
UID = rstudioapi::askForPassword("fuchs user")
PWD = rstudioapi::askForPassword("fuchs password")
}
filetarget= paste0(home,"/data/Countmatrix.RData")
if(!file.exists(filetarget)){
url="sftp://fuchs.hhlr-gu.de/scratch/fuchs/agmisc/chiocchetti/RNASeq_Data/Cepter/Output/Countmatrix.RData"
bin = getBinaryURL(url, userpw=paste0(UID,":",PWD))
writeBin(bin, filetarget)
load(filetarget)
} else {
load(filetarget)
}
Ntot= nrow(Countdata)
#merge non unique annotations
if(length(unique(rownames(Countdata))) != Ntot){
Countdata = Countdata %>% dplyr::group_by(row.names(Countdata)) %>% summarise_each(sum)
Ntot= nrow(Countdata)
}
hgnc=gconvert(query=as.numeric(rownames(Countdata)),
organism = "hsapiens",
numeric_ns = "ENTREZGENE_ACC",
target = "HGNC")
Ids = hgnc %>% dplyr::select(name, input, description) %>% group_by(input) %>%
dplyr::summarise(name=paste(name, sep="; ", collapse = ";"), description = dplyr::first(description))
rowdescription = data.frame(entrez_gene = Ids$input,
hgnc=Ids$name,
description=Ids$description)
if(! all(row.names(Countdata) %in% rowdescription$entrez_gene)){
rowdescription = rowdescription[match(row.names(Countdata), rowdescription$entrez_gene),]
rownames(rowdescription)=row.names(Countdata)
}
# load and parse sample information
SampleInfo=read.csv2(paste0(home,"/data/Sample_info_CePTER_RNASeq.csv"),
row.names = 1)
SampleInfo$Row=gsub("[0-9]*","",SampleInfo$Position)
SampleInfo$Col=as.numeric(gsub("[A-Z]*","",SampleInfo$Position))
# set factors and relevel
SampleInfo$CellLine = as.factor(SampleInfo$CellLine)
SampleInfo$gRNA = paste0("sg",SampleInfo$gRNA)
SampleInfo$gRNA = factor(SampleInfo$gRNA, levels=c("sgNTC", "sg2.1", "sg2.2"),
labels=c("sgNTC", "sg2.1", "sg2.2"))
SampleInfo$gRNA = relevel(SampleInfo$gRNA,ref="sgNTC" )
SampleInfo$KO = factor(SampleInfo$KO, levels=c(T,F), labels=c("KO", "WT"))
SampleInfo$KO = relevel(SampleInfo$KO,ref="WT" )
SampleInfo$DIFF = factor(SampleInfo$DIFF, levels=c(TRUE,FALSE),
labels=c("DIFF", "noDIFF"))
SampleInfo$DIFF = relevel(SampleInfo$DIFF,ref="noDIFF")
SampleInfo$RAPA = factor(SampleInfo$RAPA, levels=c(T,F),
labels=c("RAPA", "noRAPA"))
SampleInfo$RAPA = relevel(SampleInfo$RAPA,ref="noRAPA")
SampleInfo$label = with(SampleInfo, paste(CellLine,gRNA,DIFF,RAPA, sep="_"))
SampleInfo$fastQID = rownames(SampleInfo)
SampleInfo = SampleInfo %>% dplyr::group_by(label) %>% mutate(replicate=seq(n())) %>% as.data.frame()
SampleInfo$label_rep=with(SampleInfo, paste(label,replicate,sep="_"))
rownames(SampleInfo)=SampleInfo$fastQID
# align datasets
checkfiles = all(rownames(SampleInfo) %in% colnames(Countdata))
IDs=intersect(rownames(SampleInfo), colnames(Countdata))
Countdata = Countdata[,IDs]
SampleInfo = SampleInfo[IDs, ]
SampleInfo$reads_per_sample = colSums(Countdata)
display_tab(head(Countdata))| DE10NGSUKBR112901 | DE80NGSUKBR112902 | DE53NGSUKBR112903 | DE26NGSUKBR112904 | DE96NGSUKBR112905 | DE69NGSUKBR112906 | DE42NGSUKBR112907 | DE15NGSUKBR112908 | DE85NGSUKBR112909 | DE58NGSUKBR112910 | DE31NGSUKBR112911 | DE04NGSUKBR112912 | DE74NGSUKBR112913 | DE47NGSUKBR112914 | DE20NGSUKBR112915 | DE90NGSUKBR112916 | DE63NGSUKBR112917 | DE36NGSUKBR112918 | DE09NGSUKBR112919 | DE79NGSUKBR112920 | DE52NGSUKBR112921 | DE25NGSUKBR112922 | DE95NGSUKBR112923 | DE68NGSUKBR112924 | DE41NGSUKBR112925 | DE14NGSUKBR112926 | DE84NGSUKBR112927 | DE57NGSUKBR112928 | DE30NGSUKBR112929 | DE03NGSUKBR112930 | DE73NGSUKBR112931 | DE46NGSUKBR112932 | DE19NGSUKBR112933 | DE89NGSUKBR112934 | DE62NGSUKBR112935 | DE35NGSUKBR112936 | DE08NGSUKBR112937 | DE78NGSUKBR112938 | DE51NGSUKBR112939 | DE24NGSUKBR112940 | DE94NGSUKBR112941 | DE67NGSUKBR112942 | DE40NGSUKBR112943 | DE13NGSUKBR112944 | DE83NGSUKBR112945 | DE56NGSUKBR112946 | DE29NGSUKBR112947 | DE02NGSUKBR112948 | DE72NGSUKBR112949 | DE45NGSUKBR112950 | DE18NGSUKBR112951 | DE88NGSUKBR112952 | DE61NGSUKBR112953 | DE34NGSUKBR112954 | DE07NGSUKBR112955 | DE77NGSUKBR112956 | DE50NGSUKBR112957 | DE23NGSUKBR112958 | DE93NGSUKBR112959 | DE66NGSUKBR112960 | DE39NGSUKBR112961 | DE12NGSUKBR112962 | DE82NGSUKBR112963 | DE55NGSUKBR112964 | DE28NGSUKBR112965 | DE98NGSUKBR112966 | DE71NGSUKBR112967 | DE44NGSUKBR112968 | DE17NGSUKBR112969 | DE87NGSUKBR112970 | DE60NGSUKBR112971 | DE33NGSUKBR112972 | DE06NGSUKBR112973 | DE76NGSUKBR112974 | DE49NGSUKBR112975 | DE22NGSUKBR112976 | DE92NGSUKBR112977 | DE65NGSUKBR112978 | DE38NGSUKBR112979 | DE11NGSUKBR112980 | DE81NGSUKBR112981 | DE54NGSUKBR112982 | DE27NGSUKBR112983 | DE97NGSUKBR112984 | DE70NGSUKBR112985 | DE43NGSUKBR112986 | DE16NGSUKBR112987 | DE86NGSUKBR112988 | DE59NGSUKBR112989 | DE32NGSUKBR112990 | DE05NGSUKBR112991 | DE75NGSUKBR112992 | DE48NGSUKBR112993 | DE21NGSUKBR112994 | DE91NGSUKBR112995 | DE64NGSUKBR112996 | DE37NGSUKBR112997 | DE10NGSUKBR112998 | DE80NGSUKBR112999 | DE53NGSUKBR113000 | DE26NGSUKBR113001 | DE96NGSUKBR113002 | DE69NGSUKBR113003 | DE42NGSUKBR113004 | DE15NGSUKBR113005 | DE85NGSUKBR113006 | DE58NGSUKBR113007 | DE31NGSUKBR113008 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 100287102 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| 653635 | 46 | 82 | 133 | 121 | 84 | 78 | 31 | 21 | 32 | 48 | 26 | 47 | 59 | 18 | 113 | 66 | 71 | 99 | 101 | 59 | 55 | 75 | 77 | 42 | 27 | 32 | 41 | 85 | 90 | 112 | 75 | 0 | 32 | 25 | 42 | 19 | 42 | 1 | 20 | 33 | 7 | 47 | 64 | 28 | 32 | 42 | 66 | 21 | 25 | 75 | 60 | 41 | 18 | 99 | 33 | 11 | 7 | 55 | 29 | 16 | 53 | 75 | 95 | 2 | 31 | 121 | 1 | 36 | 43 | 20 | 30 | 82 | 56 | 109 | 92 | 27 | 62 | 17 | 35 | 43 | 15 | 42 | 26 | 17 | 112 | 81 | 126 | 137 | 31 | 64 | 64 | 58 | 47 | 16 | 84 | 93 | 31 | 65 | 55 | 38 | 20 | 12 | 38 | 48 | 51 | 60 | 58 | 80 |
| 102466751 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| 100302278 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| 645520 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| 79501 | 0 | 0 | 5 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 8 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
display_tab(SampleInfo)| Plate | Position | Row | Col | CellLine | gRNA | KO | DIFF | RAPA | label | fastQID | replicate | label_rep | reads_per_sample | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DE10NGSUKBR112901 | 1 | A01 | A | 1 | D62 | sgNTC | WT | DIFF | RAPA | D62_sgNTC_DIFF_RAPA | DE10NGSUKBR112901 | 1 | D62_sgNTC_DIFF_RAPA_1 | 8167558 |
| DE80NGSUKBR112902 | 1 | A02 | A | 2 | D62 | sgNTC | WT | DIFF | RAPA | D62_sgNTC_DIFF_RAPA | DE80NGSUKBR112902 | 2 | D62_sgNTC_DIFF_RAPA_2 | 7947513 |
| DE53NGSUKBR112903 | 1 | A03 | A | 3 | D62 | sgNTC | WT | DIFF | RAPA | D62_sgNTC_DIFF_RAPA | DE53NGSUKBR112903 | 3 | D62_sgNTC_DIFF_RAPA_3 | 8927353 |
| DE26NGSUKBR112904 | 1 | A04 | A | 4 | D62 | sgNTC | WT | DIFF | noRAPA | D62_sgNTC_DIFF_noRAPA | DE26NGSUKBR112904 | 1 | D62_sgNTC_DIFF_noRAPA_1 | 6192682 |
| DE96NGSUKBR112905 | 1 | A05 | A | 5 | D62 | sgNTC | WT | DIFF | noRAPA | D62_sgNTC_DIFF_noRAPA | DE96NGSUKBR112905 | 2 | D62_sgNTC_DIFF_noRAPA_2 | 6316070 |
| DE69NGSUKBR112906 | 1 | A06 | A | 6 | D62 | sgNTC | WT | DIFF | noRAPA | D62_sgNTC_DIFF_noRAPA | DE69NGSUKBR112906 | 3 | D62_sgNTC_DIFF_noRAPA_3 | 7211176 |
| DE42NGSUKBR112907 | 1 | A07 | A | 7 | D62 | sg2.1 | KO | DIFF | RAPA | D62_sg2.1_DIFF_RAPA | DE42NGSUKBR112907 | 1 | D62_sg2.1_DIFF_RAPA_1 | 6472088 |
| DE15NGSUKBR112908 | 1 | A08 | A | 8 | D62 | sg2.1 | KO | DIFF | RAPA | D62_sg2.1_DIFF_RAPA | DE15NGSUKBR112908 | 2 | D62_sg2.1_DIFF_RAPA_2 | 6381728 |
| DE85NGSUKBR112909 | 1 | A09 | A | 9 | D62 | sg2.1 | KO | DIFF | RAPA | D62_sg2.1_DIFF_RAPA | DE85NGSUKBR112909 | 3 | D62_sg2.1_DIFF_RAPA_3 | 7515594 |
| DE58NGSUKBR112910 | 1 | A10 | A | 10 | D62 | sg2.1 | KO | DIFF | noRAPA | D62_sg2.1_DIFF_noRAPA | DE58NGSUKBR112910 | 1 | D62_sg2.1_DIFF_noRAPA_1 | 8072060 |
| DE31NGSUKBR112911 | 1 | A11 | A | 11 | D62 | sg2.1 | KO | DIFF | noRAPA | D62_sg2.1_DIFF_noRAPA | DE31NGSUKBR112911 | 2 | D62_sg2.1_DIFF_noRAPA_2 | 9132042 |
| DE04NGSUKBR112912 | 1 | A12 | A | 12 | D62 | sg2.1 | KO | DIFF | noRAPA | D62_sg2.1_DIFF_noRAPA | DE04NGSUKBR112912 | 3 | D62_sg2.1_DIFF_noRAPA_3 | 9158749 |
| DE74NGSUKBR112913 | 1 | B01 | B | 1 | D62 | sg2.2 | KO | DIFF | RAPA | D62_sg2.2_DIFF_RAPA | DE74NGSUKBR112913 | 1 | D62_sg2.2_DIFF_RAPA_1 | 8022580 |
