WGCNA
AGC, AY
22 3 2021
Last updated: 2022-02-21
Checks: 7 0
Knit directory: CePTER_RNASeq/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20210315) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 56805fa. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/.Rhistory
Ignored: analysis/MDSplots/D62_mdsplots.RData
Ignored: data/.Rhistory
Ignored: data/Countmatrix.RData
Ignored: oldcode/
Ignored: output/D62_GOresWGCNA.xlsx
Ignored: output/D62_ResTabs_KO.RData
Ignored: output/D62_ResTabs_KO_WGCNA.RData
Ignored: output/D62_WGCNA_adj_TOM.RData
Ignored: output/D62_dds_matrix.RData
Ignored: output/GOres_Comparisons.xlsx
Ignored: output/GOres_D244DIFFRAPA.xlsx
Ignored: output/GOres_D244DIFFnoRAPA.xlsx
Ignored: output/GOres_D244noDIFFRAPA.xlsx
Ignored: output/GOres_D244noDIFFnoRAPA.xlsx
Ignored: output/GOres_D62DIFFRAPA.xlsx
Ignored: output/GOres_D62DIFFnoRAPA.xlsx
Ignored: output/GOres_D62noDIFFRAPA.xlsx
Ignored: output/GOres_D62noDIFFnoRAPA.xlsx
Ignored: output/GOres_DIFFRAPA.xlsx
Ignored: output/GOres_DIFFnoRAPA.xlsx
Ignored: output/GOres_ReNDIFFRAPA.xlsx
Ignored: output/GOres_ReNDIFFnoRAPA.xlsx
Ignored: output/GOres_ReNnoDIFFRAPA.xlsx
Ignored: output/GOres_ReNnoDIFFnoRAPA.xlsx
Ignored: output/GOres_noDIFFRAPA.xlsx
Ignored: output/GOres_noDIFFnoRAPA.xlsx
Ignored: output/ResTabs_KO.RData
Ignored: output/ResTabs_KO_WGCNA.RData
Ignored: output/Restab_D244_DIFF_RAPA.xlsx
Ignored: output/Restab_D244_DIFF_noRAPA.xlsx
Ignored: output/Restab_D244_noDIFF_RAPA.xlsx
Ignored: output/Restab_D244_noDIFF_noRAPA.xlsx
Ignored: output/Restab_D62_DIFF_RAPA.xlsx
Ignored: output/Restab_D62_DIFF_noRAPA.xlsx
Ignored: output/Restab_D62_noDIFF_RAPA.xlsx
Ignored: output/Restab_D62_noDIFF_noRAPA.xlsx
Ignored: output/Restab_ReN_DIFF_RAPA.xlsx
Ignored: output/Restab_ReN_DIFF_noRAPA.xlsx
Ignored: output/Restab_ReN_noDIFF_RAPA.xlsx
Ignored: output/Restab_ReN_noDIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_All_DIFF_RAPA.xlsx
Ignored: output/Restab_Repl_All_noDIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_D62D244_DIFF_RAPA.xlsx
Ignored: output/Restab_Repl_D62D244_DIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_D62D244_noDIFF_RAPA.xlsx
Ignored: output/Restab_Repl_D62D244_noDIFF_noRAPA.xlsx
Ignored: output/WGCNA_adj_TOM.RData
Untracked files:
Untracked: Wrapper.sh
Untracked: data/HumanM1 singlecellseq/
Untracked: data/genelists_to_test/
Untracked: mattson/
Untracked: wflow_helper.R
Unstaged changes:
Modified: analysis/MDSplots/rsconnect/shinyapps.io/molgenlab/mdsplots.dcf
Modified: output/dds_matrix.RData
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with wflow_publish() to start tracking its development.
home = getwd()WGCNA
soft thresholding
allowWGCNAThreads()Allowing multi-threading with up to 16 threads.dds2 = DESeq(ddsMat)using pre-existing size factorsestimating dispersionsfound already estimated dispersions, replacing thesegene-wise dispersion estimatesmean-dispersion relationshipfinal dispersion estimatesfitting model and testingvsd = getVarianceStabilizedData(dds2)
WGCNA_matrix <- t(log2(vsd+1)) #Need to transform for further calculations
s = abs(bicor(WGCNA_matrix)) #biweight mid-correlation
powers = c(c(1:10), seq(from = 12, to=20, by=2))
sft = pickSoftThreshold(WGCNA_matrix, powerVector = powers, verbose = 5)pickSoftThreshold: will use block size 3302.
pickSoftThreshold: calculating connectivity for given powers...
..working on genes 1 through 3302 of 13548
..working on genes 3303 through 6604 of 13548
..working on genes 6605 through 9906 of 13548
..working on genes 9907 through 13208 of 13548
..working on genes 13209 through 13548 of 13548
Power SFT.R.sq slope truncated.R.sq mean.k. median.k. max.k.
1 1 0.000357 -0.041 0.912 2560.000 2.52e+03 4570.0
2 2 0.468000 -1.240 0.908 784.000 7.09e+02 2220.0
3 3 0.688000 -1.560 0.946 310.000 2.43e+02 1270.0
4 4 0.772000 -1.690 0.965 144.000 9.51e+01 802.0
5 5 0.810000 -1.730 0.975 75.300 4.03e+01 538.0
6 6 0.827000 -1.750 0.979 42.600 1.83e+01 377.0
7 7 0.834000 -1.750 0.978 25.700 8.71e+00 273.0
8 8 0.849000 -1.750 0.984 16.300 4.34e+00 204.0
9 9 0.859000 -1.730 0.989 10.800 2.22e+00 155.0
10 10 0.869000 -1.710 0.989 7.350 1.16e+00 120.0
11 12 0.873000 -1.700 0.982 3.710 3.44e-01 76.5
12 14 0.876000 -1.700 0.990 2.040 1.10e-01 52.0
13 16 0.874000 -1.700 0.987 1.200 3.75e-02 36.8
14 18 0.878000 -1.690 0.992 0.741 1.32e-02 26.9
15 20 0.854000 -1.710 0.983 0.479 4.77e-03 20.1par(mfrow = c(1,2))
cex1 = 0.9;
plot(sft$fitIndices[,1], -sign(sft$fitIndices[,3])*sft$fitIndices[,2],
xlab="Soft Threshold (power)",ylab="Scale Free Topology Model Fit, signed R^2",
type="n", main = paste("Scale independence"));
text(sft$fitIndices[,1], -sign(sft$fitIndices[,3])*sft$fitIndices[,2],
labels=powers,cex=cex1,col="red");
abline(h=0.80,col="red")
plot(sft$fitIndices[,1], sft$fitIndices[,5],xlab="Soft Threshold (power)",ylab="Mean Connectivity", type="n",main = paste("Mean connectivity"))
text(sft$fitIndices[,1], sft$fitIndices[,5], labels=powers, cex=cex1,col="red")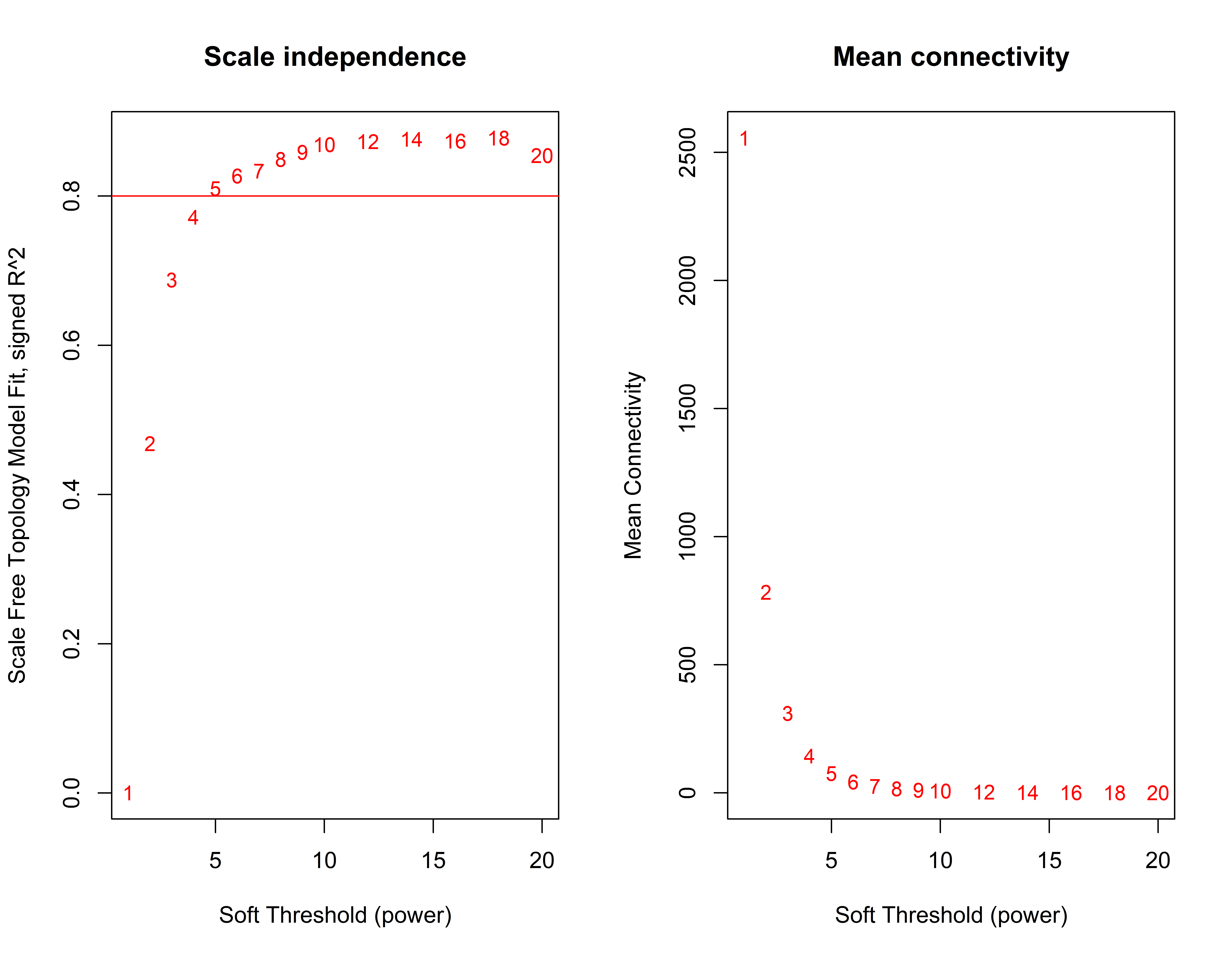
Identify Gene Modules
softPower = 6; # The point where the curve flattens
#calclute the adjacency matrix
if(reanalyze | !file.exists(paste0(output,"/WGCNA_adj_TOM.RData"))){
adj= adjacency(WGCNA_matrix,type = "unsigned", power = softPower);
#Converting adjacency matrix into To so that the noise could be reduced
TOM=TOMsimilarityFromExpr(WGCNA_matrix,networkType = "unsigned",
TOMType = "unsigned", power = softPower);
save(list = c("adj", "TOM"), file=paste0(output,"/WGCNA_adj_TOM.RData"))
} else {
load(paste0(output,"/WGCNA_adj_TOM.RData"))
}
SubGeneNames<-colnames(WGCNA_matrix)
colnames(TOM) =rownames(TOM) =SubGeneNames
dissTOM=1-TOM
diag(dissTOM) = 0
#hierarchical clustering
geneTree = flashClust(as.dist(dissTOM),method="average");
#plot the resulting clustering tree (dendrogram)
# plot(geneTree, xlab="", sub="",cex=0.3, main="Module clustering prio to merging");
#Set the minimum module size
minModuleSize = 50;
#Module identification using dynamic tree cut
dynamicMods = cutreeDynamic(dendro = geneTree,
distM = dissTOM,
cutHeight = 0.998,
minClusterSize = minModuleSize,
deepSplit=2,
pamRespectsDendro = T) ..done.#the following command gives the module labels and the size of each module. Lable 0 is reserved for unassigned genes
#table(dynamicMods)
#Plot the module assignment under the dendrogram; note: The grey color is reserved for unassigned genes
dynamicColors = labels2colors(dynamicMods)
table(dynamicColors)dynamicColors
black blue brown cyan darkgreen
451 946 640 237 164
darkgrey darkmagenta darkolivegreen darkorange darkred
159 106 127 154 173
darkturquoise green greenyellow grey grey60
164 501 321 1390 211
ivory lightcyan lightcyan1 lightgreen lightsteelblue1
62 229 66 209 68
lightyellow magenta mediumpurple3 midnightblue orange
207 361 73 235 158
orangered4 paleturquoise pink plum1 purple
73 133 412 74 357
red royalblue saddlebrown salmon sienna3
465 204 142 282 95
skyblue skyblue3 steelblue tan turquoise
151 80 140 304 2233
violet white yellow yellowgreen
127 153 621 90 plotDendroAndColors(geneTree, dynamicColors, "Dynamic Tree Cut",
dendroLabels = FALSE, hang = 0.03,
addGuide = TRUE,
guideHang = 0.05,
main = "Gene dendrogram and module colors")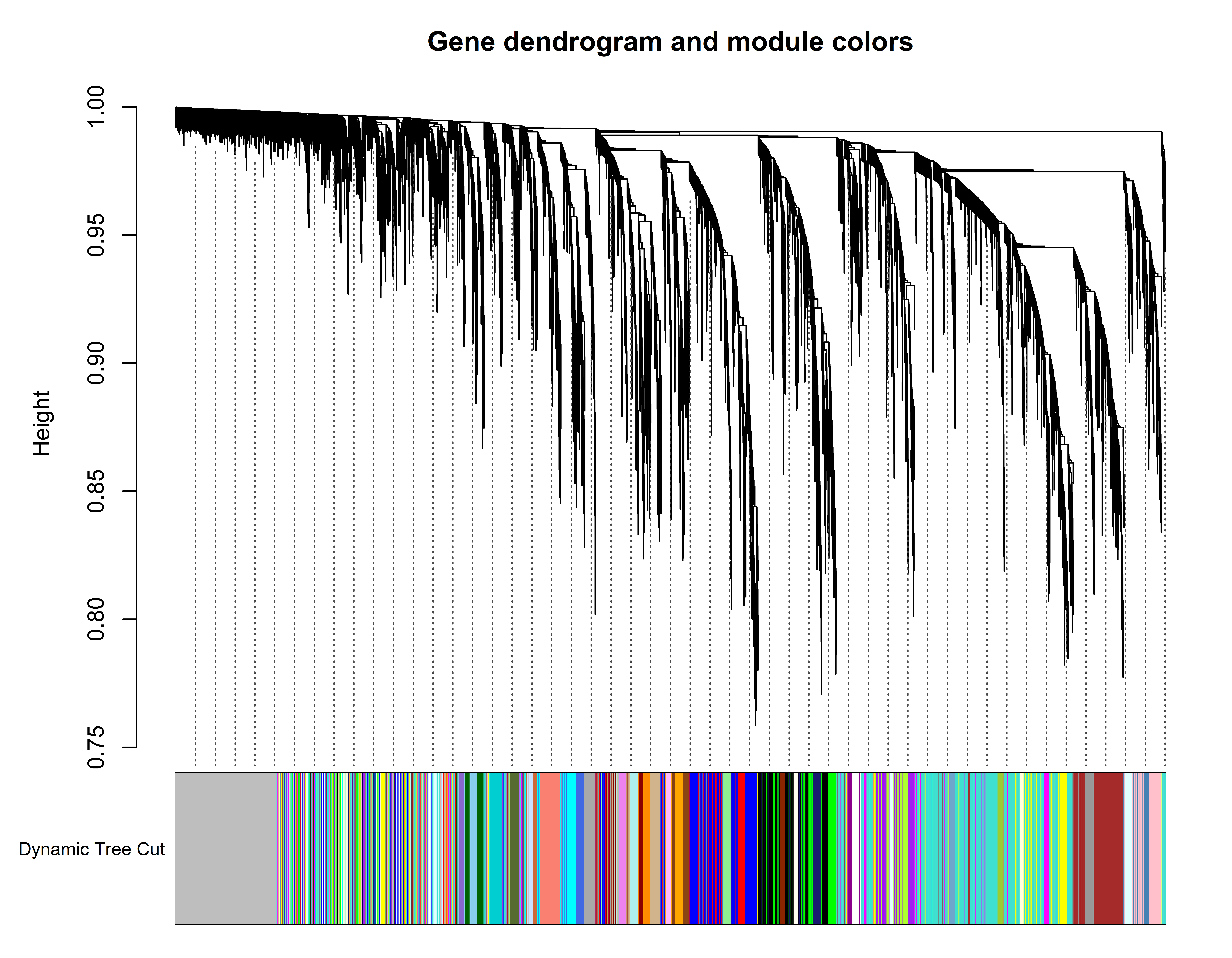
diag(dissTOM) = NA;#Visualize the Tom plot. Raise the dissimilarity matrix to the power of 4 to bring out the module structure
TOMplot(dissTOM^4, geneTree, as.character(dynamicColors), main="weighted distance of Topological overlap Matrix")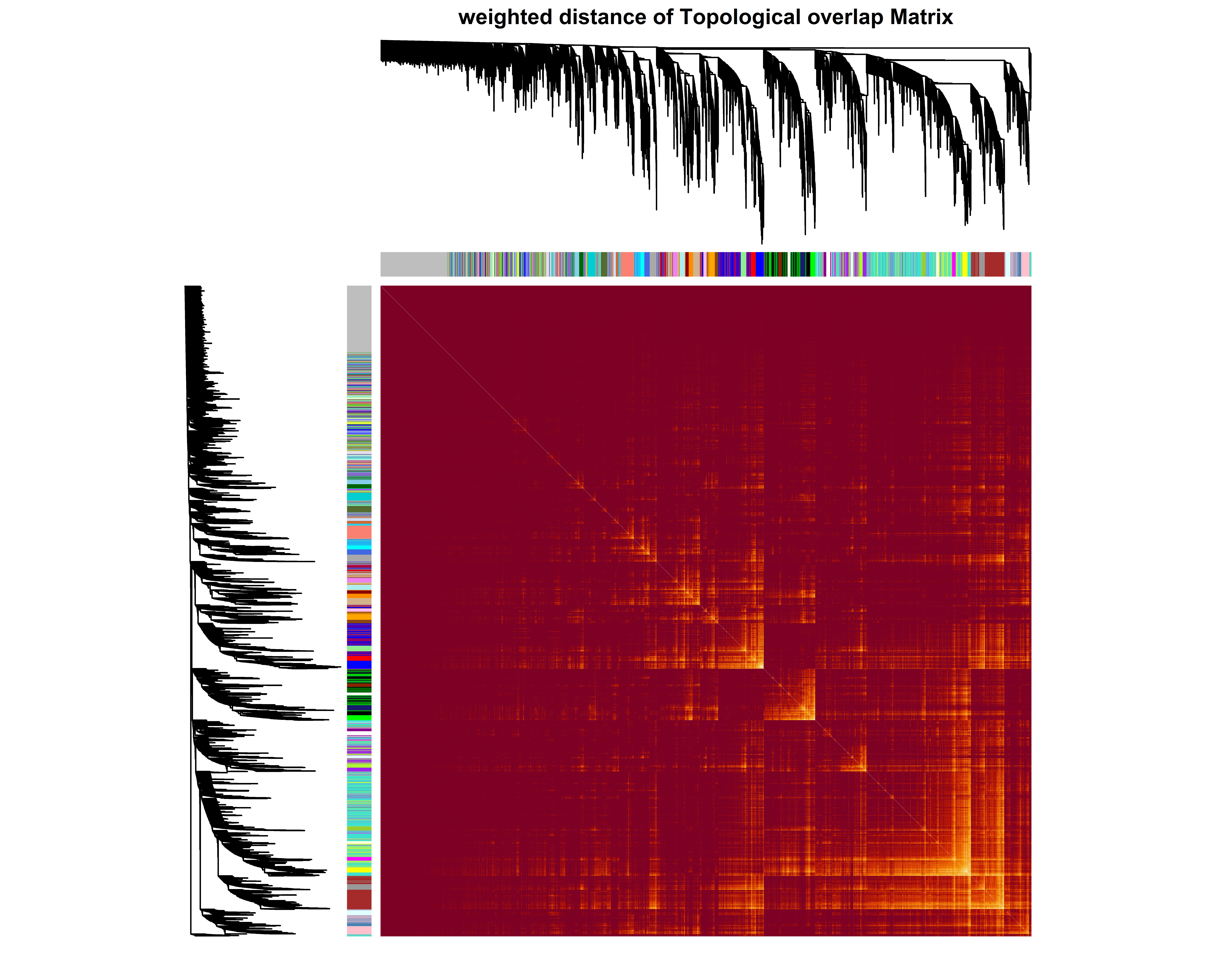
Module eigengenes
Heatmaps MEs
#colors for plotting heatmap
colors <- rev(colorRampPalette(brewer.pal(9, "Spectral"))(255))
cellcol = Dark8[1:nlevels(SampleInfo$CellLine)]
names(cellcol) = levels(SampleInfo$CellLine)
gRNAcol = Dark8[c(1:nlevels(SampleInfo$gRNA))+nlevels(SampleInfo$CellLine)]
names(gRNAcol) = levels(SampleInfo$gRNA)
diffcol = brewer.pal(3,"Set1")[1:nlevels(SampleInfo$DIFF)]
names(diffcol) = levels(SampleInfo$DIFF)
rapacol = brewer.pal(3,"Set2")[1:nlevels(SampleInfo$RAPA)]
names(rapacol) = levels(SampleInfo$RAPA)
clustcol = gplots::col2hex(unique(as.character(colors_new)))
names(clustcol) = unique(as.character(colors_new))
rownames(WGCNA_matrix)=SampleInfo[rownames(WGCNA_matrix), "label_rep"]
ann_colors = list(
DIFF = diffcol,
RAPA = rapacol,
gRNA = gRNAcol,
CellLine=cellcol,
cluster = clustcol)
idx=order(SampleInfo$gRNA, SampleInfo$CellLine, SampleInfo$DIFF,SampleInfo$RAPA)
WGCNA_matrix_sorted=WGCNA_matrix[SampleInfo$label_rep[idx], order(colors_new)]
collabels = SampleInfo[idx,c("CellLine","gRNA","DIFF", "RAPA")] %>%
mutate_all(as.character) %>% as.data.frame()
rownames(collabels)=SampleInfo$label_rep[idx]
genlabels = data.frame(cluster = as.character(colors_new)[order(colors_new)])
rownames(genlabels) = colnames(WGCNA_matrix_sorted)
# pheatmap(t(WGCNA_matrix_sorted),
# cluster_rows= F,
# cluster_cols = F,
# border_color = NA, show_rownames = F, show_colnames = F,
# clustering_method = "ward.D2",
# annotation_row = genlabels,
# annotation_col = collabels,
# annotation_colors = ann_colors,
# col = colors,
# scale = "row",
# main = "Distances normalized log2 counts")
MElabels = data.frame(cluster = gsub("ME", "",colnames(MEs)))
rownames(MElabels) = colnames(MEs)
clustcol = gplots::col2hex(unique(as.character(MElabels$cluster)))
names(clustcol) = as.character(MElabels$cluster)
ann_colors = list(
DIFF = diffcol,
RAPA = rapacol,
gRNA = gRNAcol,
CellLine=cellcol,
cluster = clustcol)
rownames(MEs) = SampleInfo[rownames(MEs),"label_rep"]
pheatmap(t(MEs[idx,]),
border_color = NA,
annotation_row = MElabels,
annotation_col = collabels,
cluster_cols = F,
show_rownames = F, show_colnames = F,
clustering_method = "ward.D2",
annotation_colors = ann_colors,
scale="row",
breaks = seq(-2, 2,length.out=255),
col = colors,
main = "eigengene values")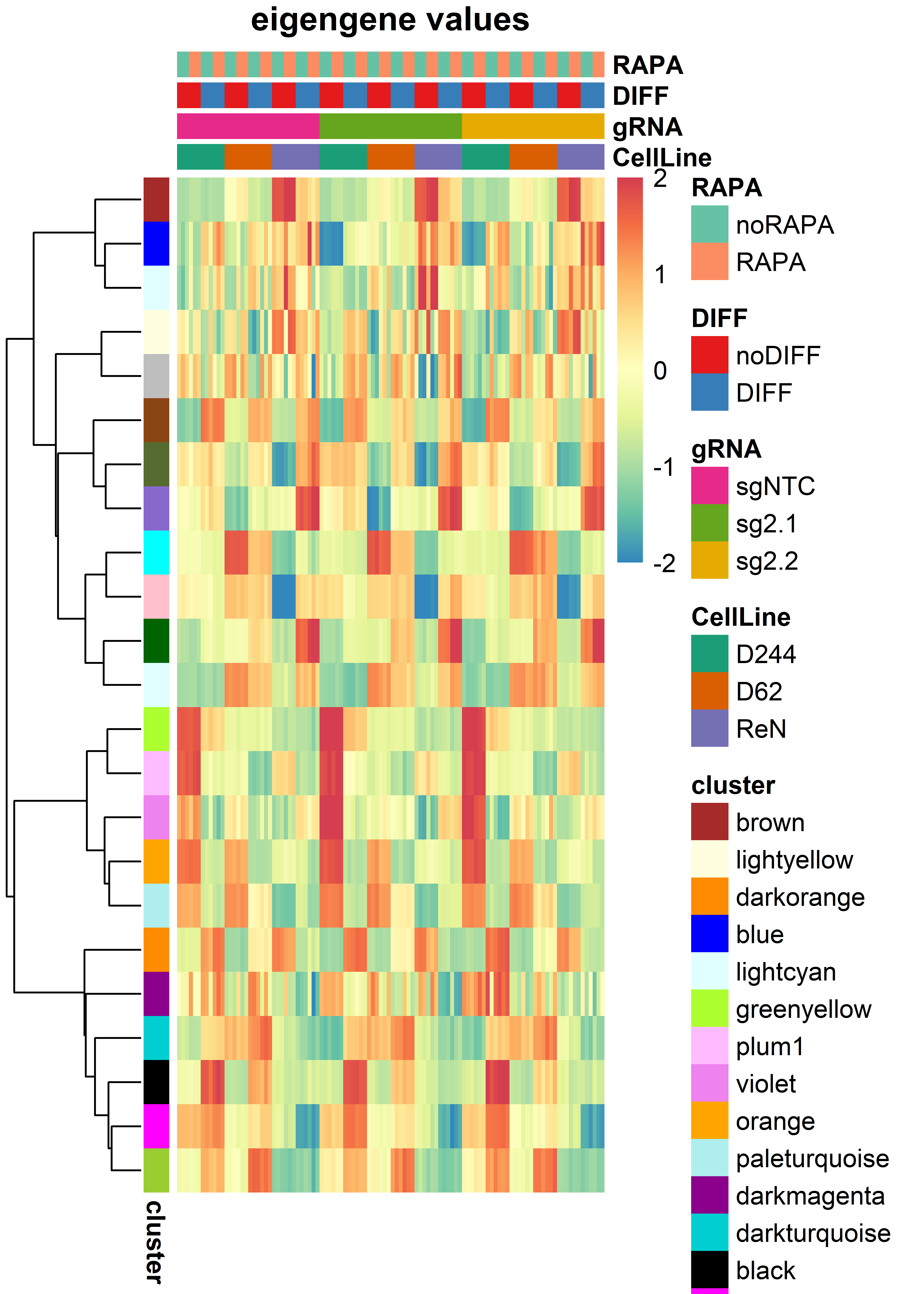
SampleInfo = as.data.frame(colData(ddsMat))
MEMat = SampleInfo[,grep("ME", colnames(SampleInfo))]
## herlper functions#test differences
testit=function(Dataset, samples = Set,
depvar){
data = Dataset[samples,]
res=list()
for(i in grep("ME", colnames(data), value = T)){
res[[i]]=lm(as.formula(paste0(i,"~1+",depvar)), data)
}
return(res)
}
# extract coefficents
getcoefff=function(x){
res = summary(x)$coefficients[2,]
return(res)
}
# comparisonME
comparisonME = function(SampleInfo, Set, target){
LMlist=testit(Dataset = SampleInfo, samples = Set, depvar=target)
coeff = as.data.frame(lapply(LMlist, getcoefff) %>% do.call(rbind, .))