| DE47NGSUKBR112914 | 1 | B02 | B | 2 | D62 | sg2.2 | KO | DIFF | RAPA | D62_sg2.2_DIFF_RAPA | DE47NGSUKBR112914 | 2 | D62_sg2.2_DIFF_RAPA_2 | 5290014 |
| DE20NGSUKBR112915 | 1 | B03 | B | 3 | D62 | sg2.2 | KO | DIFF | RAPA | D62_sg2.2_DIFF_RAPA | DE20NGSUKBR112915 | 3 | D62_sg2.2_DIFF_RAPA_3 | 7350899 |
| DE90NGSUKBR112916 | 1 | B04 | B | 4 | D62 | sg2.2 | KO | DIFF | noRAPA | D62_sg2.2_DIFF_noRAPA | DE90NGSUKBR112916 | 1 | D62_sg2.2_DIFF_noRAPA_1 | 7142950 |
| DE63NGSUKBR112917 | 1 | B05 | B | 5 | D62 | sg2.2 | KO | DIFF | noRAPA | D62_sg2.2_DIFF_noRAPA | DE63NGSUKBR112917 | 2 | D62_sg2.2_DIFF_noRAPA_2 | 6379496 |
| DE36NGSUKBR112918 | 1 | B06 | B | 6 | D62 | sg2.2 | KO | DIFF | noRAPA | D62_sg2.2_DIFF_noRAPA | DE36NGSUKBR112918 | 3 | D62_sg2.2_DIFF_noRAPA_3 | 6345821 |
| DE09NGSUKBR112919 | 1 | B07 | B | 7 | D62 | sgNTC | WT | noDIFF | RAPA | D62_sgNTC_noDIFF_RAPA | DE09NGSUKBR112919 | 1 | D62_sgNTC_noDIFF_RAPA_1 | 6954777 |
| DE79NGSUKBR112920 | 1 | B08 | B | 8 | D62 | sgNTC | WT | noDIFF | RAPA | D62_sgNTC_noDIFF_RAPA | DE79NGSUKBR112920 | 2 | D62_sgNTC_noDIFF_RAPA_2 | 6247879 |
| DE52NGSUKBR112921 | 1 | B09 | B | 9 | D62 | sgNTC | WT | noDIFF | RAPA | D62_sgNTC_noDIFF_RAPA | DE52NGSUKBR112921 | 3 | D62_sgNTC_noDIFF_RAPA_3 | 7651123 |
| DE25NGSUKBR112922 | 1 | B10 | B | 10 | D62 | sgNTC | WT | noDIFF | noRAPA | D62_sgNTC_noDIFF_noRAPA | DE25NGSUKBR112922 | 1 | D62_sgNTC_noDIFF_noRAPA_1 | 8143934 |
| DE95NGSUKBR112923 | 1 | B11 | B | 11 | D62 | sgNTC | WT | noDIFF | noRAPA | D62_sgNTC_noDIFF_noRAPA | DE95NGSUKBR112923 | 2 | D62_sgNTC_noDIFF_noRAPA_2 | 7710977 |
| DE68NGSUKBR112924 | 1 | B12 | B | 12 | D62 | sgNTC | WT | noDIFF | noRAPA | D62_sgNTC_noDIFF_noRAPA | DE68NGSUKBR112924 | 3 | D62_sgNTC_noDIFF_noRAPA_3 | 9158766 |
| DE41NGSUKBR112925 | 1 | C01 | C | 1 | D62 | sg2.1 | KO | noDIFF | RAPA | D62_sg2.1_noDIFF_RAPA | DE41NGSUKBR112925 | 1 | D62_sg2.1_noDIFF_RAPA_1 | 7883862 |
| DE14NGSUKBR112926 | 1 | C02 | C | 2 | D62 | sg2.1 | KO | noDIFF | RAPA | D62_sg2.1_noDIFF_RAPA | DE14NGSUKBR112926 | 2 | D62_sg2.1_noDIFF_RAPA_2 | 7134773 |
| DE84NGSUKBR112927 | 1 | C03 | C | 3 | D62 | sg2.1 | KO | noDIFF | RAPA | D62_sg2.1_noDIFF_RAPA | DE84NGSUKBR112927 | 3 | D62_sg2.1_noDIFF_RAPA_3 | 8617208 |
| DE57NGSUKBR112928 | 1 | C04 | C | 4 | D62 | sg2.1 | KO | noDIFF | noRAPA | D62_sg2.1_noDIFF_noRAPA | DE57NGSUKBR112928 | 1 | D62_sg2.1_noDIFF_noRAPA_1 | 7544453 |
| DE30NGSUKBR112929 | 1 | C05 | C | 5 | D62 | sg2.1 | KO | noDIFF | noRAPA | D62_sg2.1_noDIFF_noRAPA | DE30NGSUKBR112929 | 2 | D62_sg2.1_noDIFF_noRAPA_2 | 7622380 |
| DE03NGSUKBR112930 | 1 | C06 | C | 6 | D62 | sg2.1 | KO | noDIFF | noRAPA | D62_sg2.1_noDIFF_noRAPA | DE03NGSUKBR112930 | 3 | D62_sg2.1_noDIFF_noRAPA_3 | 7939374 |
| DE73NGSUKBR112931 | 1 | C07 | C | 7 | D62 | sg2.2 | KO | noDIFF | RAPA | D62_sg2.2_noDIFF_RAPA | DE73NGSUKBR112931 | 1 | D62_sg2.2_noDIFF_RAPA_1 | 7320431 |
| DE46NGSUKBR112932 | 1 | C08 | C | 8 | D62 | sg2.2 | KO | noDIFF | RAPA | D62_sg2.2_noDIFF_RAPA | DE46NGSUKBR112932 | 2 | D62_sg2.2_noDIFF_RAPA_2 | 6532522 |
| DE19NGSUKBR112933 | 1 | C09 | C | 9 | D62 | sg2.2 | KO | noDIFF | RAPA | D62_sg2.2_noDIFF_RAPA | DE19NGSUKBR112933 | 3 | D62_sg2.2_noDIFF_RAPA_3 | 7115292 |
| DE89NGSUKBR112934 | 1 | C10 | C | 10 | D62 | sg2.2 | KO | noDIFF | noRAPA | D62_sg2.2_noDIFF_noRAPA | DE89NGSUKBR112934 | 1 | D62_sg2.2_noDIFF_noRAPA_1 | 7618310 |
| DE62NGSUKBR112935 | 1 | C11 | C | 11 | D62 | sg2.2 | KO | noDIFF | noRAPA | D62_sg2.2_noDIFF_noRAPA | DE62NGSUKBR112935 | 2 | D62_sg2.2_noDIFF_noRAPA_2 | 7387684 |
| DE35NGSUKBR112936 | 1 | C12 | C | 12 | D62 | sg2.2 | KO | noDIFF | noRAPA | D62_sg2.2_noDIFF_noRAPA | DE35NGSUKBR112936 | 3 | D62_sg2.2_noDIFF_noRAPA_3 | 7440730 |
| DE08NGSUKBR112937 | 1 | D01 | D | 1 | ReN | sgNTC | WT | DIFF | RAPA | ReN_sgNTC_DIFF_RAPA | DE08NGSUKBR112937 | 1 | ReN_sgNTC_DIFF_RAPA_1 | 6178329 |
| DE78NGSUKBR112938 | 1 | D02 | D | 2 | ReN | sgNTC | WT | DIFF | RAPA | ReN_sgNTC_DIFF_RAPA | DE78NGSUKBR112938 | 2 | ReN_sgNTC_DIFF_RAPA_2 | 7406448 |
| DE51NGSUKBR112939 | 1 | D03 | D | 3 | ReN | sgNTC | WT | DIFF | RAPA | ReN_sgNTC_DIFF_RAPA | DE51NGSUKBR112939 | 3 | ReN_sgNTC_DIFF_RAPA_3 | 6338186 |