coeff$padj = p.adjust(coeff$`Pr(>|t|)`, "bonferroni")
return(coeff)}
CellLines=c("D62","D244","ReN")
Rapamycin=c("noRAPA", "RAPA")
Differentiation=c("noDIFF", "DIFF")
Type_sgRNA<-list(c("sgNTC","sg2.1"),c("sgNTC","sg2.2"))
target="KO"
r = Rapamycin[1]
Cl=CellLines[1]
d = Differentiation[1]
Tp = Type_sgRNA[[1]]
# no random effects included
for(r in Rapamycin){
Rapafilter = SampleInfo$RAPA %in% r
for(d in Differentiation){
Difffilter = SampleInfo$DIFF %in% d
for(Tp in Type_sgRNA){
sgRNAfilter = SampleInfo$gRNA %in% Tp
vs_label=paste0(Tp, sep="", collapse="_")
for(Cl in CellLines){
Celllinefilter = SampleInfo$CellLine %in% Cl
Set = rownames(SampleInfo)[Rapafilter&Difffilter&
sgRNAfilter&Celllinefilter]
lab = paste("restabWGCNA", Cl, vs_label, d,r, sep="_")
assign(lab, comparisonME(SampleInfo, Set, target))
}
}
}
}
comparisons= apropos("restabWGCNA")
save(list = comparisons, file = paste0(home,"/output/ResTabs_KO_WGCNA.RData"))mypval=0.05
MEIds = rownames(restabWGCNA_D62_sgNTC_sg2.1_noDIFF_noRAPA)
getWGCNAoutputs = function(targetline,targetdiff,targetrapa){
samplesincl = SampleInfo$DIFF==targetdiff &
SampleInfo$RAPA==targetrapa &