| DE24NGSUKBR112940 | 1 | D04 | D | 4 | ReN | sgNTC | WT | DIFF | noRAPA | ReN_sgNTC_DIFF_noRAPA | DE24NGSUKBR112940 | 1 | ReN_sgNTC_DIFF_noRAPA_1 | 6274167 |
| DE94NGSUKBR112941 | 1 | D05 | D | 5 | ReN | sgNTC | WT | DIFF | noRAPA | ReN_sgNTC_DIFF_noRAPA | DE94NGSUKBR112941 | 2 | ReN_sgNTC_DIFF_noRAPA_2 | 5928527 |
| DE67NGSUKBR112942 | 1 | D06 | D | 6 | ReN | sgNTC | WT | DIFF | noRAPA | ReN_sgNTC_DIFF_noRAPA | DE67NGSUKBR112942 | 3 | ReN_sgNTC_DIFF_noRAPA_3 | 6267235 |
| DE40NGSUKBR112943 | 1 | D07 | D | 7 | ReN | sg2.1 | KO | DIFF | RAPA | ReN_sg2.1_DIFF_RAPA | DE40NGSUKBR112943 | 1 | ReN_sg2.1_DIFF_RAPA_1 | 7119065 |
| DE13NGSUKBR112944 | 1 | D08 | D | 8 | ReN | sg2.1 | KO | DIFF | RAPA | ReN_sg2.1_DIFF_RAPA | DE13NGSUKBR112944 | 2 | ReN_sg2.1_DIFF_RAPA_2 | 7634764 |
| DE83NGSUKBR112945 | 1 | D09 | D | 9 | ReN | sg2.1 | KO | DIFF | RAPA | ReN_sg2.1_DIFF_RAPA | DE83NGSUKBR112945 | 3 | ReN_sg2.1_DIFF_RAPA_3 | 7137636 |
| DE56NGSUKBR112946 | 1 | D10 | D | 10 | ReN | sg2.1 | KO | DIFF | noRAPA | ReN_sg2.1_DIFF_noRAPA | DE56NGSUKBR112946 | 1 | ReN_sg2.1_DIFF_noRAPA_1 | 7764289 |
| DE29NGSUKBR112947 | 1 | D11 | D | 11 | ReN | sg2.1 | KO | DIFF | noRAPA | ReN_sg2.1_DIFF_noRAPA | DE29NGSUKBR112947 | 2 | ReN_sg2.1_DIFF_noRAPA_2 | 7877670 |
| DE02NGSUKBR112948 | 1 | D12 | D | 12 | ReN | sg2.1 | KO | DIFF | noRAPA | ReN_sg2.1_DIFF_noRAPA | DE02NGSUKBR112948 | 3 | ReN_sg2.1_DIFF_noRAPA_3 | 8604849 |
| DE72NGSUKBR112949 | 1 | E01 | E | 1 | ReN | sg2.2 | KO | DIFF | RAPA | ReN_sg2.2_DIFF_RAPA | DE72NGSUKBR112949 | 1 | ReN_sg2.2_DIFF_RAPA_1 | 6814818 |
| DE45NGSUKBR112950 | 1 | E02 | E | 2 | ReN | sg2.2 | KO | DIFF | RAPA | ReN_sg2.2_DIFF_RAPA | DE45NGSUKBR112950 | 2 | ReN_sg2.2_DIFF_RAPA_2 | 7117729 |
| DE18NGSUKBR112951 | 1 | E03 | E | 3 | ReN | sg2.2 | KO | DIFF | RAPA | ReN_sg2.2_DIFF_RAPA | DE18NGSUKBR112951 | 3 | ReN_sg2.2_DIFF_RAPA_3 | 5945129 |
| DE88NGSUKBR112952 | 1 | E04 | E | 4 | ReN | sg2.2 | KO | DIFF | noRAPA | ReN_sg2.2_DIFF_noRAPA | DE88NGSUKBR112952 | 1 | ReN_sg2.2_DIFF_noRAPA_1 | 7519852 |
| DE61NGSUKBR112953 | 1 | E05 | E | 5 | ReN | sg2.2 | KO | DIFF | noRAPA | ReN_sg2.2_DIFF_noRAPA | DE61NGSUKBR112953 | 2 | ReN_sg2.2_DIFF_noRAPA_2 | 5783946 |
| DE34NGSUKBR112954 | 1 | E06 | E | 6 | ReN | sg2.2 | KO | DIFF | noRAPA | ReN_sg2.2_DIFF_noRAPA | DE34NGSUKBR112954 | 3 | ReN_sg2.2_DIFF_noRAPA_3 | 6347263 |
| DE07NGSUKBR112955 | 1 | E07 | E | 7 | ReN | sgNTC | WT | noDIFF | RAPA | ReN_sgNTC_noDIFF_RAPA | DE07NGSUKBR112955 | 1 | ReN_sgNTC_noDIFF_RAPA_1 | 7676047 |
| DE77NGSUKBR112956 | 1 | E08 | E | 8 | ReN | sgNTC | WT | noDIFF | RAPA | ReN_sgNTC_noDIFF_RAPA | DE77NGSUKBR112956 | 2 | ReN_sgNTC_noDIFF_RAPA_2 | 6505596 |
| DE50NGSUKBR112957 | 1 | E09 | E | 9 | ReN | sgNTC | WT | noDIFF | RAPA | ReN_sgNTC_noDIFF_RAPA | DE50NGSUKBR112957 | 3 | ReN_sgNTC_noDIFF_RAPA_3 | 6630480 |
| DE23NGSUKBR112958 | 1 | E10 | E | 10 | ReN | sgNTC | WT | noDIFF | noRAPA | ReN_sgNTC_noDIFF_noRAPA | DE23NGSUKBR112958 | 1 | ReN_sgNTC_noDIFF_noRAPA_1 | 6984521 |
| DE93NGSUKBR112959 | 1 | E11 | E | 11 | ReN | sgNTC | WT | noDIFF | noRAPA | ReN_sgNTC_noDIFF_noRAPA | DE93NGSUKBR112959 | 2 | ReN_sgNTC_noDIFF_noRAPA_2 | 8152264 |
| DE66NGSUKBR112960 | 1 | E12 | E | 12 | ReN | sgNTC | WT | noDIFF | noRAPA | ReN_sgNTC_noDIFF_noRAPA | DE66NGSUKBR112960 | 3 | ReN_sgNTC_noDIFF_noRAPA_3 | 7595849 |
| DE39NGSUKBR112961 | 1 | F01 | F | 1 | ReN | sg2.1 | KO | noDIFF | RAPA | ReN_sg2.1_noDIFF_RAPA | DE39NGSUKBR112961 | 1 | ReN_sg2.1_noDIFF_RAPA_1 | 6704413 |
| DE12NGSUKBR112962 | 1 | F02 | F | 2 | ReN | sg2.1 | KO | noDIFF | RAPA | ReN_sg2.1_noDIFF_RAPA | DE12NGSUKBR112962 | 2 | ReN_sg2.1_noDIFF_RAPA_2 | 5751386 |
| DE82NGSUKBR112963 | 1 | F03 | F | 3 | ReN | sg2.1 | KO | noDIFF | RAPA | ReN_sg2.1_noDIFF_RAPA | DE82NGSUKBR112963 | 3 | ReN_sg2.1_noDIFF_RAPA_3 | 5811217 |
| DE55NGSUKBR112964 | 1 | F04 | F | 4 | ReN | sg2.1 | KO | noDIFF | noRAPA | ReN_sg2.1_noDIFF_noRAPA | DE55NGSUKBR112964 | 1 | ReN_sg2.1_noDIFF_noRAPA_1 | 7189688 |