SampleInfo$CellLine==targetline
pvalrep= get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj<=mypval &
get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj<=mypval
betarep = apply(cbind(get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$Estimate,
get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$Estimate), 1,
samesign)
idx=which(betarep & pvalrep)
hits=MEIds[idx]
restab=data.frame(
Module = MEIds,
beta_2.1 = get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$Estimate,
bonferroni_2.1 = get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj,
beta_2.2 = get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$Estimate,
bonferroni_2.2 = get(paste0("restabWGCNA_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj)
print(restab)
write.xlsx(restab, file=paste0(output, "/Restab_",
targetline,"_",
targetdiff, "_",
targetrapa, ".xlsx"))
SamplesSet=SampleInfo[samplesincl,] %>% select(all_of(hits))
EigengenePlot(SamplesSet, SampleInfo, samplesincl)
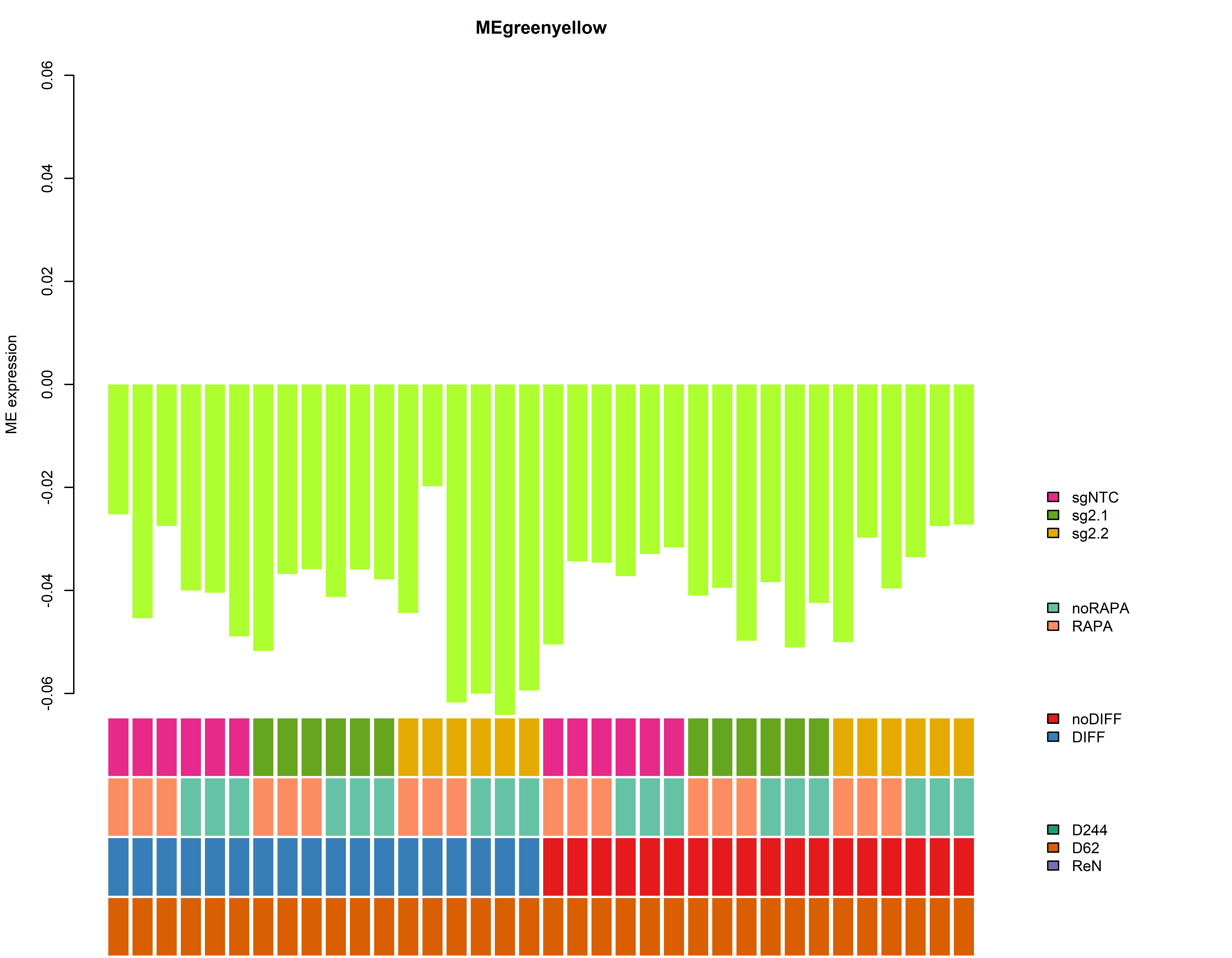
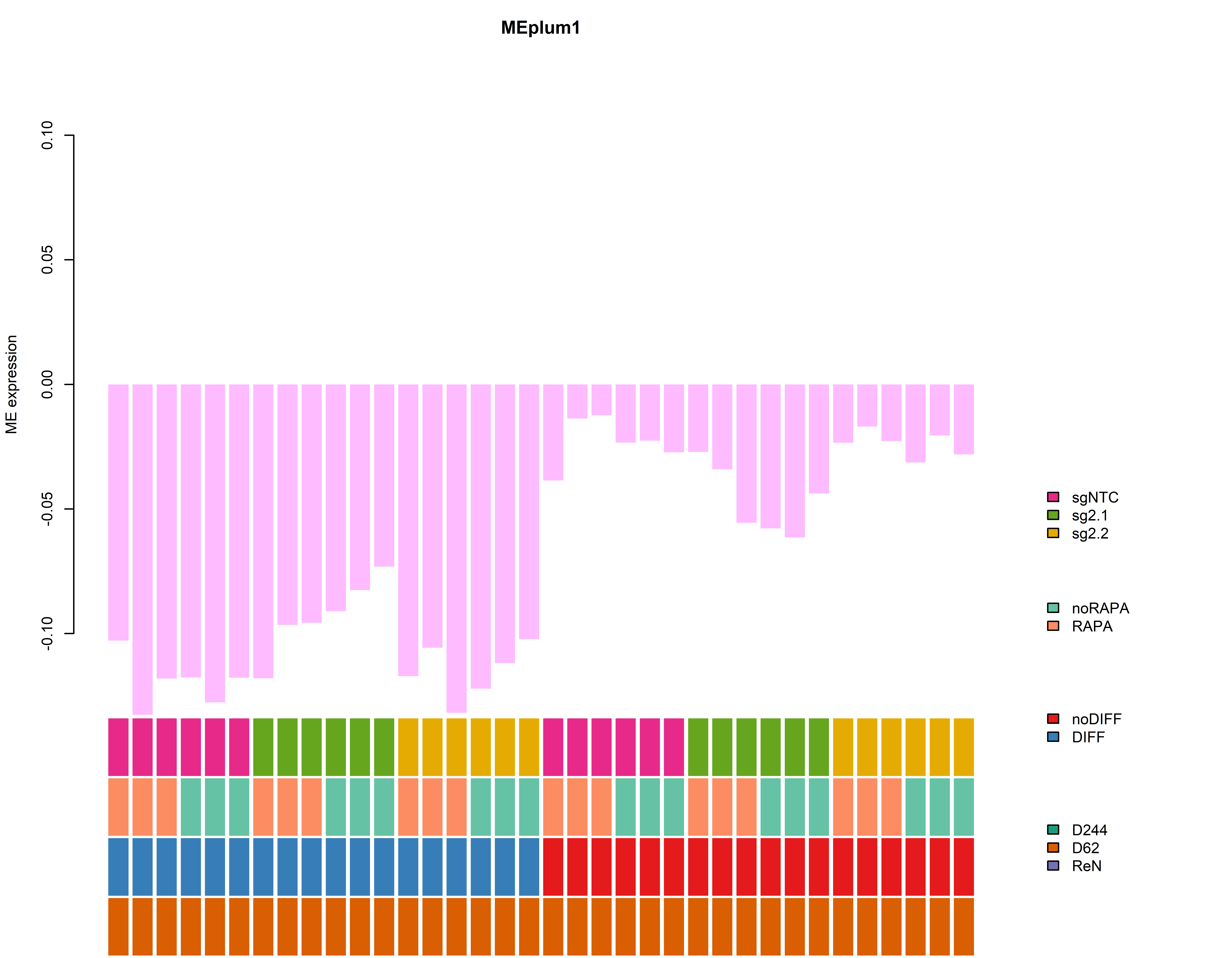
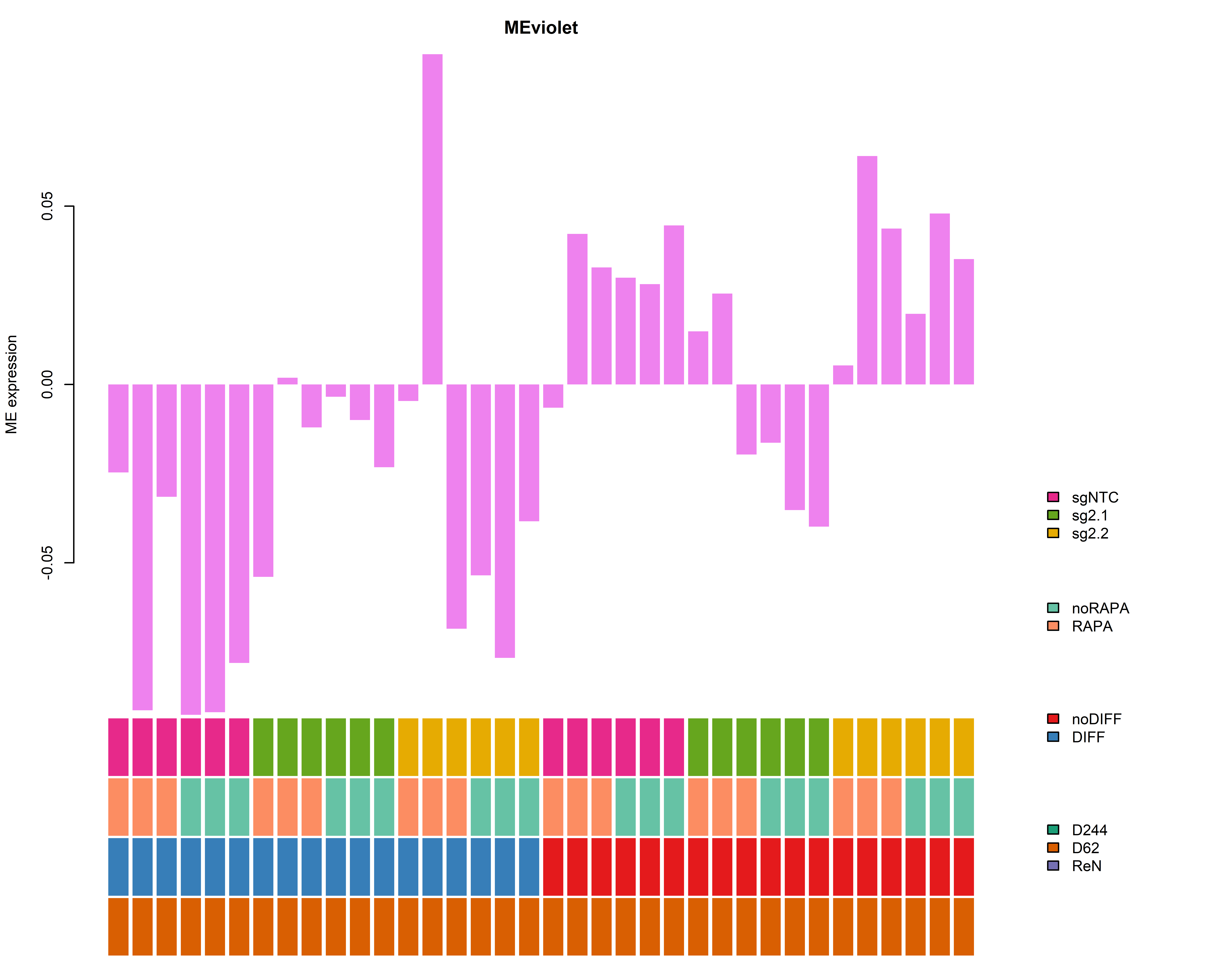