| DE28NGSUKBR112965 | 1 | F05 | F | 5 | ReN | sg2.1 | KO | noDIFF | noRAPA | ReN_sg2.1_noDIFF_noRAPA | DE28NGSUKBR112965 | 2 | ReN_sg2.1_noDIFF_noRAPA_2 | 6200389 |
| DE98NGSUKBR112966 | 1 | F06 | F | 6 | ReN | sg2.1 | KO | noDIFF | noRAPA | ReN_sg2.1_noDIFF_noRAPA | DE98NGSUKBR112966 | 3 | ReN_sg2.1_noDIFF_noRAPA_3 | 7186721 |
| DE71NGSUKBR112967 | 1 | F07 | F | 7 | ReN | sg2.2 | KO | noDIFF | RAPA | ReN_sg2.2_noDIFF_RAPA | DE71NGSUKBR112967 | 1 | ReN_sg2.2_noDIFF_RAPA_1 | 6307822 |
| DE44NGSUKBR112968 | 1 | F08 | F | 8 | ReN | sg2.2 | KO | noDIFF | RAPA | ReN_sg2.2_noDIFF_RAPA | DE44NGSUKBR112968 | 2 | ReN_sg2.2_noDIFF_RAPA_2 | 6891060 |
| DE17NGSUKBR112969 | 1 | F09 | F | 9 | ReN | sg2.2 | KO | noDIFF | RAPA | ReN_sg2.2_noDIFF_RAPA | DE17NGSUKBR112969 | 3 | ReN_sg2.2_noDIFF_RAPA_3 | 6789988 |
| DE87NGSUKBR112970 | 1 | F10 | F | 10 | ReN | sg2.2 | KO | noDIFF | noRAPA | ReN_sg2.2_noDIFF_noRAPA | DE87NGSUKBR112970 | 1 | ReN_sg2.2_noDIFF_noRAPA_1 | 7050910 |
| DE60NGSUKBR112971 | 1 | F11 | F | 11 | ReN | sg2.2 | KO | noDIFF | noRAPA | ReN_sg2.2_noDIFF_noRAPA | DE60NGSUKBR112971 | 2 | ReN_sg2.2_noDIFF_noRAPA_2 | 8331662 |
| DE33NGSUKBR112972 | 1 | F12 | F | 12 | ReN | sg2.2 | KO | noDIFF | noRAPA | ReN_sg2.2_noDIFF_noRAPA | DE33NGSUKBR112972 | 3 | ReN_sg2.2_noDIFF_noRAPA_3 | 8271624 |
| DE06NGSUKBR112973 | 1 | G01 | G | 1 | D244 | sgNTC | WT | DIFF | RAPA | D244_sgNTC_DIFF_RAPA | DE06NGSUKBR112973 | 1 | D244_sgNTC_DIFF_RAPA_1 | 6649365 |
| DE76NGSUKBR112974 | 1 | G02 | G | 2 | D244 | sgNTC | WT | DIFF | RAPA | D244_sgNTC_DIFF_RAPA | DE76NGSUKBR112974 | 2 | D244_sgNTC_DIFF_RAPA_2 | 7645043 |
| DE49NGSUKBR112975 | 1 | G03 | G | 3 | D244 | sgNTC | WT | DIFF | RAPA | D244_sgNTC_DIFF_RAPA | DE49NGSUKBR112975 | 3 | D244_sgNTC_DIFF_RAPA_3 | 7704010 |
| DE22NGSUKBR112976 | 1 | G04 | G | 4 | D244 | sgNTC | WT | DIFF | noRAPA | D244_sgNTC_DIFF_noRAPA | DE22NGSUKBR112976 | 1 | D244_sgNTC_DIFF_noRAPA_1 | 2696422 |
| DE92NGSUKBR112977 | 1 | G05 | G | 5 | D244 | sgNTC | WT | DIFF | noRAPA | D244_sgNTC_DIFF_noRAPA | DE92NGSUKBR112977 | 2 | D244_sgNTC_DIFF_noRAPA_2 | 7988548 |
| DE65NGSUKBR112978 | 1 | G06 | G | 6 | D244 | sgNTC | WT | DIFF | noRAPA | D244_sgNTC_DIFF_noRAPA | DE65NGSUKBR112978 | 3 | D244_sgNTC_DIFF_noRAPA_3 | 5755497 |
| DE38NGSUKBR112979 | 1 | G07 | G | 7 | D244 | sg2.1 | KO | DIFF | RAPA | D244_sg2.1_DIFF_RAPA | DE38NGSUKBR112979 | 1 | D244_sg2.1_DIFF_RAPA_1 | 9078711 |
| DE11NGSUKBR112980 | 1 | G08 | G | 8 | D244 | sg2.1 | KO | DIFF | RAPA | D244_sg2.1_DIFF_RAPA | DE11NGSUKBR112980 | 2 | D244_sg2.1_DIFF_RAPA_2 | 8222243 |
| DE81NGSUKBR112981 | 1 | G09 | G | 9 | D244 | sg2.1 | KO | DIFF | RAPA | D244_sg2.1_DIFF_RAPA | DE81NGSUKBR112981 | 3 | D244_sg2.1_DIFF_RAPA_3 | 6513540 |
| DE54NGSUKBR112982 | 1 | G10 | G | 10 | D244 | sg2.1 | KO | DIFF | noRAPA | D244_sg2.1_DIFF_noRAPA | DE54NGSUKBR112982 | 1 | D244_sg2.1_DIFF_noRAPA_1 | 7754476 |
| DE27NGSUKBR112983 | 1 | G11 | G | 11 | D244 | sg2.1 | KO | DIFF | noRAPA | D244_sg2.1_DIFF_noRAPA | DE27NGSUKBR112983 | 2 | D244_sg2.1_DIFF_noRAPA_2 | 8398511 |
| DE97NGSUKBR112984 | 1 | G12 | G | 12 | D244 | sg2.1 | KO | DIFF | noRAPA | D244_sg2.1_DIFF_noRAPA | DE97NGSUKBR112984 | 3 | D244_sg2.1_DIFF_noRAPA_3 | 7432891 |
| DE70NGSUKBR112985 | 1 | H01 | H | 1 | D244 | sg2.2 | KO | DIFF | RAPA | D244_sg2.2_DIFF_RAPA | DE70NGSUKBR112985 | 1 | D244_sg2.2_DIFF_RAPA_1 | 8760362 |
| DE43NGSUKBR112986 | 1 | H02 | H | 2 | D244 | sg2.2 | KO | DIFF | RAPA | D244_sg2.2_DIFF_RAPA | DE43NGSUKBR112986 | 2 | D244_sg2.2_DIFF_RAPA_2 | 8888690 |
| DE16NGSUKBR112987 | 1 | H03 | H | 3 | D244 | sg2.2 | KO | DIFF | RAPA | D244_sg2.2_DIFF_RAPA | DE16NGSUKBR112987 | 3 | D244_sg2.2_DIFF_RAPA_3 | 8598242 |
| DE86NGSUKBR112988 | 1 | H04 | H | 4 | D244 | sg2.2 | KO | DIFF | noRAPA | D244_sg2.2_DIFF_noRAPA | DE86NGSUKBR112988 | 1 | D244_sg2.2_DIFF_noRAPA_1 | 9274217 |
| DE59NGSUKBR112989 | 1 | H05 | H | 5 | D244 | sg2.2 | KO | DIFF | noRAPA | D244_sg2.2_DIFF_noRAPA | DE59NGSUKBR112989 | 2 | D244_sg2.2_DIFF_noRAPA_2 | 8086371 |
| DE32NGSUKBR112990 | 1 | H06 | H | 6 | D244 | sg2.2 | KO | DIFF | noRAPA | D244_sg2.2_DIFF_noRAPA | DE32NGSUKBR112990 | 3 | D244_sg2.2_DIFF_noRAPA_3 | 6898881 |
| DE05NGSUKBR112991 | 1 | H07 | H | 7 | D244 | sgNTC | WT | noDIFF | RAPA | D244_sgNTC_noDIFF_RAPA | DE05NGSUKBR112991 | 1 | D244_sgNTC_noDIFF_RAPA_1 | 7527521 |
| DE75NGSUKBR112992 | 1 | H08 | H | 8 | D244 | sgNTC | WT | noDIFF | RAPA | D244_sgNTC_noDIFF_RAPA | DE75NGSUKBR112992 | 2 | D244_sgNTC_noDIFF_RAPA_2 | 7444314 |
| DE48NGSUKBR112993 | 1 | H09 | H | 9 | D244 | sgNTC | WT | noDIFF | RAPA | D244_sgNTC_noDIFF_RAPA | DE48NGSUKBR112993 | 3 | D244_sgNTC_noDIFF_RAPA_3 | 6600900 |
| DE21NGSUKBR112994 | 1 | H10 | H | 10 | D244 | sgNTC | WT | noDIFF | noRAPA | D244_sgNTC_noDIFF_noRAPA | DE21NGSUKBR112994 | 1 | D244_sgNTC_noDIFF_noRAPA_1 | 4543120 |
| DE91NGSUKBR112995 | 1 | H11 | H | 11 | D244 | sgNTC | WT | noDIFF | noRAPA | D244_sgNTC_noDIFF_noRAPA | DE91NGSUKBR112995 | 2 | D244_sgNTC_noDIFF_noRAPA_2 | 8465146 |
| DE64NGSUKBR112996 | 1 | H12 | H | 12 | D244 | sgNTC | WT | noDIFF | noRAPA | D244_sgNTC_noDIFF_noRAPA | DE64NGSUKBR112996 | 3 | D244_sgNTC_noDIFF_noRAPA_3 | 8262382 |
| DE37NGSUKBR112997 | 2 | A1 | A | 1 | D244 | sg2.1 | KO | noDIFF | RAPA | D244_sg2.1_noDIFF_RAPA | DE37NGSUKBR112997 | 1 | D244_sg2.1_noDIFF_RAPA_1 | 6632796 |
| DE10NGSUKBR112998 | 2 | A2 | A | 2 | D244 | sg2.1 | KO | noDIFF | RAPA | D244_sg2.1_noDIFF_RAPA | DE10NGSUKBR112998 | 2 | D244_sg2.1_noDIFF_RAPA_2 | 6565291 |
| DE80NGSUKBR112999 | 2 | A3 | A | 3 | D244 | sg2.1 | KO | noDIFF | RAPA | D244_sg2.1_noDIFF_RAPA | DE80NGSUKBR112999 | 3 | D244_sg2.1_noDIFF_RAPA_3 | 7135203 |
| DE53NGSUKBR113000 | 2 | A4 | A | 4 | D244 | sg2.1 | KO | noDIFF | noRAPA | D244_sg2.1_noDIFF_noRAPA | DE53NGSUKBR113000 | 1 | D244_sg2.1_noDIFF_noRAPA_1 | 6769717 |
| DE26NGSUKBR113001 | 2 | A5 | A | 5 | D244 | sg2.1 | KO | noDIFF | noRAPA | D244_sg2.1_noDIFF_noRAPA | DE26NGSUKBR113001 | 2 | D244_sg2.1_noDIFF_noRAPA_2 | 6858477 |
| DE96NGSUKBR113002 | 2 | A6 | A | 6 | D244 | sg2.1 | KO | noDIFF | noRAPA | D244_sg2.1_noDIFF_noRAPA | DE96NGSUKBR113002 | 3 | D244_sg2.1_noDIFF_noRAPA_3 | 7163942 |
| DE69NGSUKBR113003 | 2 | A7 | A | 7 | D244 | sg2.2 | KO | noDIFF | RAPA | D244_sg2.2_noDIFF_RAPA | DE69NGSUKBR113003 | 1 | D244_sg2.2_noDIFF_RAPA_1 | 6244331 |
| DE42NGSUKBR113004 | 2 | A8 | A | 8 | D244 | sg2.2 | KO | noDIFF | RAPA | D244_sg2.2_noDIFF_RAPA | DE42NGSUKBR113004 | 2 | D244_sg2.2_noDIFF_RAPA_2 | 5077409 |
| DE15NGSUKBR113005 | 2 | A9 | A | 9 | D244 | sg2.2 | KO | noDIFF | RAPA | D244_sg2.2_noDIFF_RAPA | DE15NGSUKBR113005 | 3 | D244_sg2.2_noDIFF_RAPA_3 | 6114278 |
| DE85NGSUKBR113006 | 2 | A10 | A | 10 | D244 | sg2.2 | KO | noDIFF | noRAPA | D244_sg2.2_noDIFF_noRAPA | DE85NGSUKBR113006 | 1 | D244_sg2.2_noDIFF_noRAPA_1 | 6591276 |
| DE58NGSUKBR113007 | 2 | A11 | A | 11 | D244 | sg2.2 | KO | noDIFF | noRAPA | D244_sg2.2_noDIFF_noRAPA | DE58NGSUKBR113007 | 2 | D244_sg2.2_noDIFF_noRAPA_2 | 7619920 |
| DE31NGSUKBR113008 | 2 | A12 | A | 12 | D244 | sg2.2 | KO | noDIFF | noRAPA | D244_sg2.2_noDIFF_noRAPA | DE31NGSUKBR113008 | 3 | D244_sg2.2_noDIFF_noRAPA_3 | 6147586 |
Total number of samples overlapping between Counts and SampleInfo: 108
boxplot_counts = function(plotsubset, maintitle, colorcode) {
par(mar=c(3,3,5,7))
a =boxplot(log2(plotsubset+1), main = maintitle,
col = Dark8[as.factor(SampleInfo[,colorcode])], names=NA,
ylab = "log2 transformed", xlab="samples")
legend("bottomleft", legend = levels(SampleInfo[,colorcode]),
bty = "n", bg="white",
pch = 16, col = Dark8[1:length(unique(SampleInfo[,colorcode]))])
}
barplot_counts = function(DF, maintitle, colorcode) {
barplot(log2(DF[,"reads_per_sample"]), main = maintitle,
col = Dark8[as.factor(DF[,colorcode])], names="",
ylab = "log2 transformed",xlab="samples")
legend("bottomleft", legend = levels(DF[,colorcode]), pch = 16,
bty = "n", bg="white",
col = Dark8[1:length(unique(DF[,colorcode]))])
}
boxplot_counts(Countdata, "raw counts", "CellLine")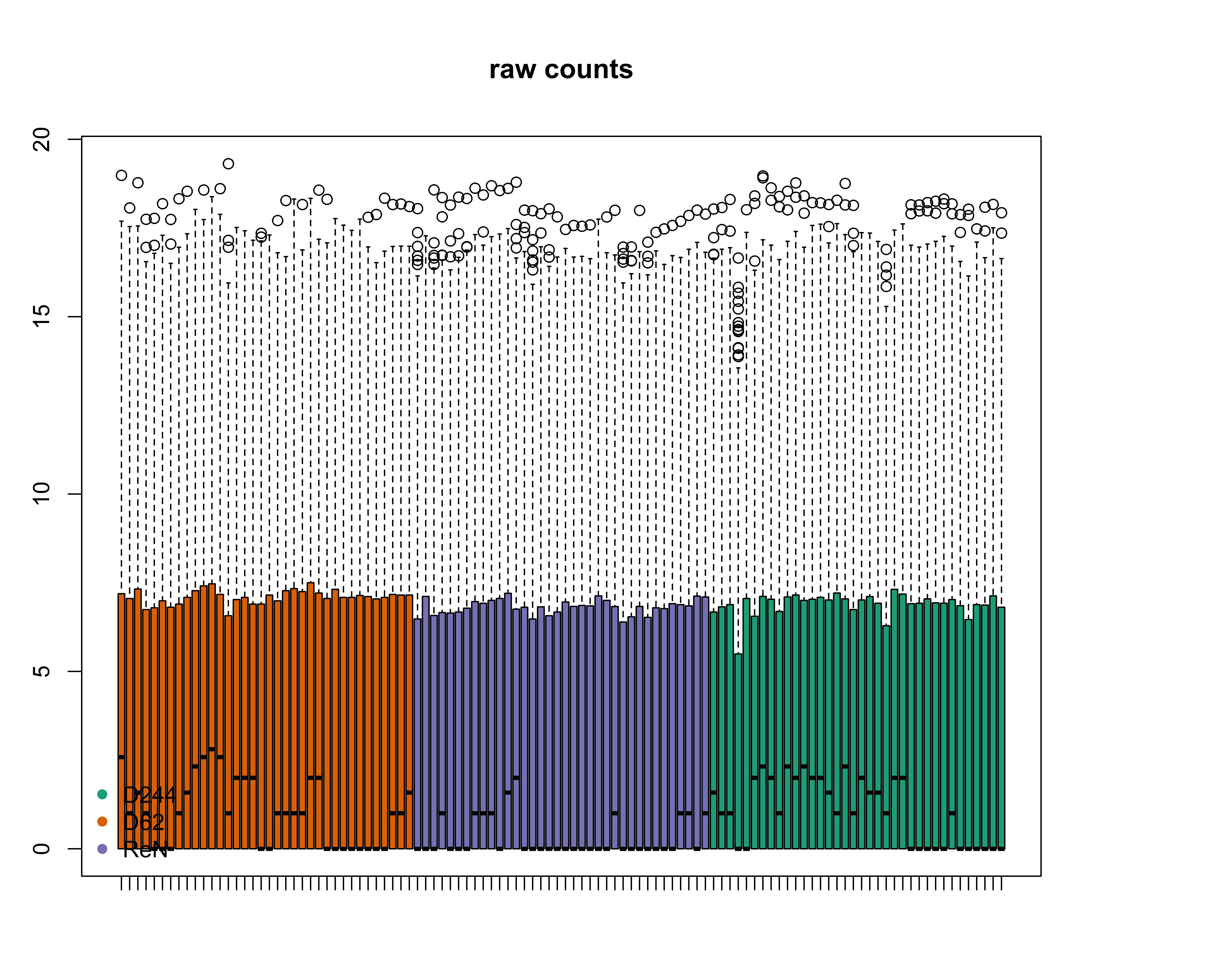
boxplot_counts(Countdata, "raw counts", "gRNA")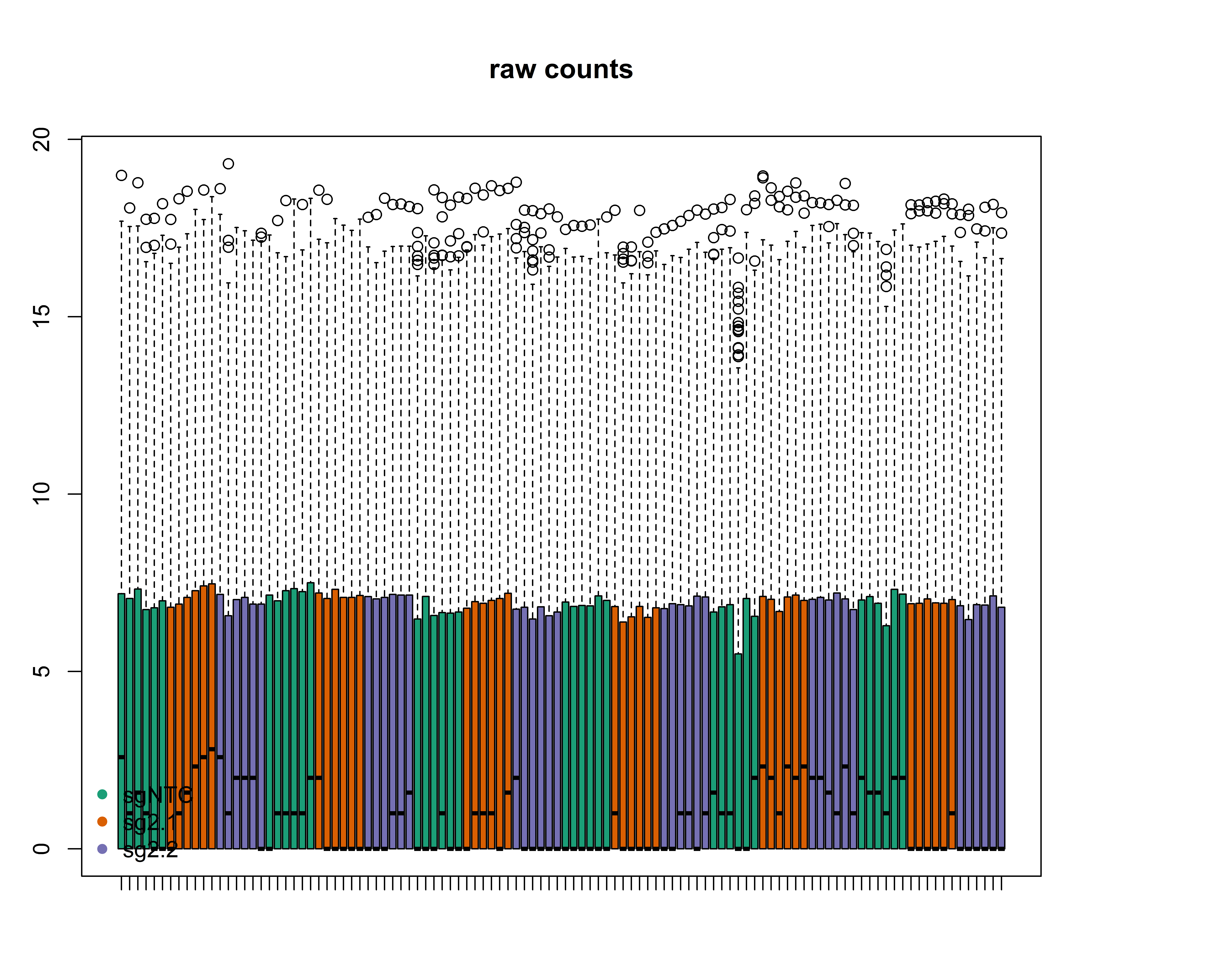
barplot_counts(SampleInfo, "total reads", "CellLine")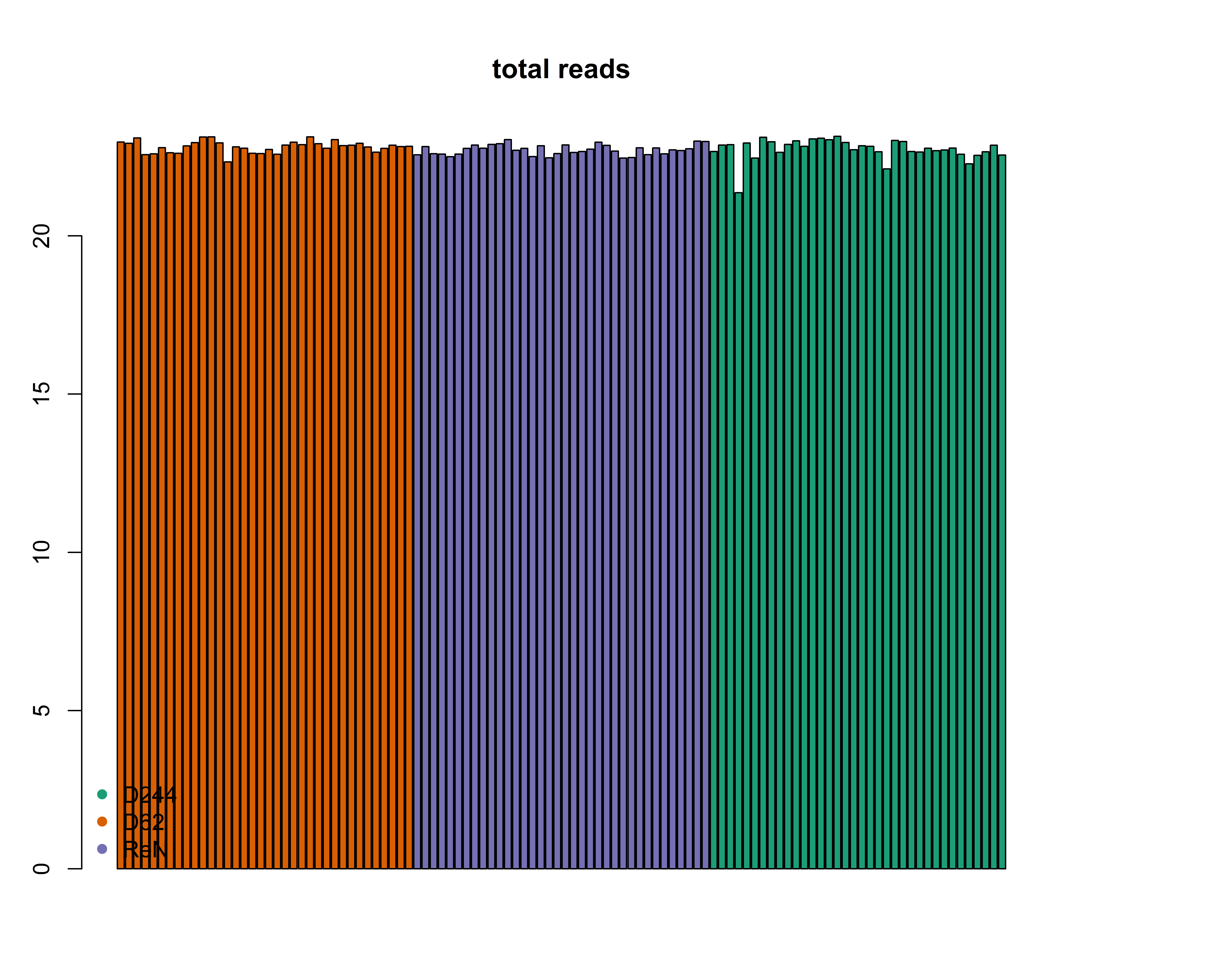