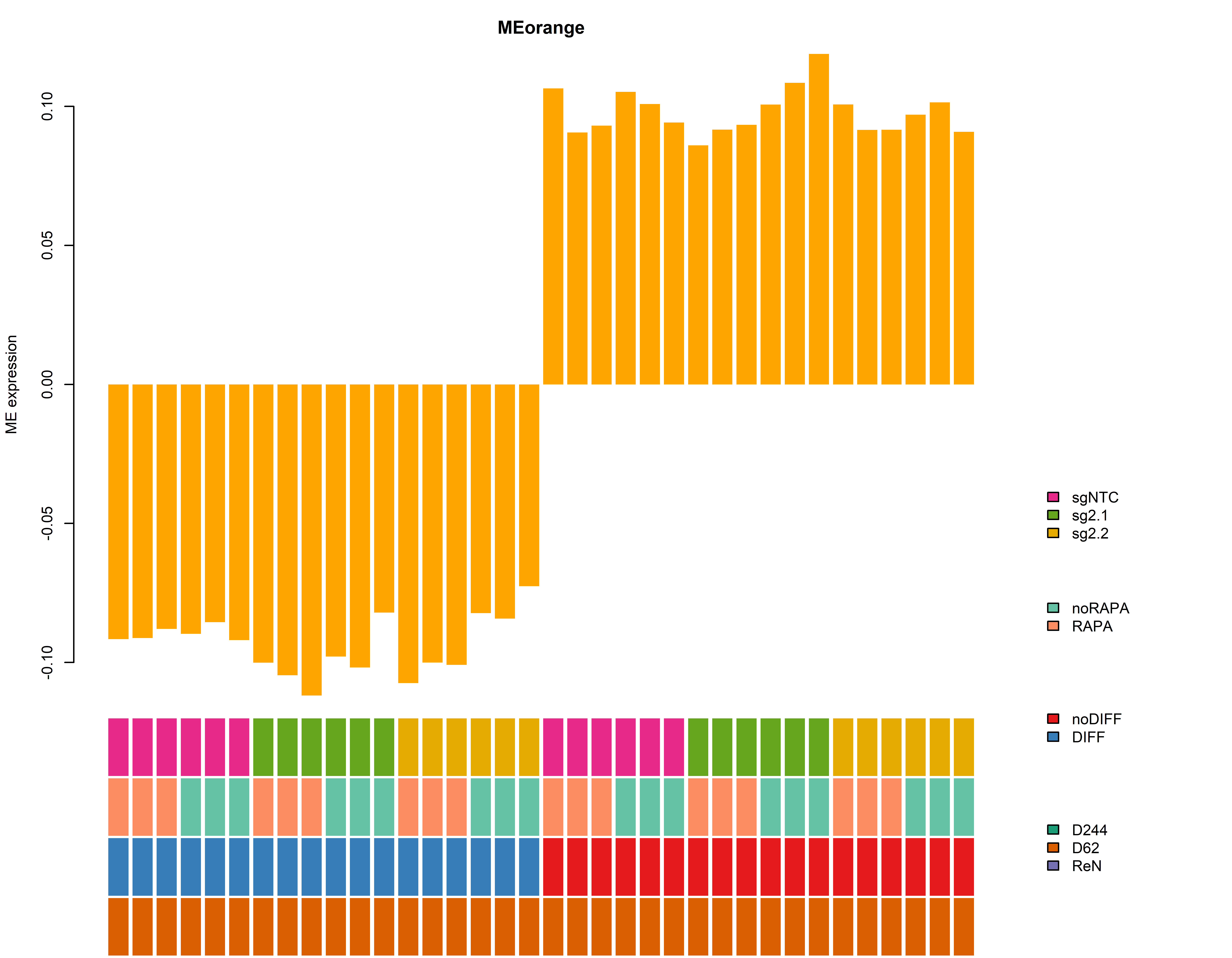
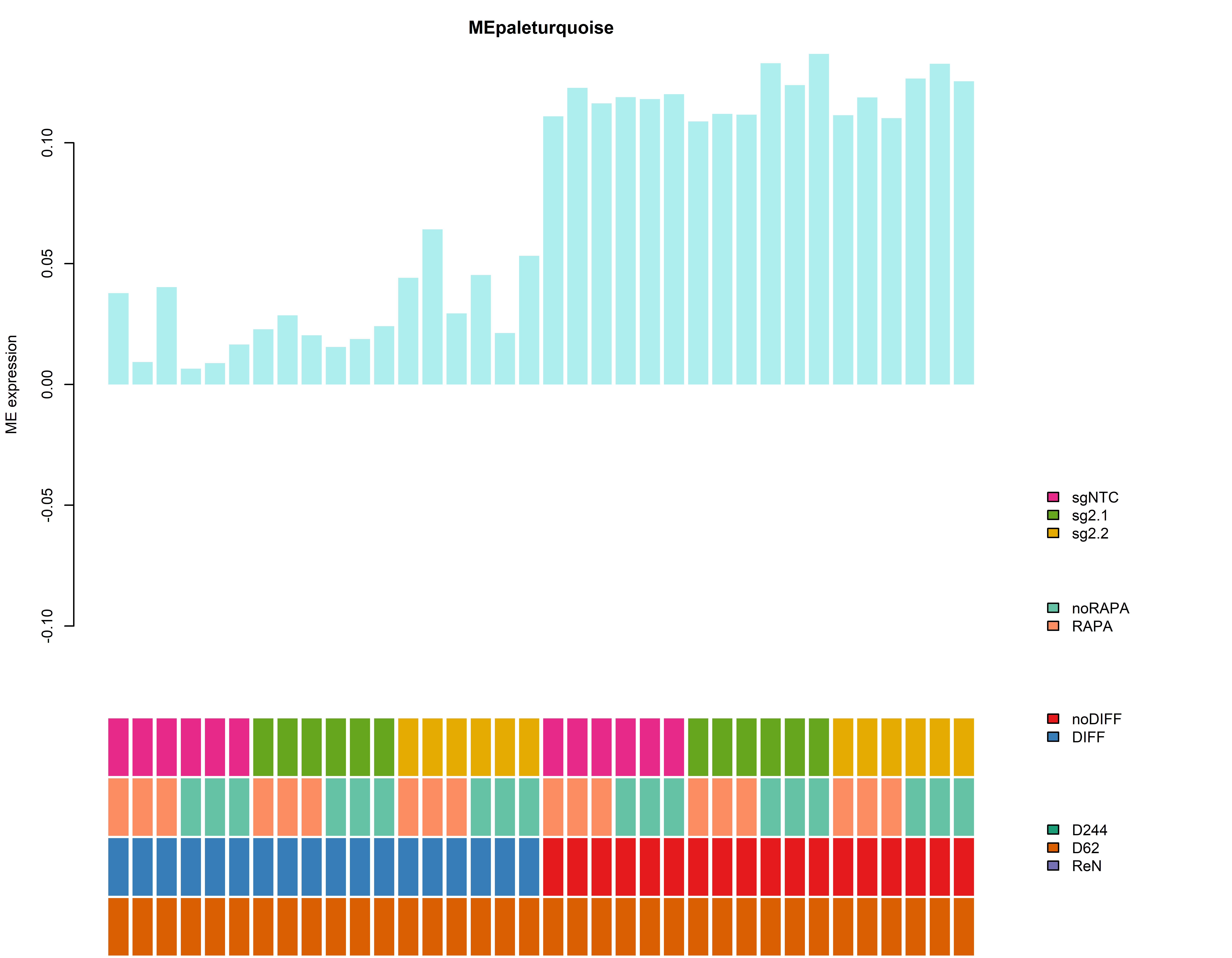
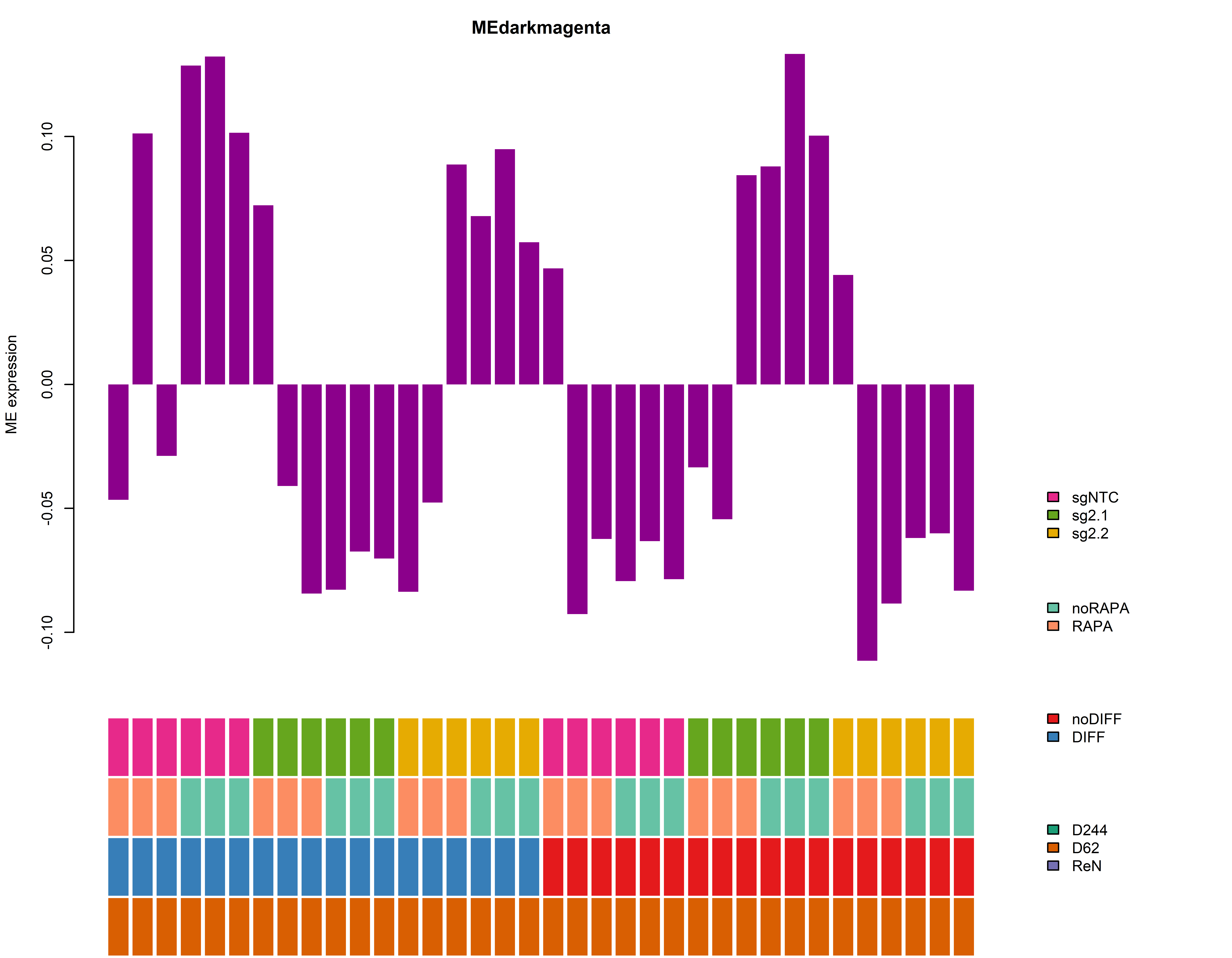
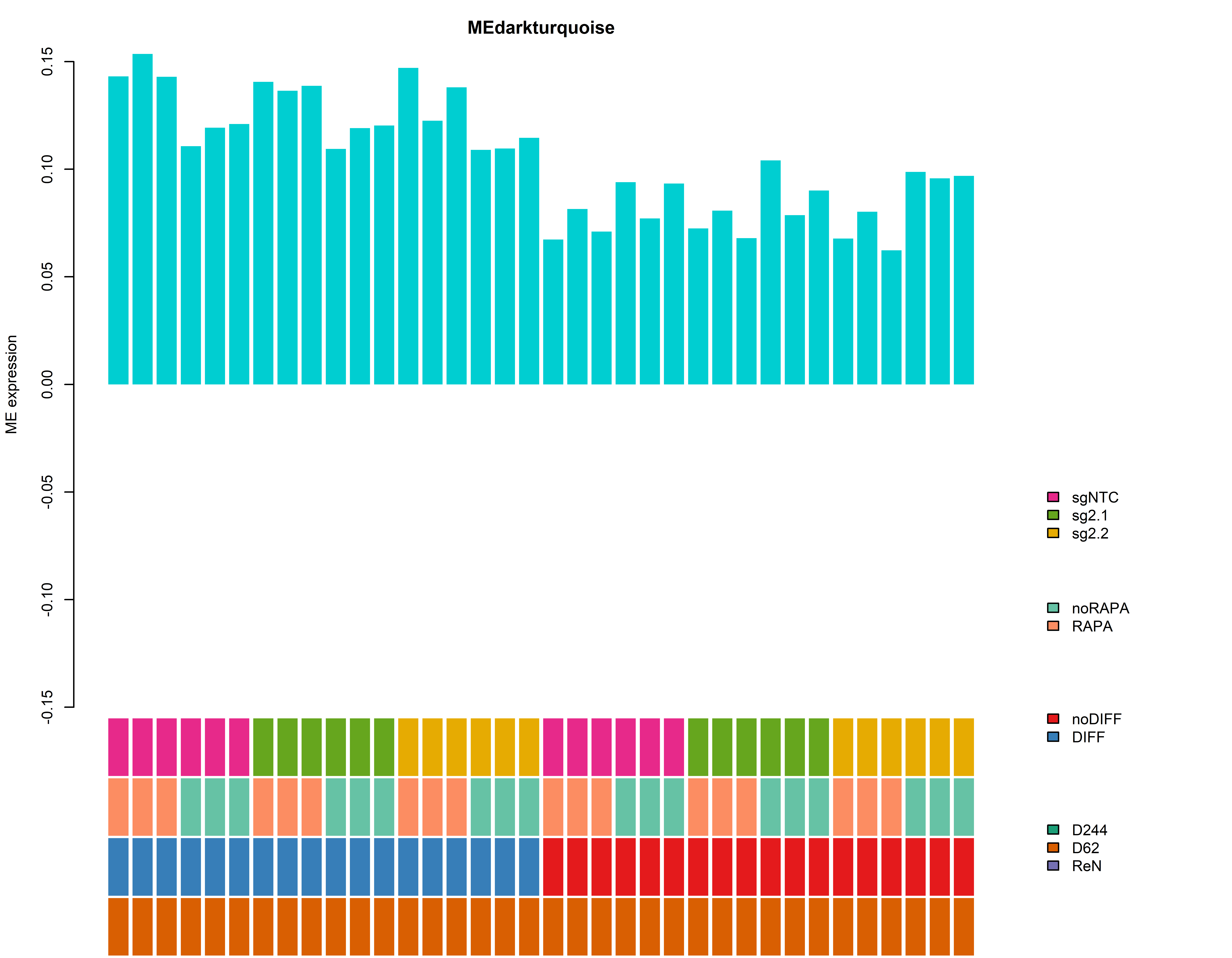
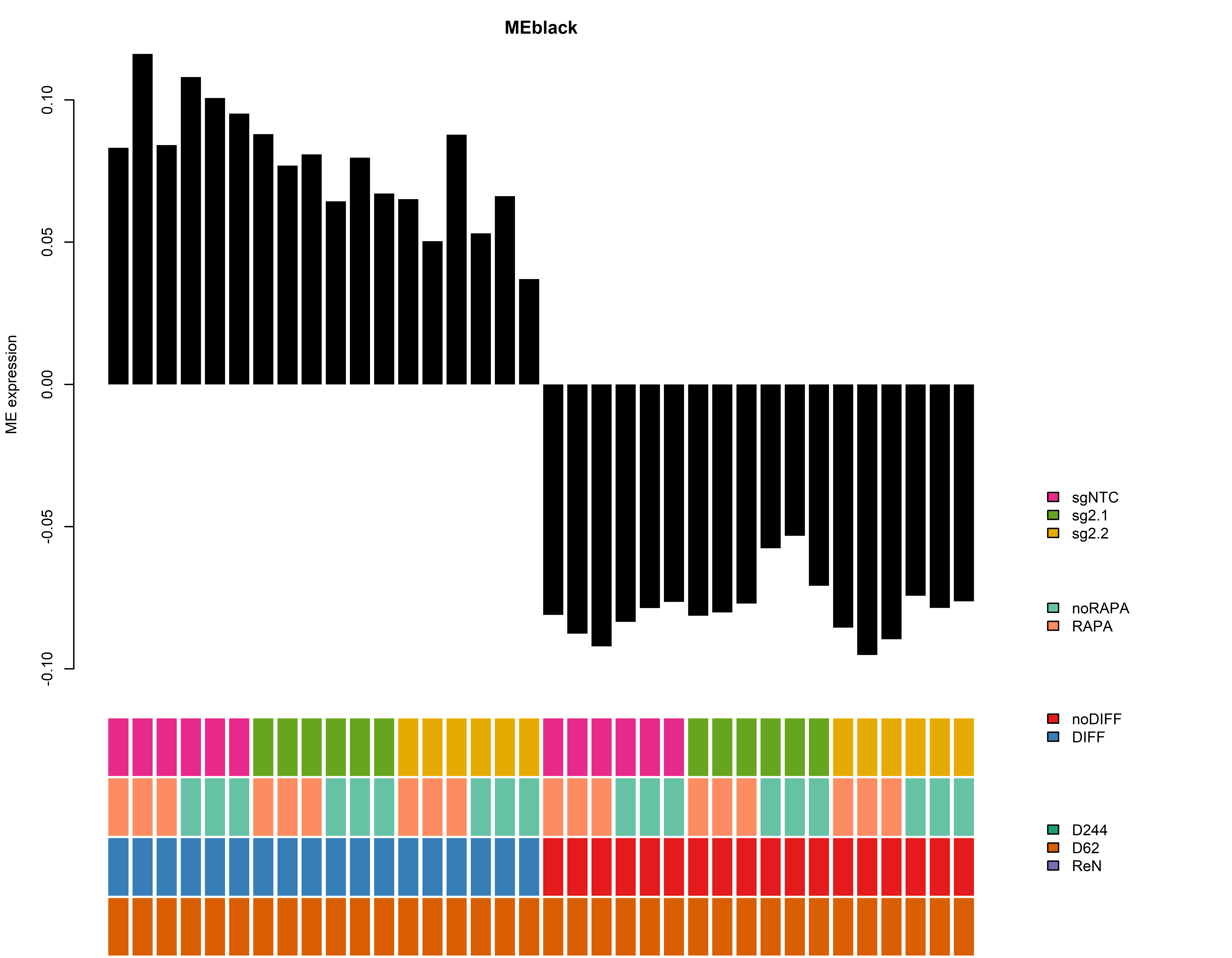
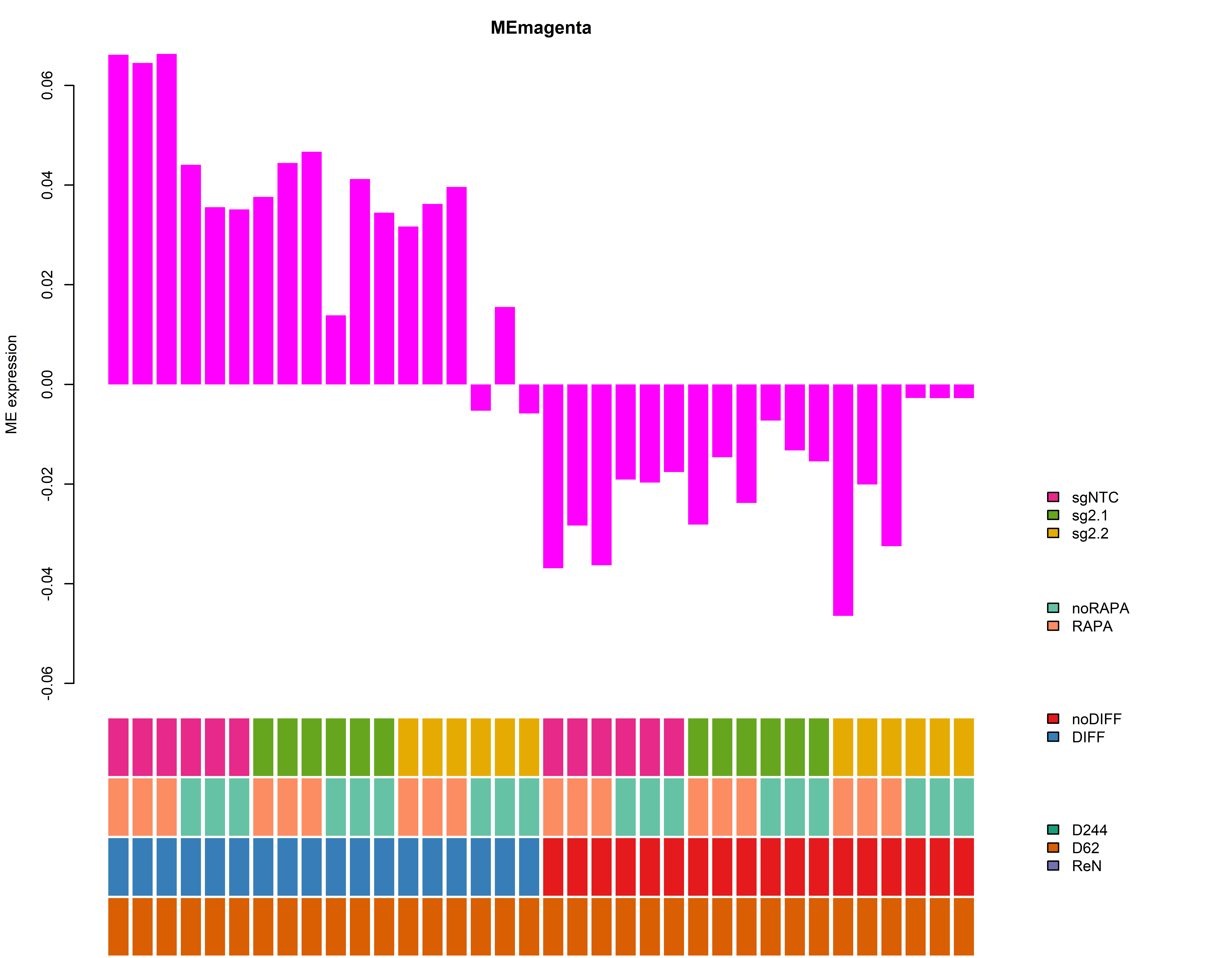
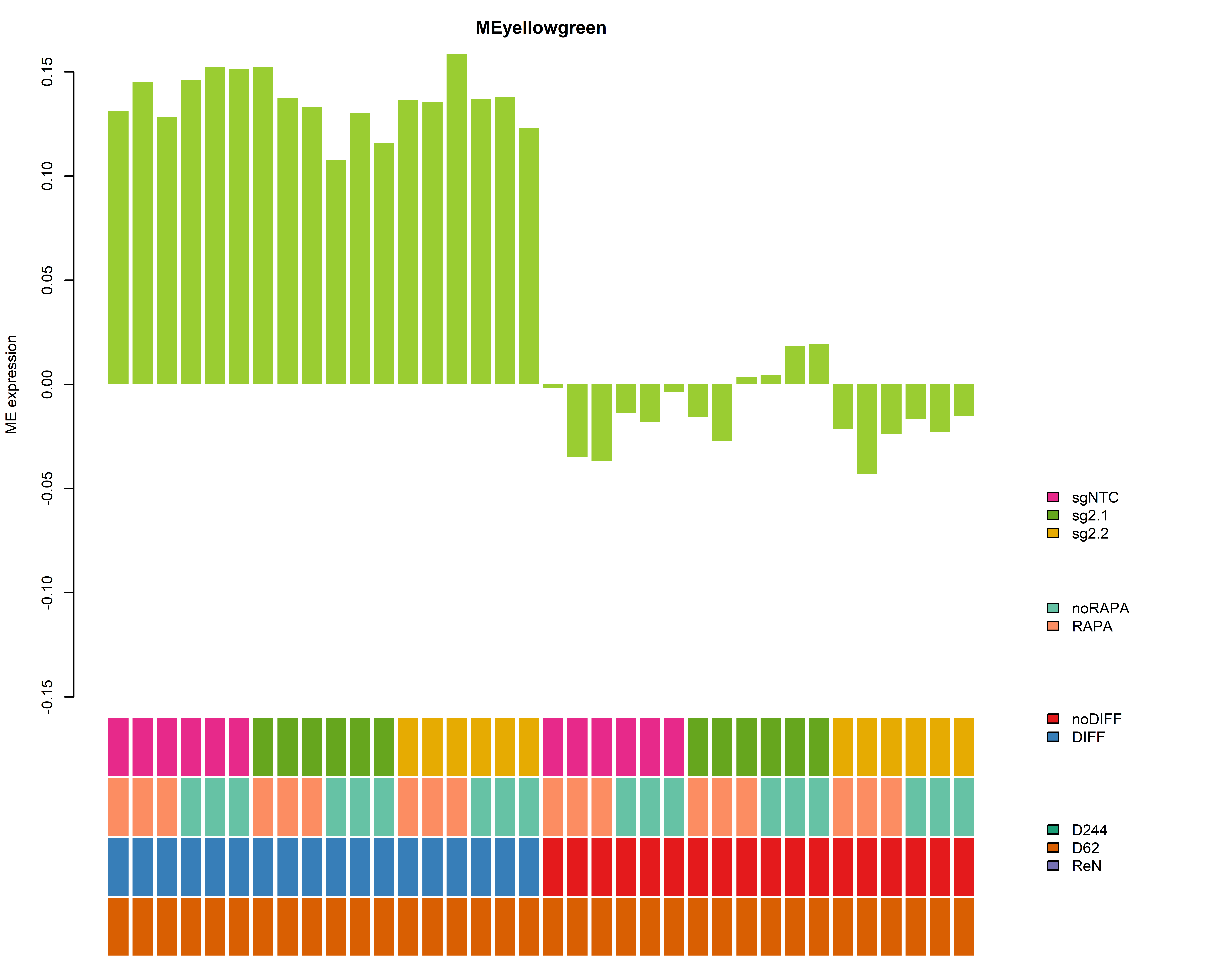
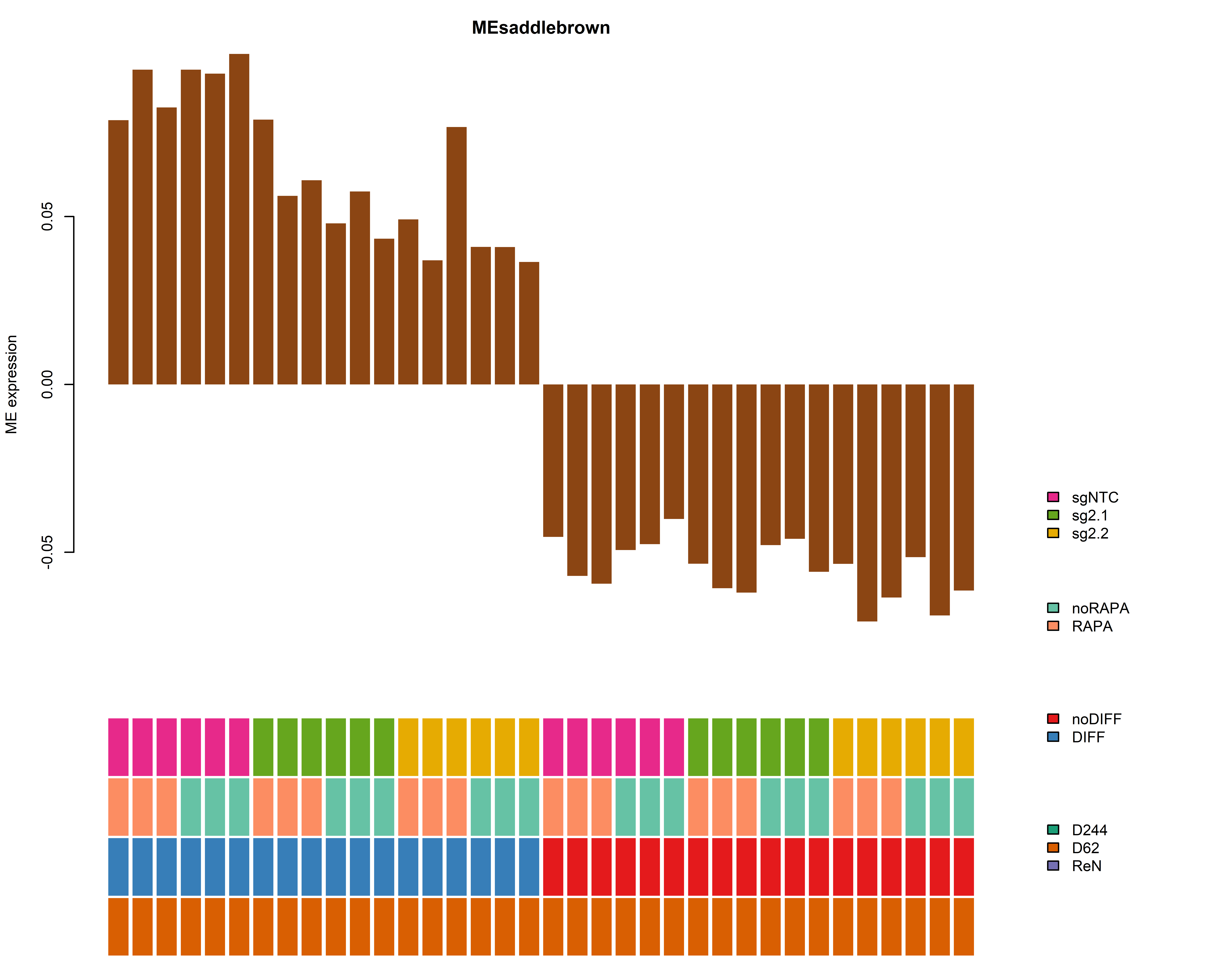
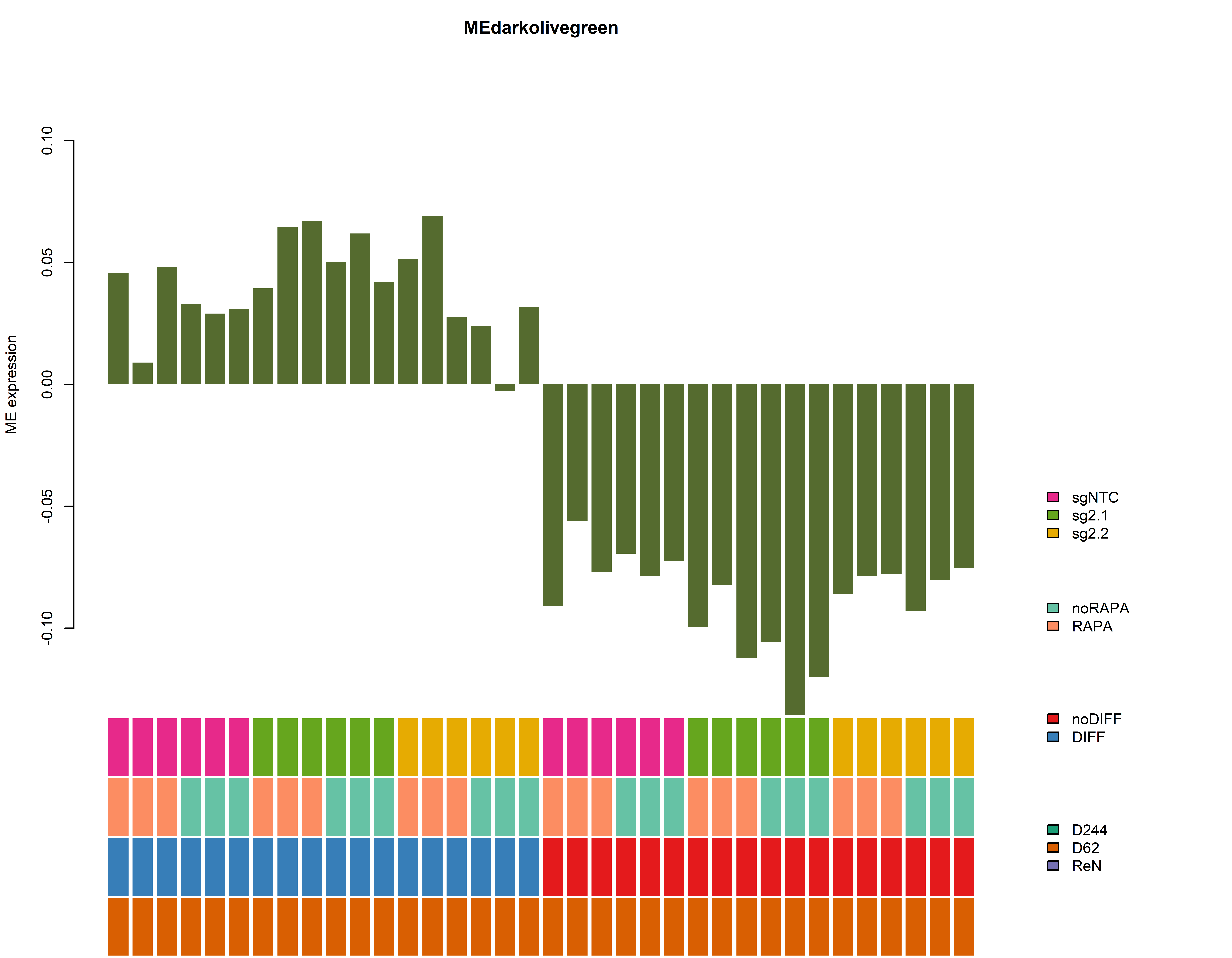
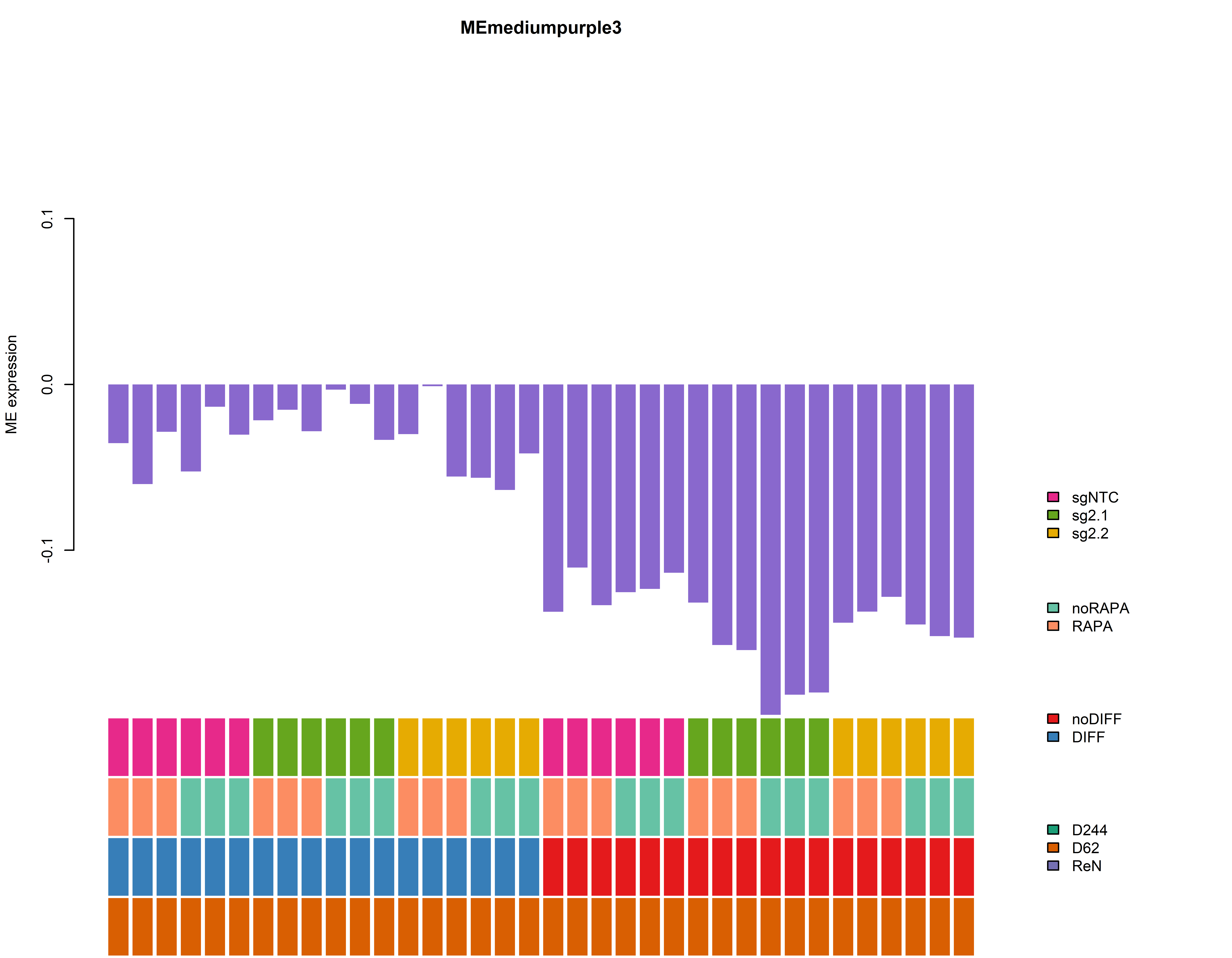
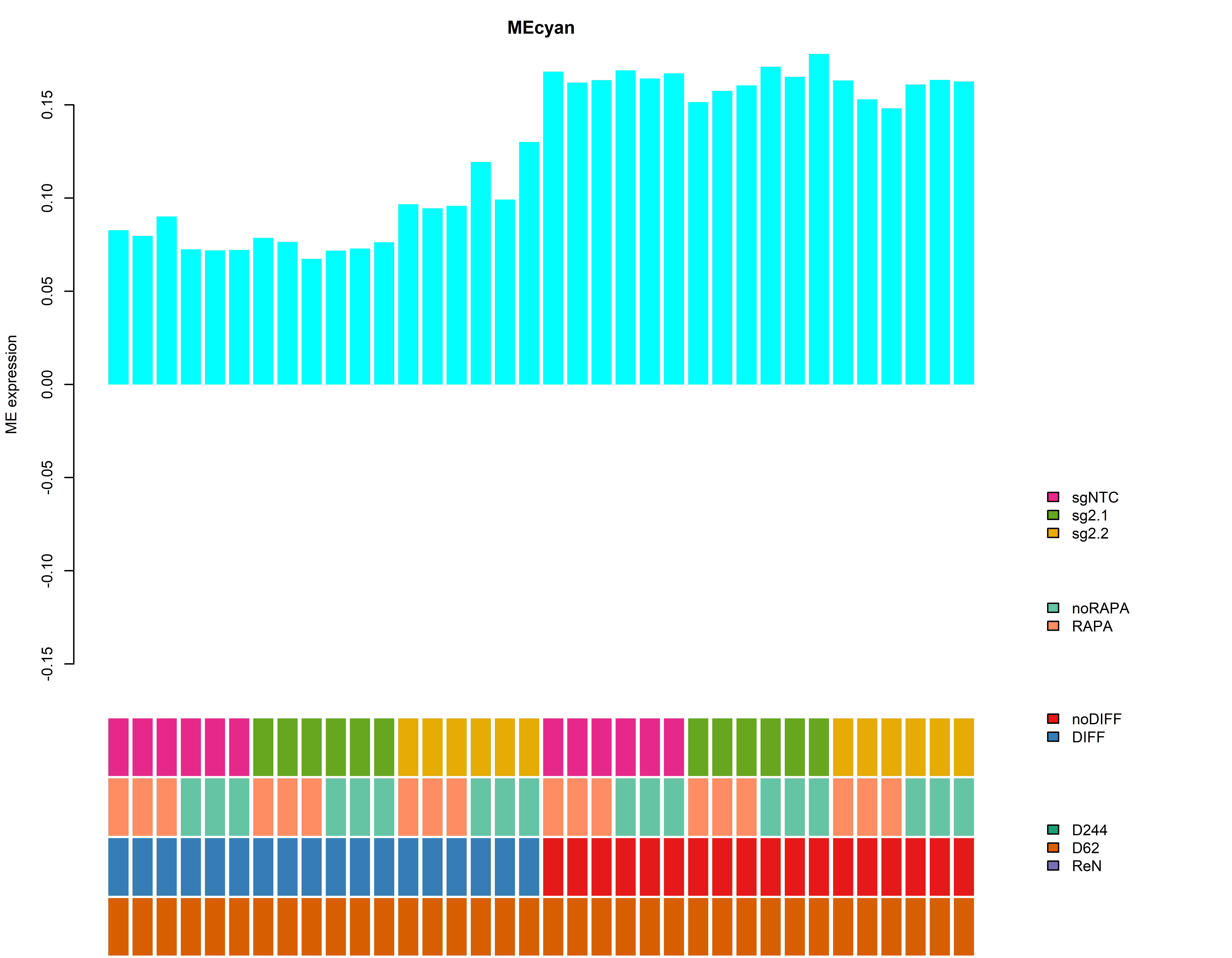
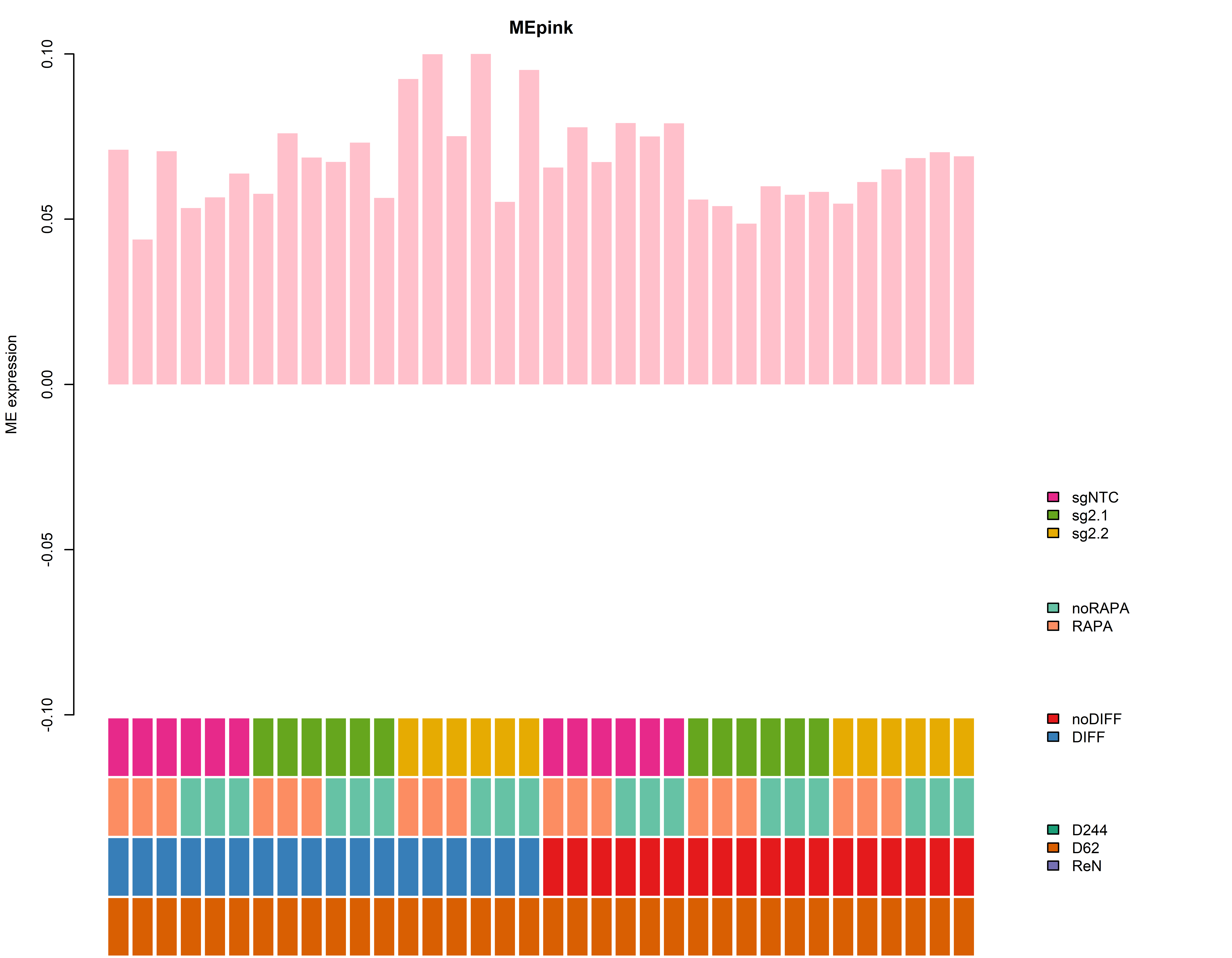
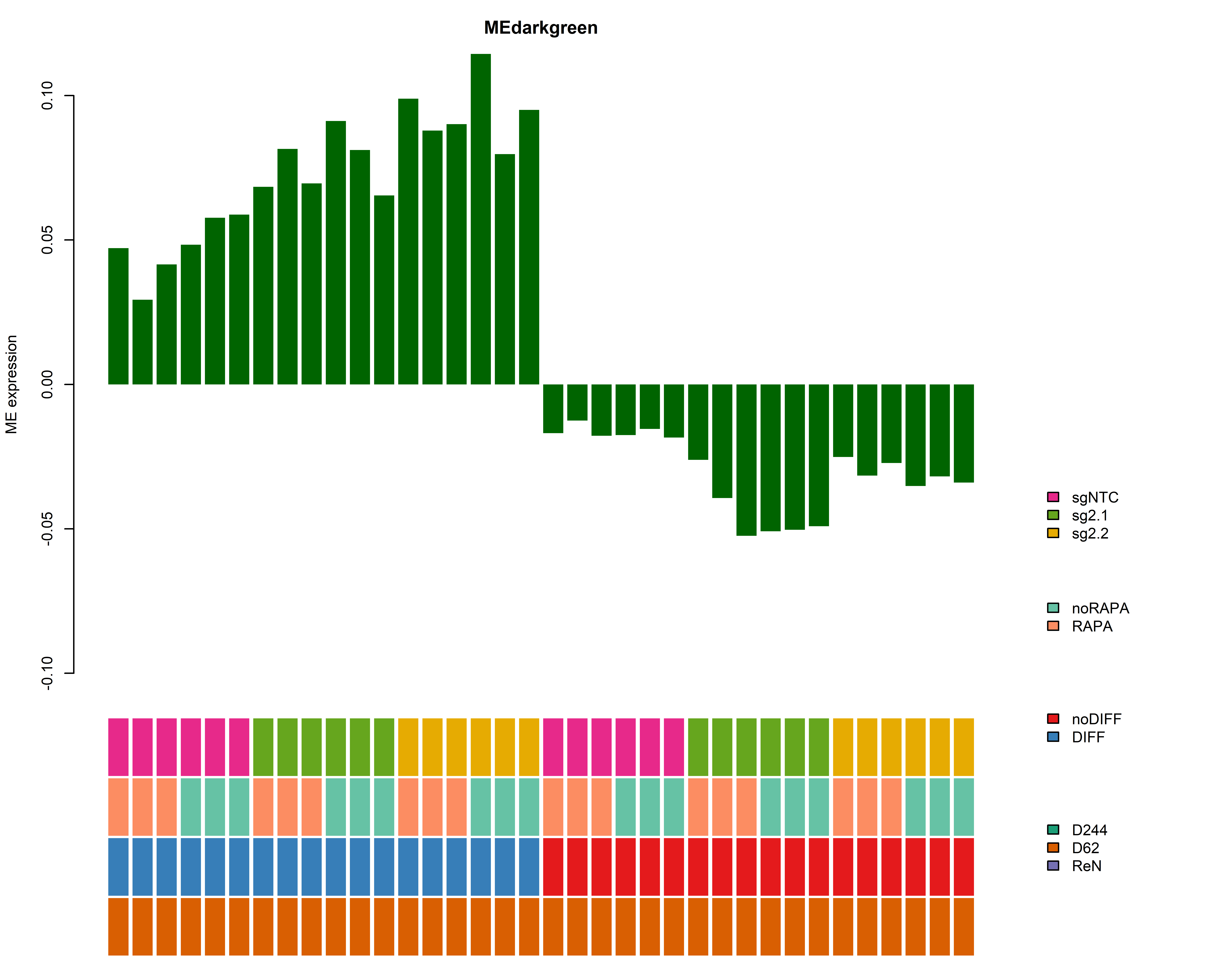
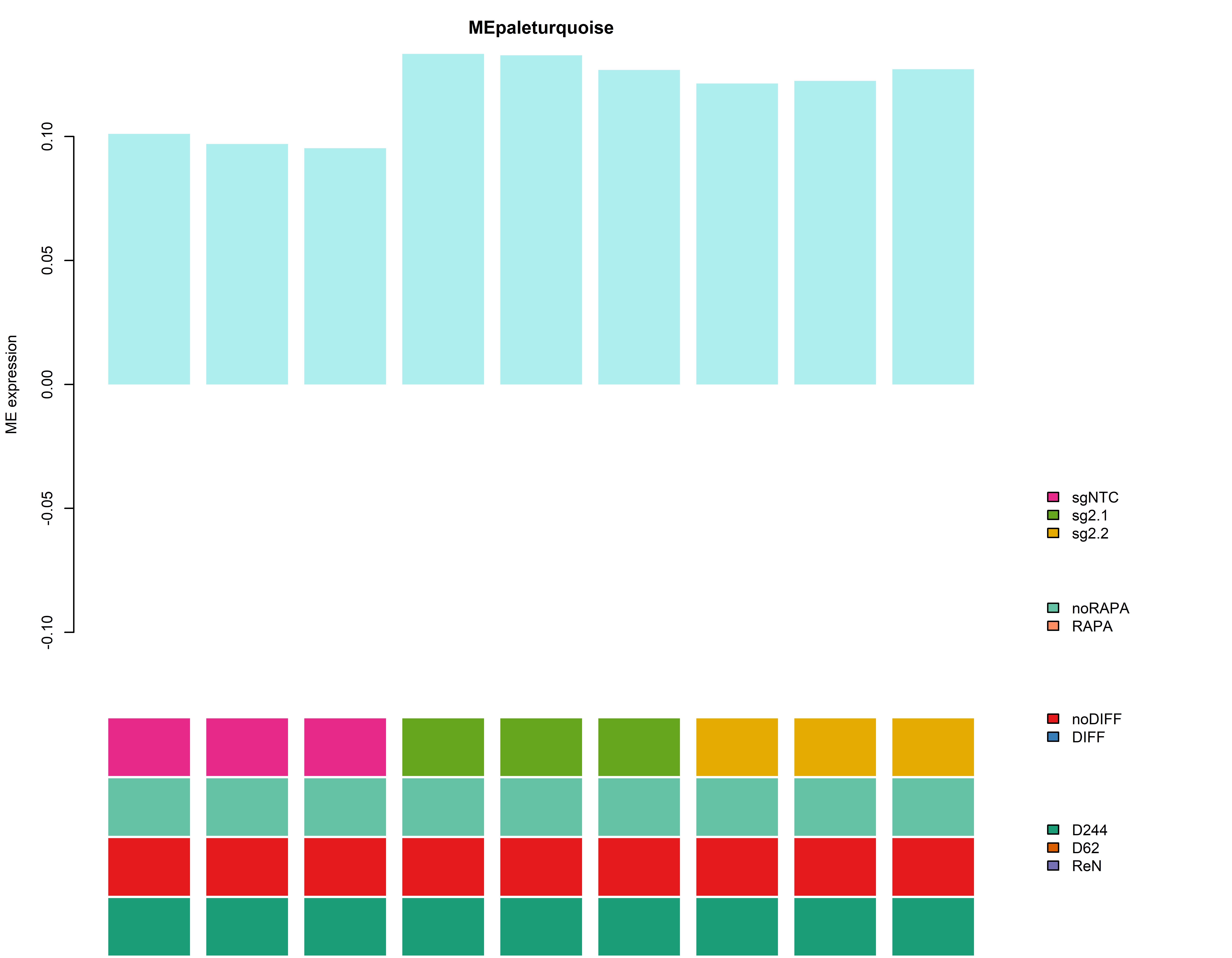