barplot_counts(SampleInfo, "total reads", "gRNA")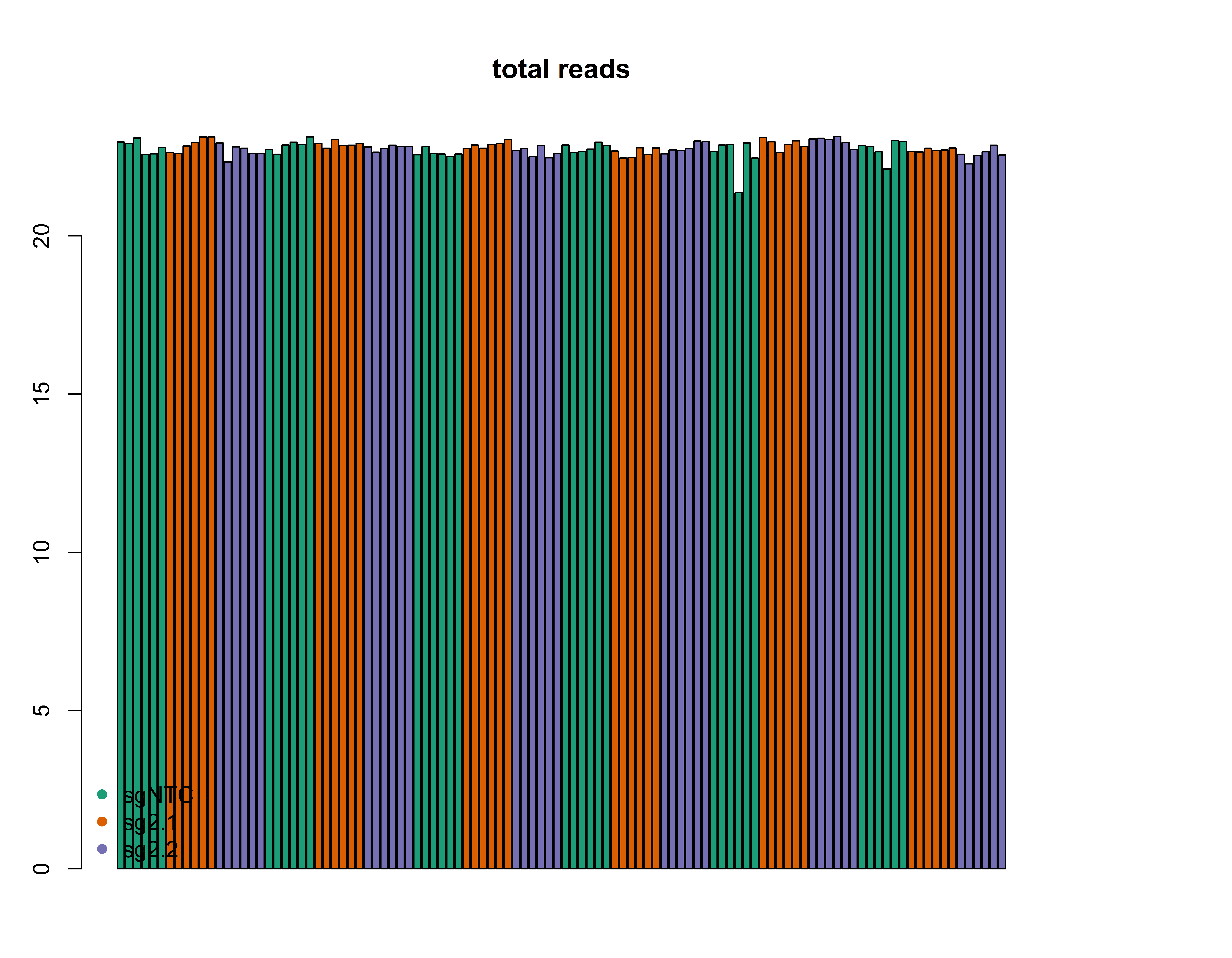
plot(density(log2(rowMeans(Countdata))), main="distribution of gene expression",
xlab="mean log2(counts +1)")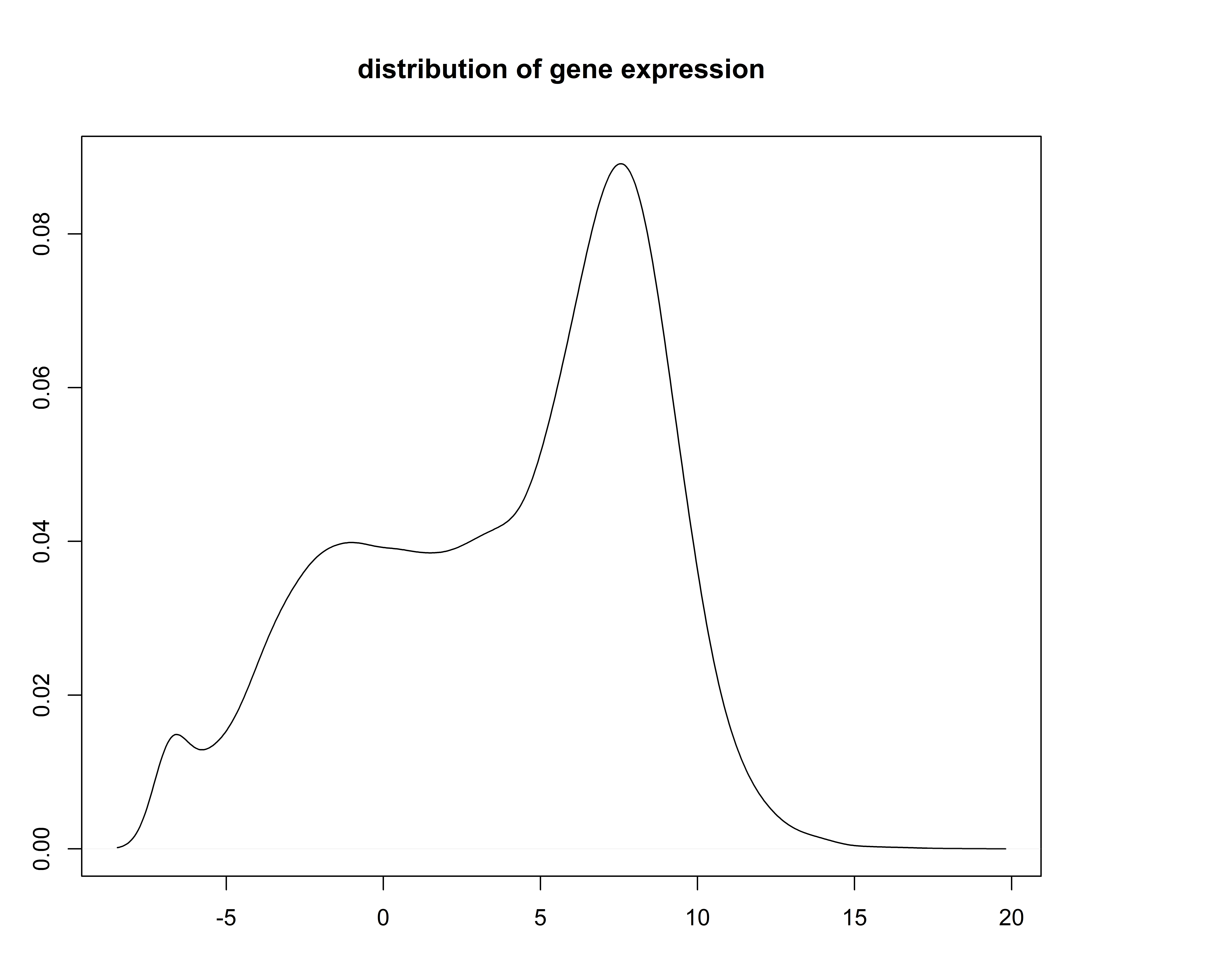
# remove genes wich were not detected in at least 50% of the samples
keeperidx = rowSums(Countdata>1)>nrow(SampleInfo)/2
Countdata_cl = Countdata[keeperidx, ]
rowdescription = rowdescription[as.character(row.names(Countdata_cl)),]
fullmodel = as.formula("~CellLine+gRNA+DIFF+RAPA")
ddsMat <- DESeqDataSetFromMatrix(countData = Countdata_cl,
colData = SampleInfo,
rowData = rowdescription,
design = fullmodel)
ddsMat = estimateSizeFactors(ddsMat)
ddsMat = estimateDispersions(ddsMat)gene-wise dispersion estimatesmean-dispersion relationshipfinal dispersion estimatesreads = as.data.frame(counts(ddsMat, normalized=T))
SDs = apply(reads, 1, sd)
keepvar = SDs>0
ddsMat <- ddsMat[keepvar,]
Nfilt = length(ddsMat)
reads = as.data.frame(counts(ddsMat, normalized=T))
SampleInfo$reads_per_sample_cl= colSums(reads)before cleaning
- Average reads per samples: 7 187 519
- Standard deviation reads per samples: 1 050 507
- Total genes mapped: 28 395
after cleaning
- Average reads per samples: 6 902 029
- Standard deviation reads per samples: 491 463.8
- Genes removed due to low reads: 14 847
- Total genes included after filtering: 13 548
Clustering
hierarchical clustering based on the top 2000 genes by variance
log2_cpm = log2(reads+1)
varsset=apply(log2_cpm, 1, var)
cpm.sel.trans = t(log2_cpm[order(varsset,decreasing = T)[1:2000],])
rownames(cpm.sel.trans)=SampleInfo$label_rep
distance = dist(cpm.sel.trans)
hc = stats::hclust(distance, method="ward.D2")
cutN=24
clusters = cutree(hc, k=cutN)
Colors=sample(jetcolors(cutN))[clusters]
myLetters <- LETTERS[1:26]
numRow=match(SampleInfo$Row, myLetters)
numRow=numRow+(SampleInfo$Plate-1)*8
addRow=LETTERS[numRow]
Plotdata=data.frame(Rows=addRow, numRow = numRow, Cols = SampleInfo$Col,
Group=clusters, Colors=Colors)
par(mar=c(15,3,5,3))
plot(as.dendrogram(hc), main=paste("Similairtiy by gene expression, guessed",cutN,"clusters"), cex=0.7)
colored_dots(colors = Colors, dend = as.dendrogram(hc), rowLabels = "cluster")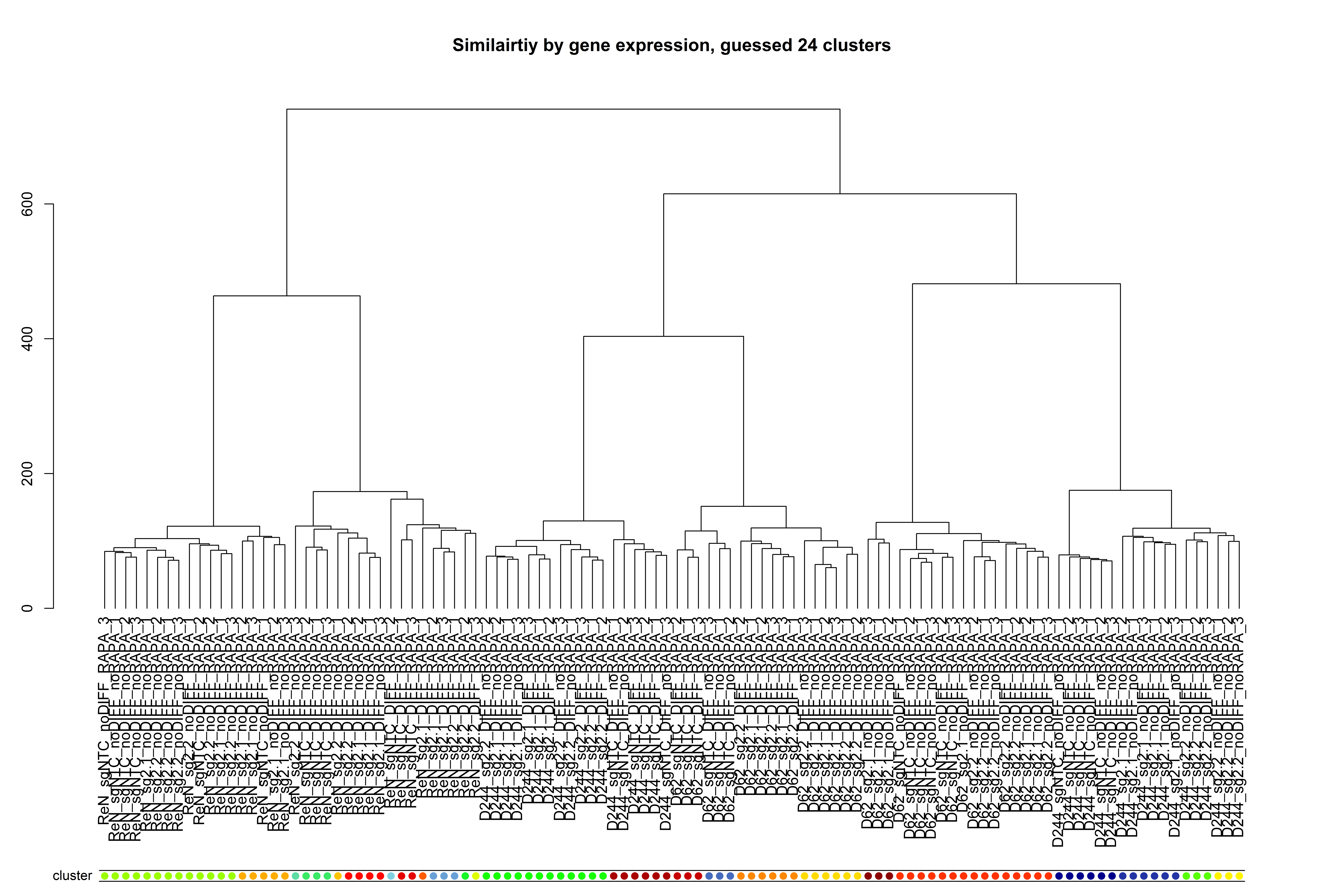
Similarity based on hcluster plot
par(mar=c(2,5,8,3))
plot(0,0, type="n", ylab="", xlab="",
ylim=rev(range(Plotdata$numRow))+c(1,-1),
xlim=range(Plotdata$Cols)+c(-1,1), xaxt="n",yaxt="n" ,
main="plate similarity plot")
points(y=Plotdata$numRow, x=Plotdata$Cols, pch=16, cex=4, col=Plotdata$Colors)
text(y=Plotdata$numRow, x=Plotdata$Cols, labels = Plotdata$Group)
text(y=Plotdata$numRow, x=Plotdata$Cols, labels = Plotdata$Group)
axis(2, at=1:9, labels = c(paste0("P1_", LETTERS[1:8]), "P2_A"), las=1)
axis(3, at=1:12, labels = c(paste0("Col_", 1:12)), las=3)
abline(h=8.5)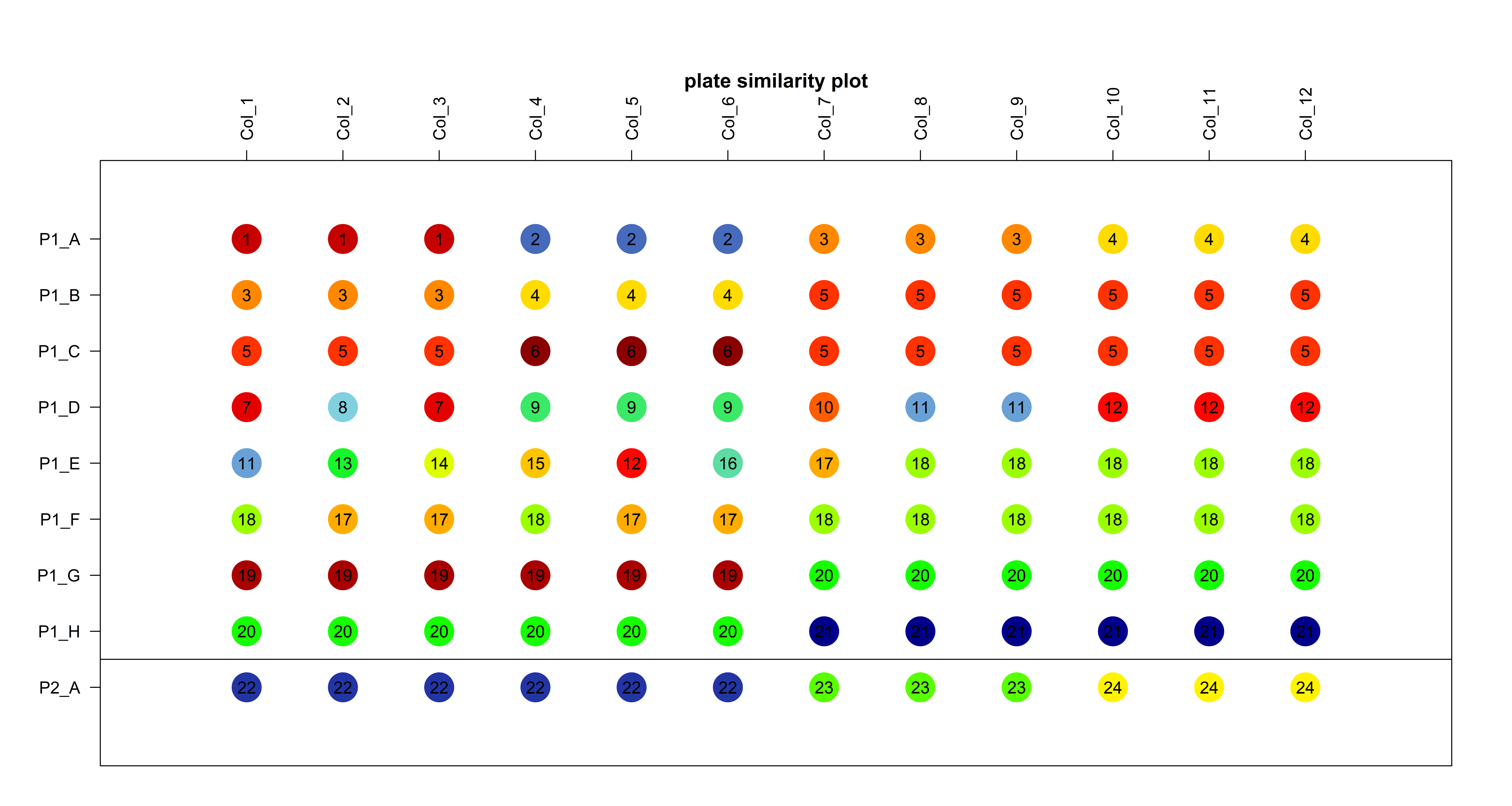
sampleDistMatrix <- as.matrix(distance)
#colors for plotting heatmap
colors <- colorRampPalette(brewer.pal(9, "Spectral"))(255)
cellcol = Dark8[1:nlevels(SampleInfo$CellLine)]
names(cellcol) = levels(SampleInfo$CellLine)
gRNAcol = Dark8[c(1:nlevels(SampleInfo$gRNA))+nlevels(SampleInfo$CellLine)]
names(gRNAcol) = levels(SampleInfo$gRNA)
diffcol = brewer.pal(3,"Set1")[1:nlevels(SampleInfo$DIFF)]
names(diffcol) = levels(SampleInfo$DIFF)
rapacol = brewer.pal(3,"Set2")[1:nlevels(SampleInfo$RAPA)]
names(rapacol) = levels(SampleInfo$RAPA)
ann_colors = list(
DIFF = diffcol,
RAPA = rapacol,
gRNA = gRNAcol,
CellLine=cellcol)
labels = SampleInfo[,c("CellLine","gRNA","DIFF", "RAPA")] %>%
mutate_all(as.character) %>% as.data.frame()
rownames(labels)=SampleInfo$label_rep
pheatmap(sampleDistMatrix,
clustering_distance_rows = distance,
clustering_distance_cols = distance,
clustering_method = "ward.D2",
scale ="row",
border_color = NA,
annotation_row = labels,
annotation_col = labels,
annotation_colors = ann_colors,
col = colors,
main = "Distances normalized log2 counts")
save(ddsMat, file=paste0(output,"/dds_matrix.RData"))PCA and MDS
# PCA
gpca <- glmpca(t(cpm.sel.trans), L = 2)
gpca.dat <- gpca$factors
gpca.dat$CellLine <- SampleInfo$CellLine
gpca.dat$gRNA <- SampleInfo$gRNA
gpca.dat$KO<- SampleInfo$KO
gpca.dat$DIFF <- SampleInfo$DIFF
gpca.dat$RAPA<- SampleInfo$RAPA
rownames(gpca.dat) = SampleInfo$labels
mds = as.data.frame(SampleInfo) %>% cbind(cmdscale(distance))
save(mds, gpca.dat, file=paste0(home, "/analysis/MDSplots/mdsplots.RData"))
#
# ggplot(gpca.dat, aes(x = dim1, y = dim2, color = CellLine, shape = DIFF)) +
# geom_point(size = 2) + ggtitle("PCA with log2 counts")
#
# ggplot(mds, aes(x = `1`, y = `2`, color = CellLine, shape = DIFF)) +
# geom_point(size = 2) + ggtitle("MDS with log2 counts")
rsconnect::setAccountInfo(name='molgenlab',
token='86875F8B6550C3A26488035E69B1F18D',
secret=shinySECRET)
rsconnect::deployApp(paste0(home, "/analysis/MDSplots"))Preparing to deploy application...DONE
Uploading bundle for application: 3794945...DONE
Deploying bundle: 5601810 for application: 3794945 ...
Waiting for task: 1104549867
building: Building image: 6510542
building: Fetching packages
building: Installing packages
building: Installing files
building: Pushing image: 6510542
deploying: Starting instances
rollforward: Activating new instances
unstaging: Stopping old instances
Application successfully deployed to https://molgenlab.shinyapps.io/mdsplots/
sessionInfo()R version 4.1.2 (2021-11-01)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 18363)
Matrix products: default
locale:
[1] LC_COLLATE=German_Germany.1252 LC_CTYPE=German_Germany.1252
[3] LC_MONETARY=German_Germany.1252 LC_NUMERIC=C
[5] LC_TIME=German_Germany.1252
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] gprofiler2_0.2.1 dendextend_1.15.2
[3] glmpca_0.2.0 RCurl_1.98-1.5
[5] knitr_1.37 DESeq2_1.34.0
[7] SummarizedExperiment_1.24.0 Biobase_2.54.0
[9] MatrixGenerics_1.6.0 matrixStats_0.61.0
[11] GenomicRanges_1.46.1 GenomeInfoDb_1.30.1
[13] IRanges_2.28.0 S4Vectors_0.32.3
[15] BiocGenerics_0.40.0 pheatmap_1.0.12
[17] RColorBrewer_1.1-2 compareGroups_4.5.1
[19] forcats_0.5.1 stringr_1.4.0
[21] dplyr_1.0.7 purrr_0.3.4
[23] readr_2.1.2 tidyr_1.1.4
[25] tibble_3.1.6 ggplot2_3.3.5
[27] tidyverse_1.3.1 kableExtra_1.3.4
[29] workflowr_1.7.0
loaded via a namespace (and not attached):
[1] readxl_1.3.1 uuid_1.0-3 backports_1.4.1
[4] systemfonts_1.0.3 lazyeval_0.2.2 splines_4.1.2
[7] BiocParallel_1.28.3 digest_0.6.29 htmltools_0.5.2
[10] rsconnect_0.8.25 viridis_0.6.2 fansi_1.0.2
[13] magrittr_2.0.2 Rsolnp_1.16 memoise_2.0.1
[16] tzdb_0.2.0 Biostrings_2.62.0 annotate_1.72.0
[19] modelr_0.1.8 officer_0.4.1 svglite_2.0.0
[22] askpass_1.1 colorspace_2.0-2 blob_1.2.2
[25] rvest_1.0.2 haven_2.4.3 xfun_0.29
[28] callr_3.7.0 crayon_1.4.2 jsonlite_1.7.3
[31] genefilter_1.76.0 survival_3.2-13 glue_1.6.1
[34] gtable_0.3.0 zlibbioc_1.40.0 XVector_0.34.0
[37] webshot_0.5.2 DelayedArray_0.20.0 scales_1.1.1
[40] DBI_1.1.2 Rcpp_1.0.8 viridisLite_0.4.0
[43] xtable_1.8-4 bit_4.0.4 truncnorm_1.0-8
[46] htmlwidgets_1.5.4 httr_1.4.2 ellipsis_0.3.2
[49] mice_3.14.0 farver_2.1.0 pkgconfig_2.0.3
[52] XML_3.99-0.8 nnet_7.3-17 sass_0.4.0
[55] dbplyr_2.1.1 locfit_1.5-9.4 utf8_1.2.2
[58] tidyselect_1.1.1 rlang_1.0.0 later_1.3.0
[61] AnnotationDbi_1.56.2 munsell_0.5.0 cellranger_1.1.0
[64] tools_4.1.2 cachem_1.0.6 cli_3.1.1
[67] generics_0.1.2 RSQLite_2.2.9 broom_0.7.12
[70] evaluate_0.14 fastmap_1.1.0 yaml_2.2.2
[73] processx_3.5.2 bit64_4.0.5 fs_1.5.2
[76] zip_2.2.0 packrat_0.7.0 KEGGREST_1.34.0
[79] whisker_0.4 xml2_1.3.3 compiler_4.1.2
[82] rstudioapi_0.13 curl_4.3.2 plotly_4.10.0
[85] png_0.1-7 reprex_2.0.1 geneplotter_1.72.0
[88] bslib_0.3.1 stringi_1.7.6 HardyWeinberg_1.7.4
[91] highr_0.9 ps_1.6.0 gdtools_0.2.3
[94] lattice_0.20-45 Matrix_1.4-0 vctrs_0.3.8
[97] pillar_1.7.0 lifecycle_1.0.1 BiocManager_1.30.16
[100] jquerylib_0.1.4 data.table_1.14.2 bitops_1.0-7
[103] flextable_0.6.10 httpuv_1.6.5 R6_2.5.1
[106] promises_1.2.0.1 gridExtra_2.3 writexl_1.4.0
[109] MASS_7.3-55 assertthat_0.2.1 chron_2.3-56
[112] openssl_1.4.6 rprojroot_2.0.2 withr_2.4.3
[115] GenomeInfoDbData_1.2.7 parallel_4.1.2 hms_1.1.1
[118] grid_4.1.2 rmarkdown_2.11 git2r_0.29.0
[121] getPass_0.2-2 lubridate_1.8.0 base64enc_0.1-3