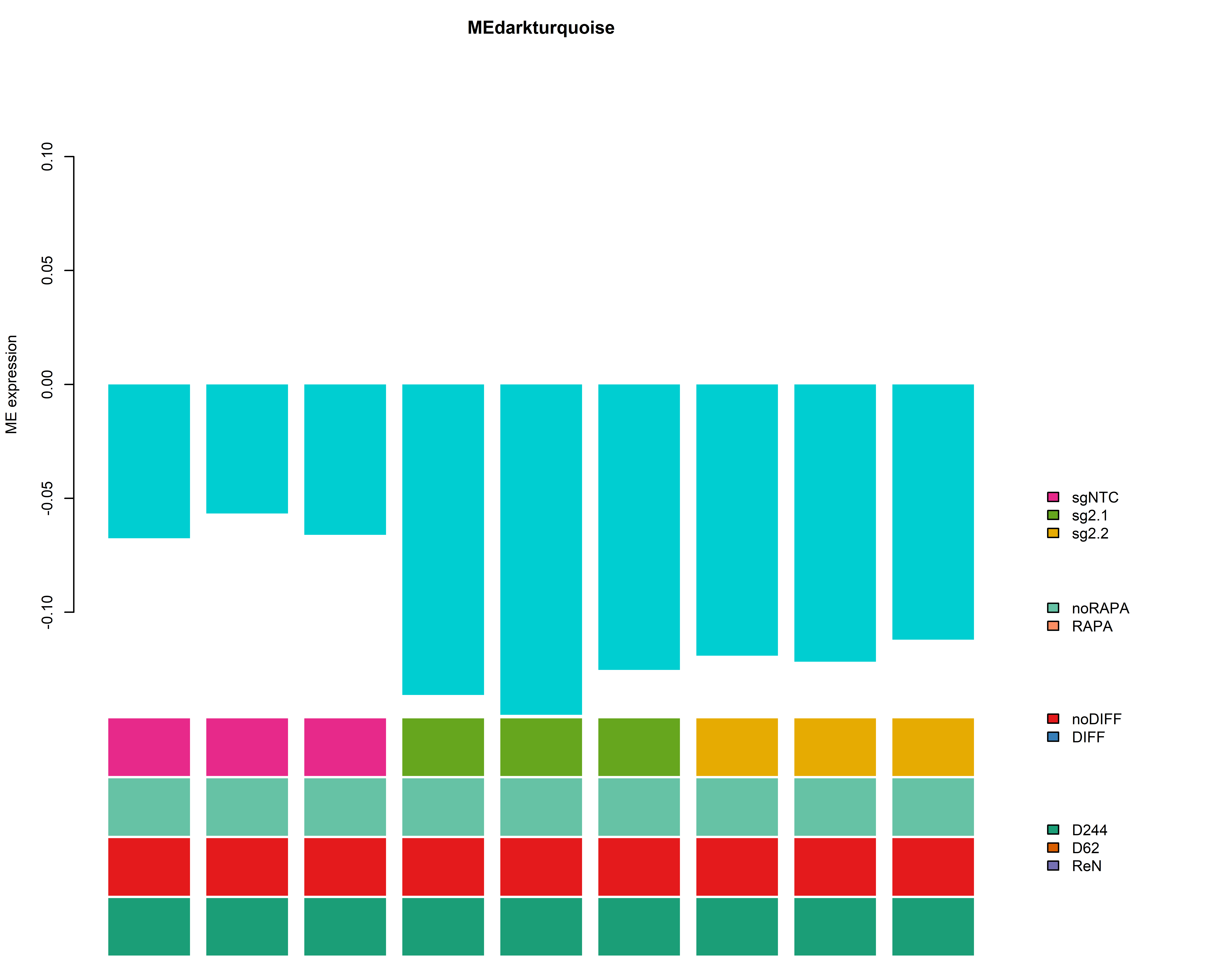
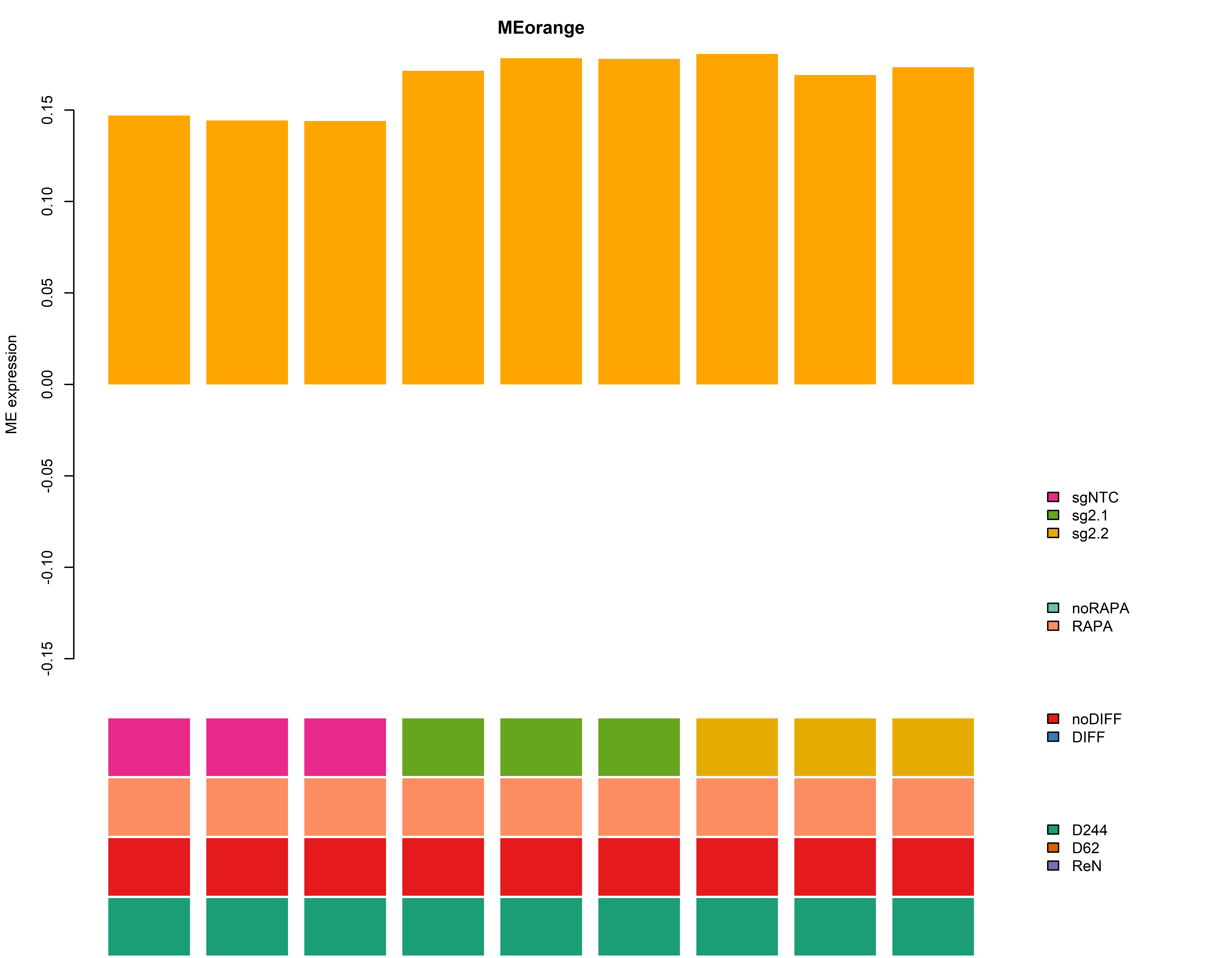
}Eigengene plots all modules
EigengenePlot(data=SampleInfo[,grep("ME", colnames(SampleInfo))],
Sampledata = SampleInfo,
samplesincl=rep(T, nrow(SampleInfo)))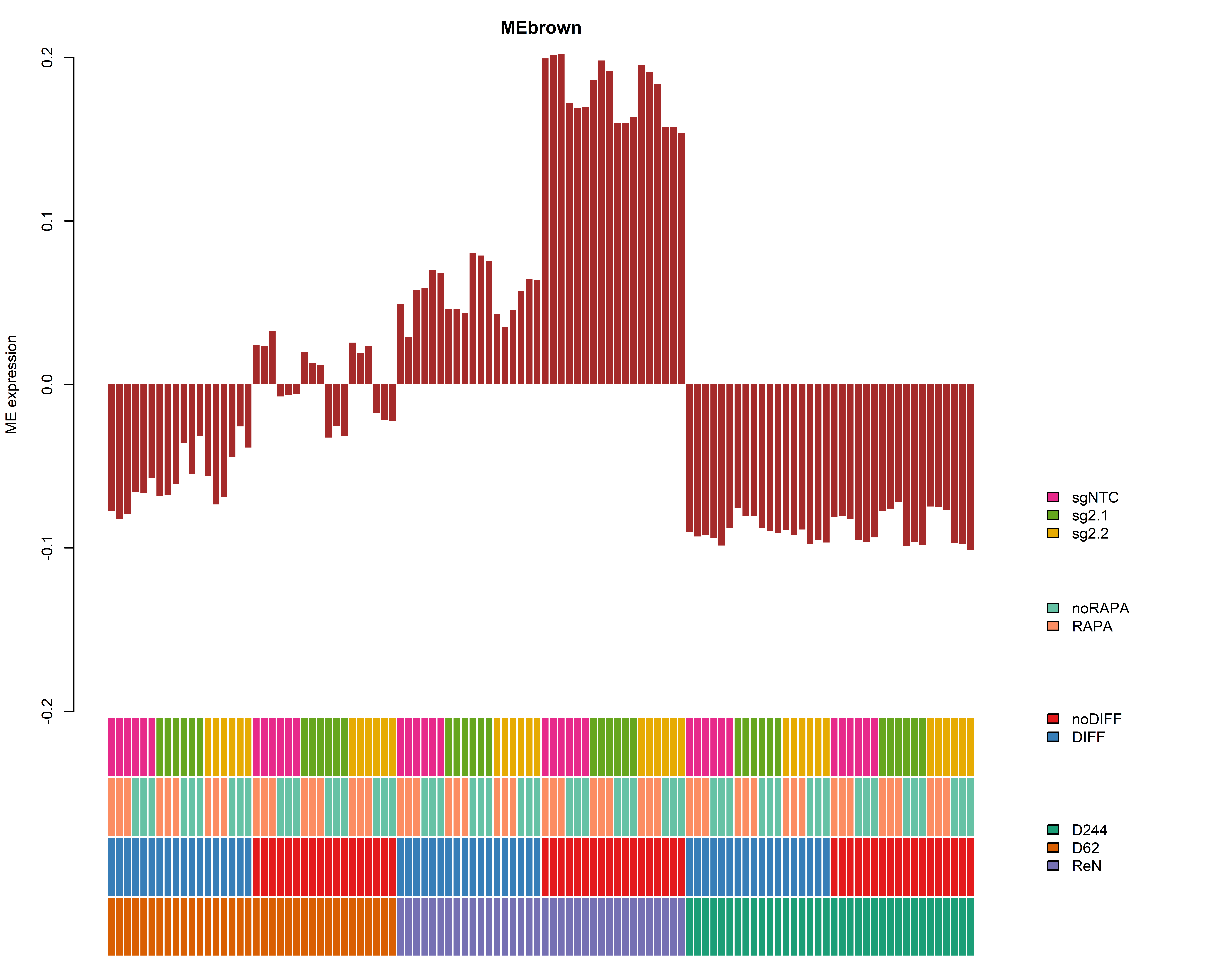
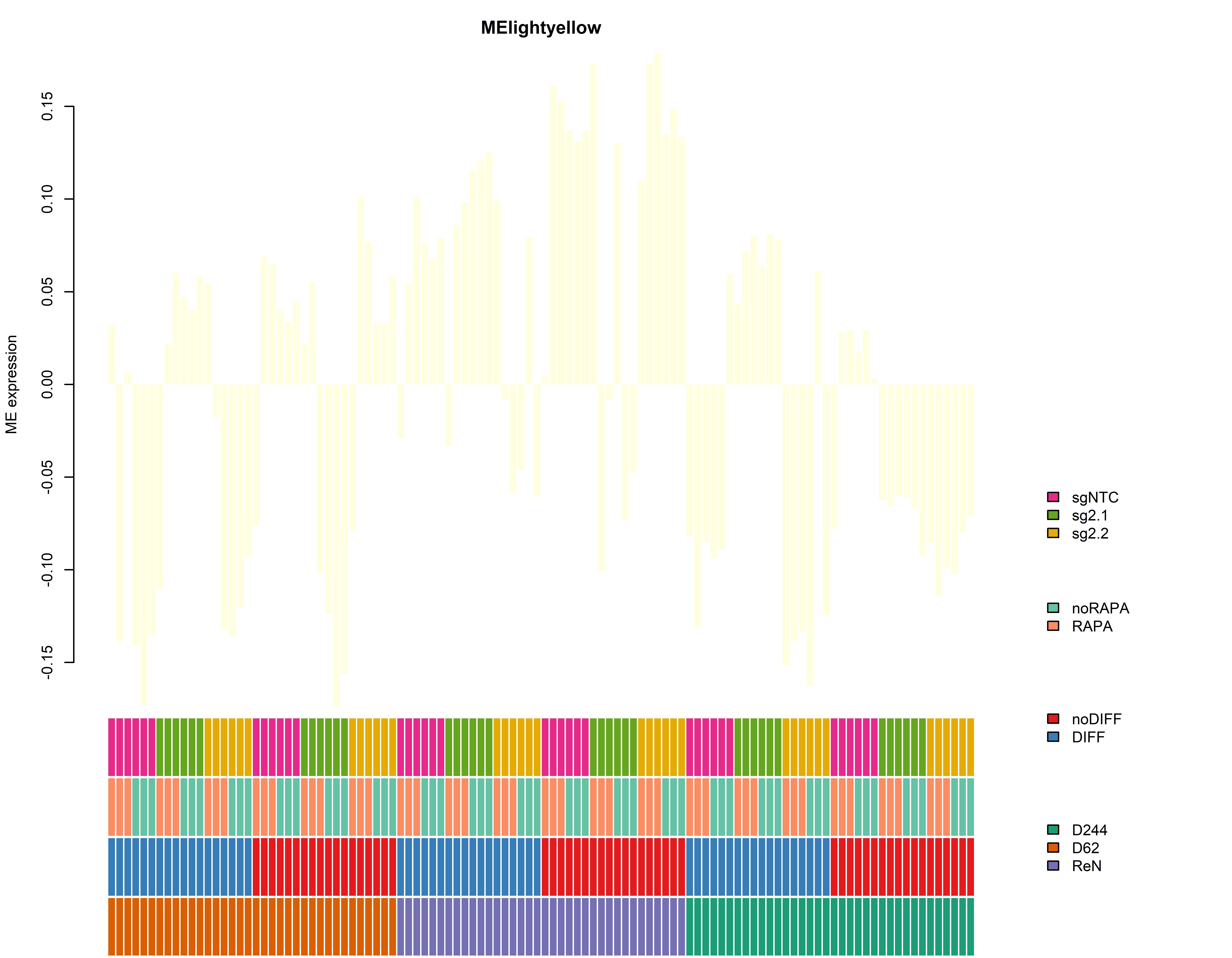
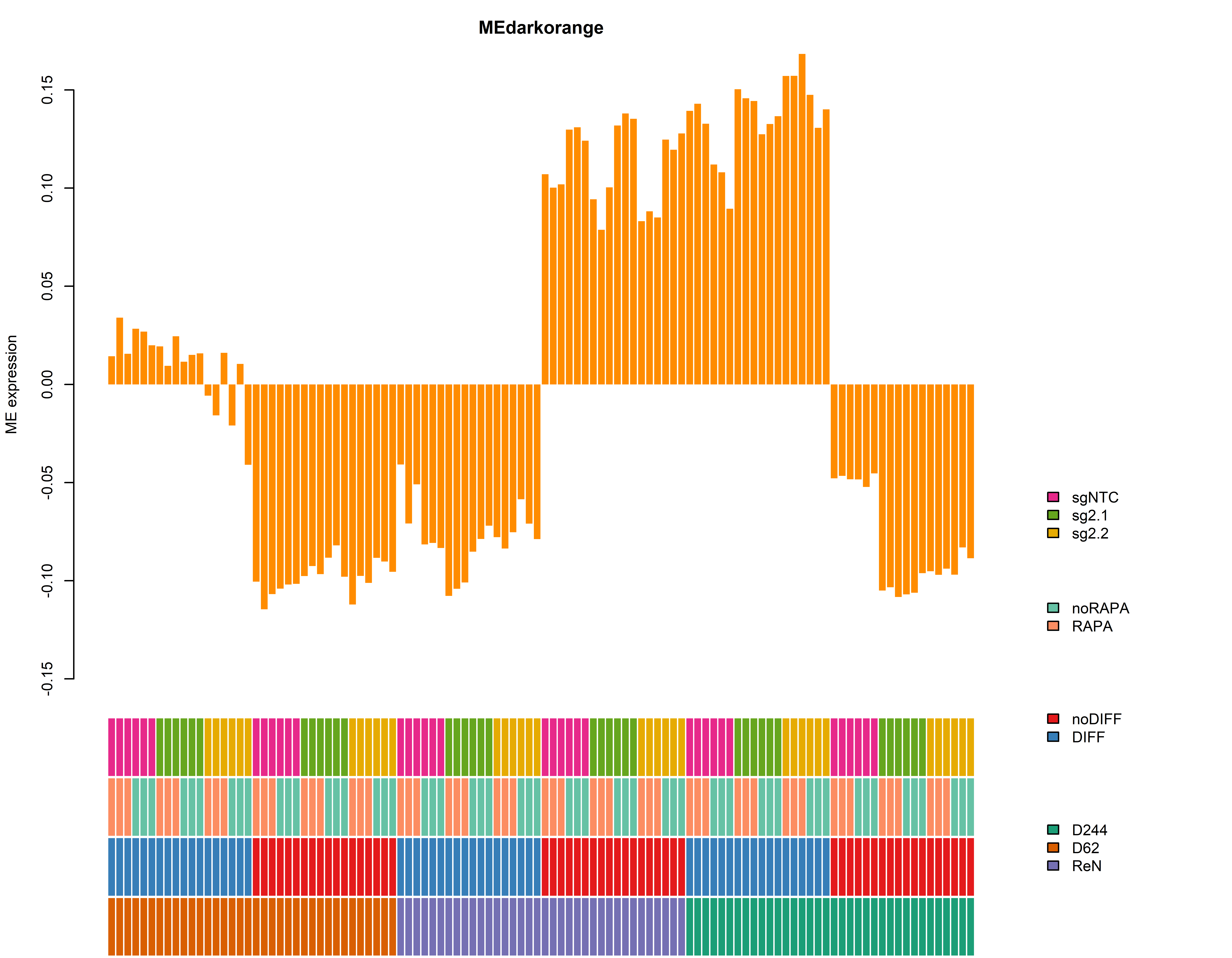
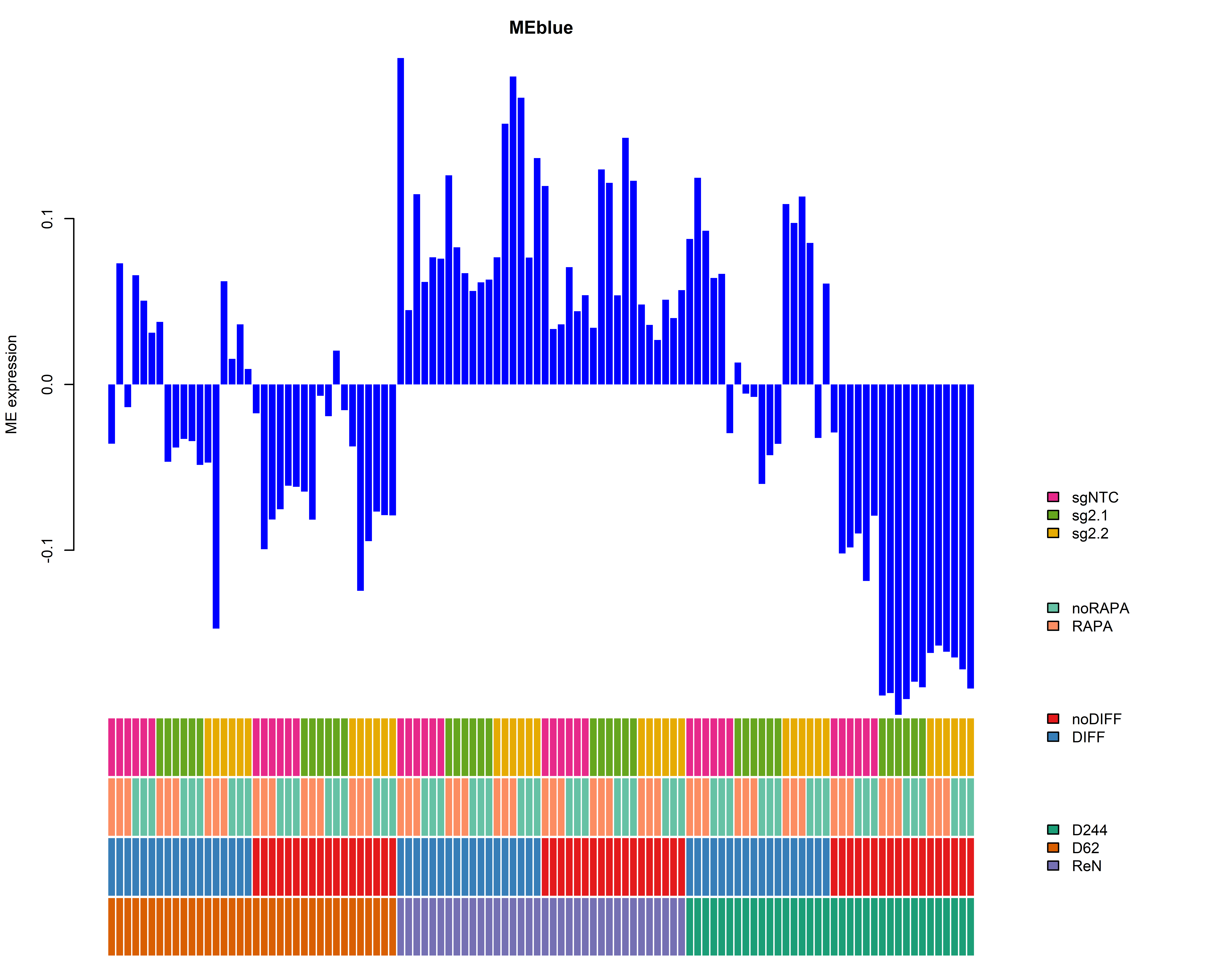
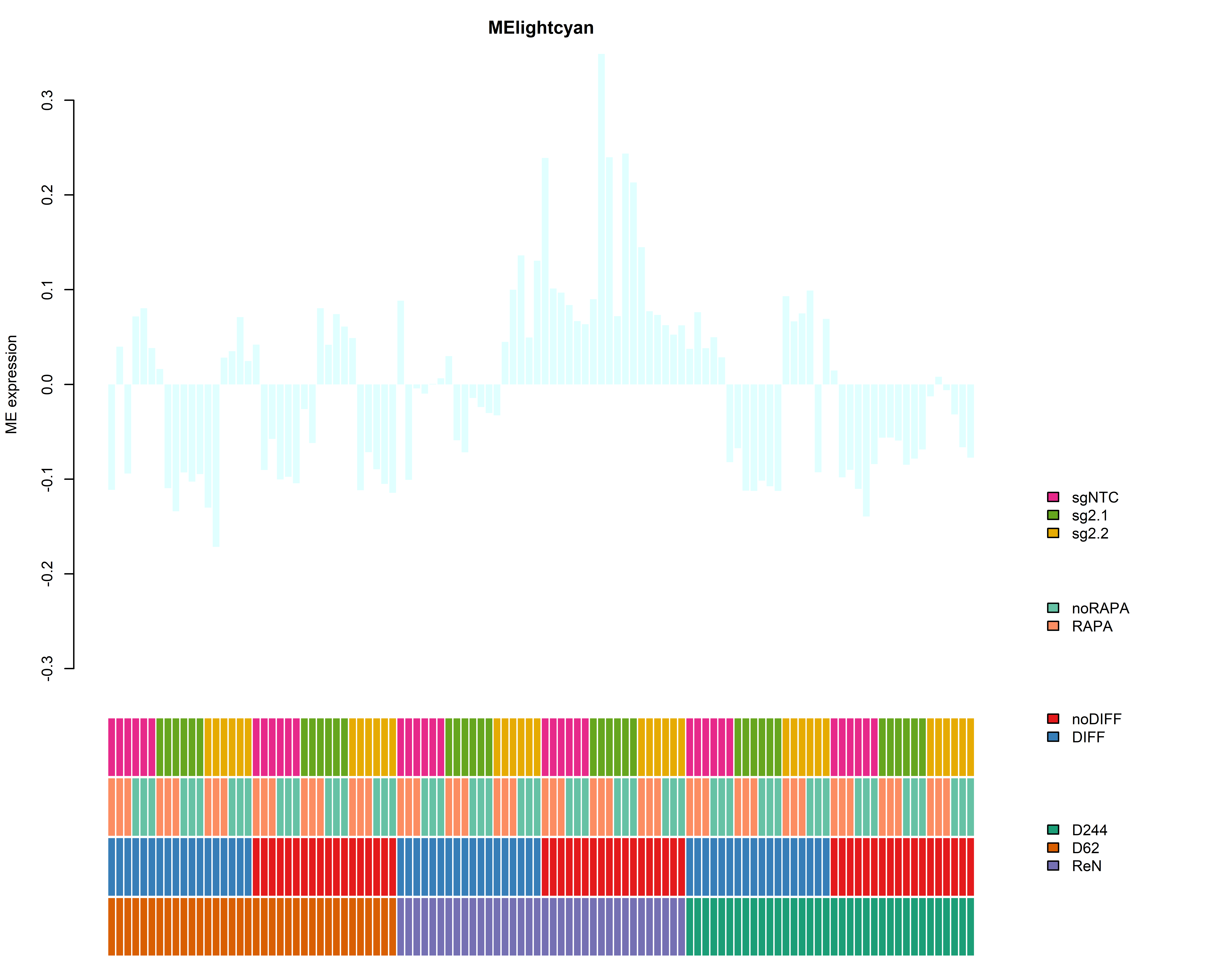
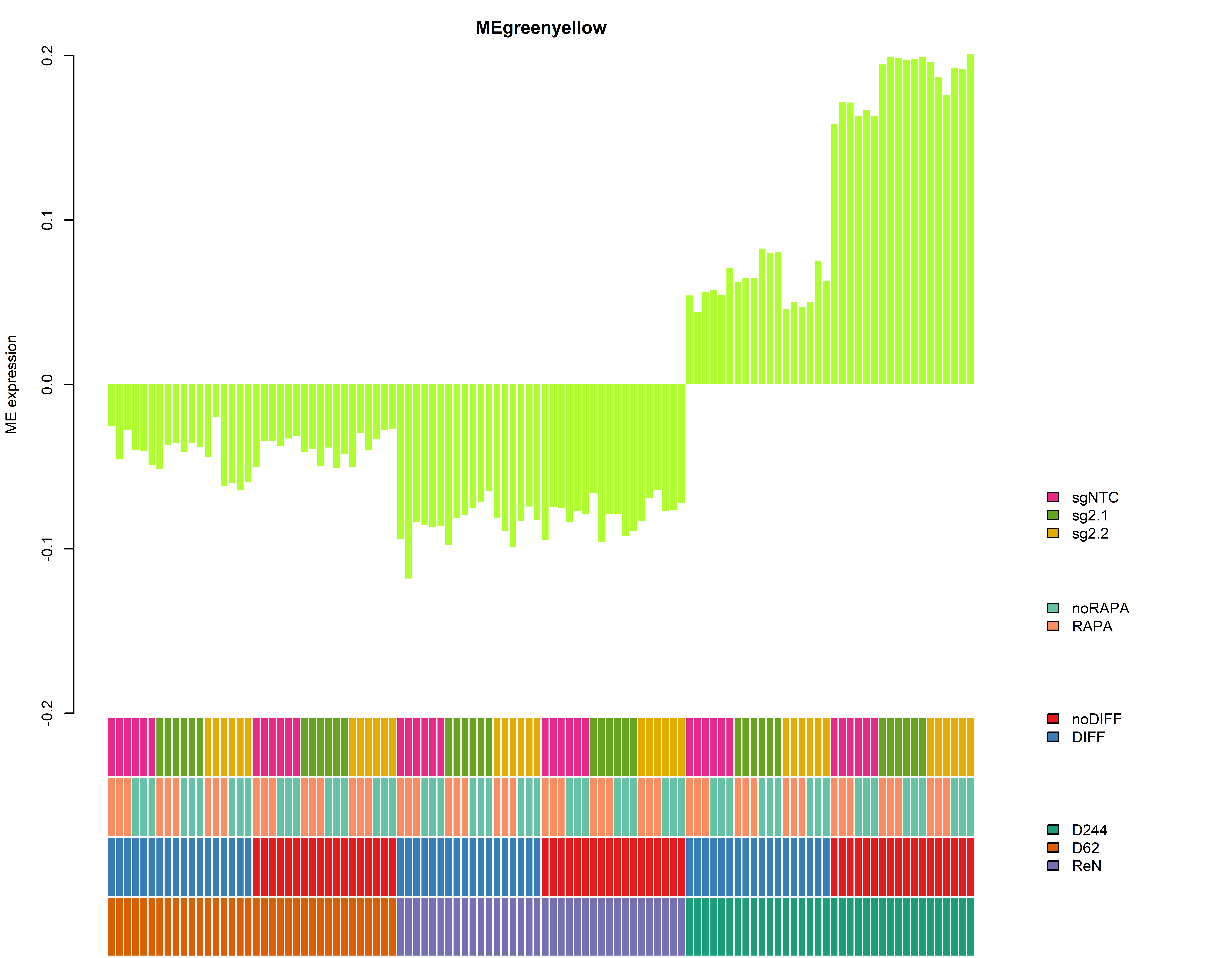
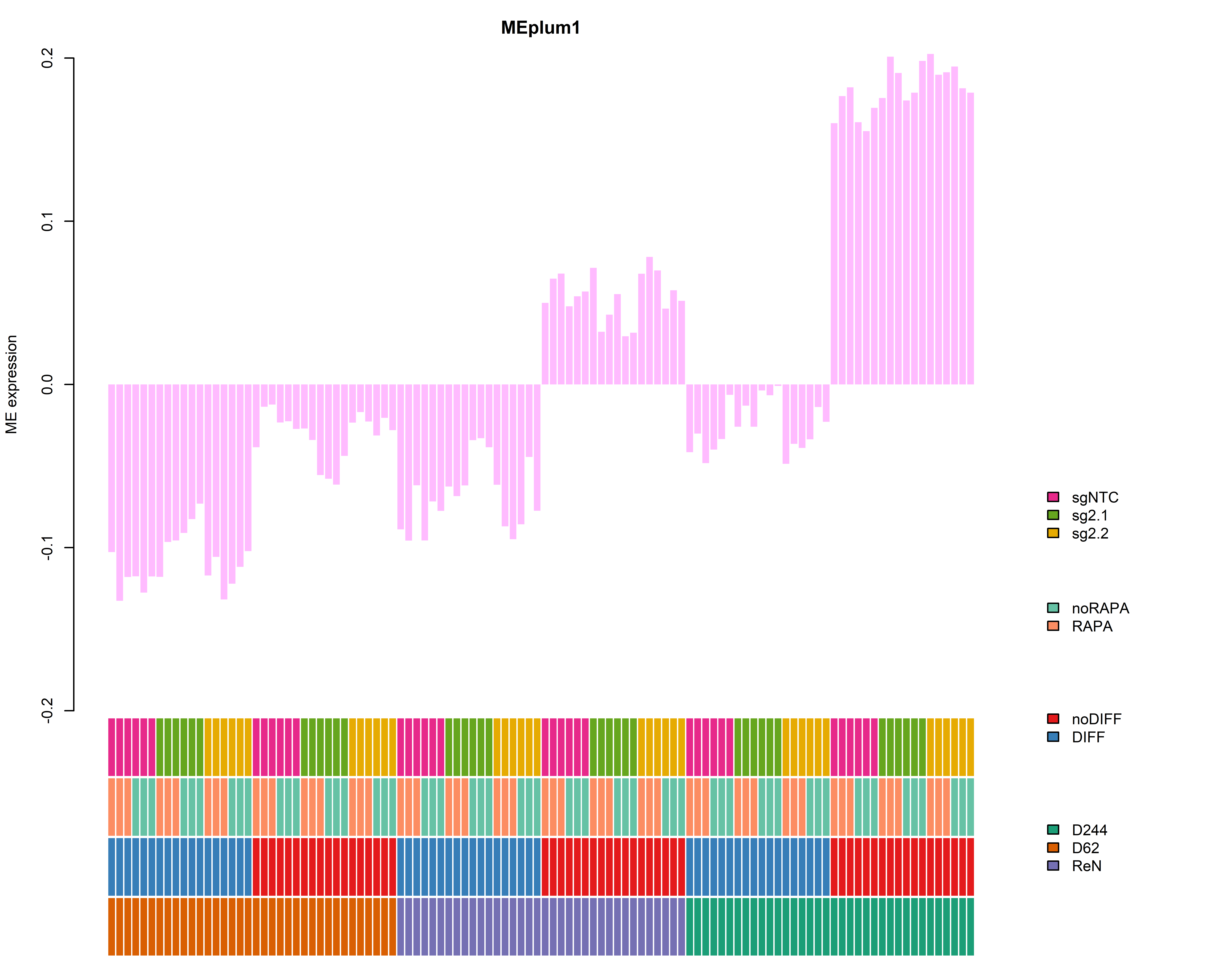
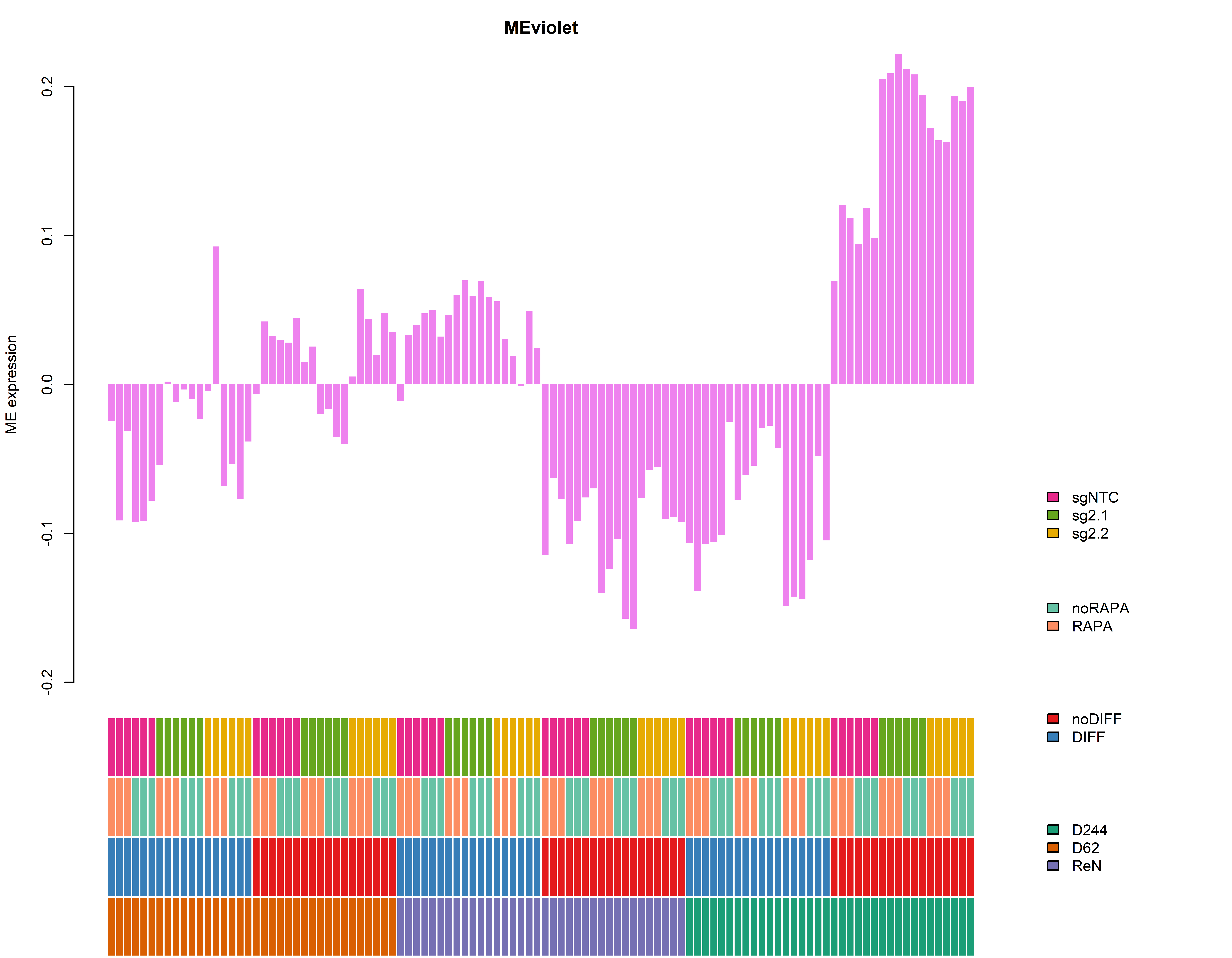
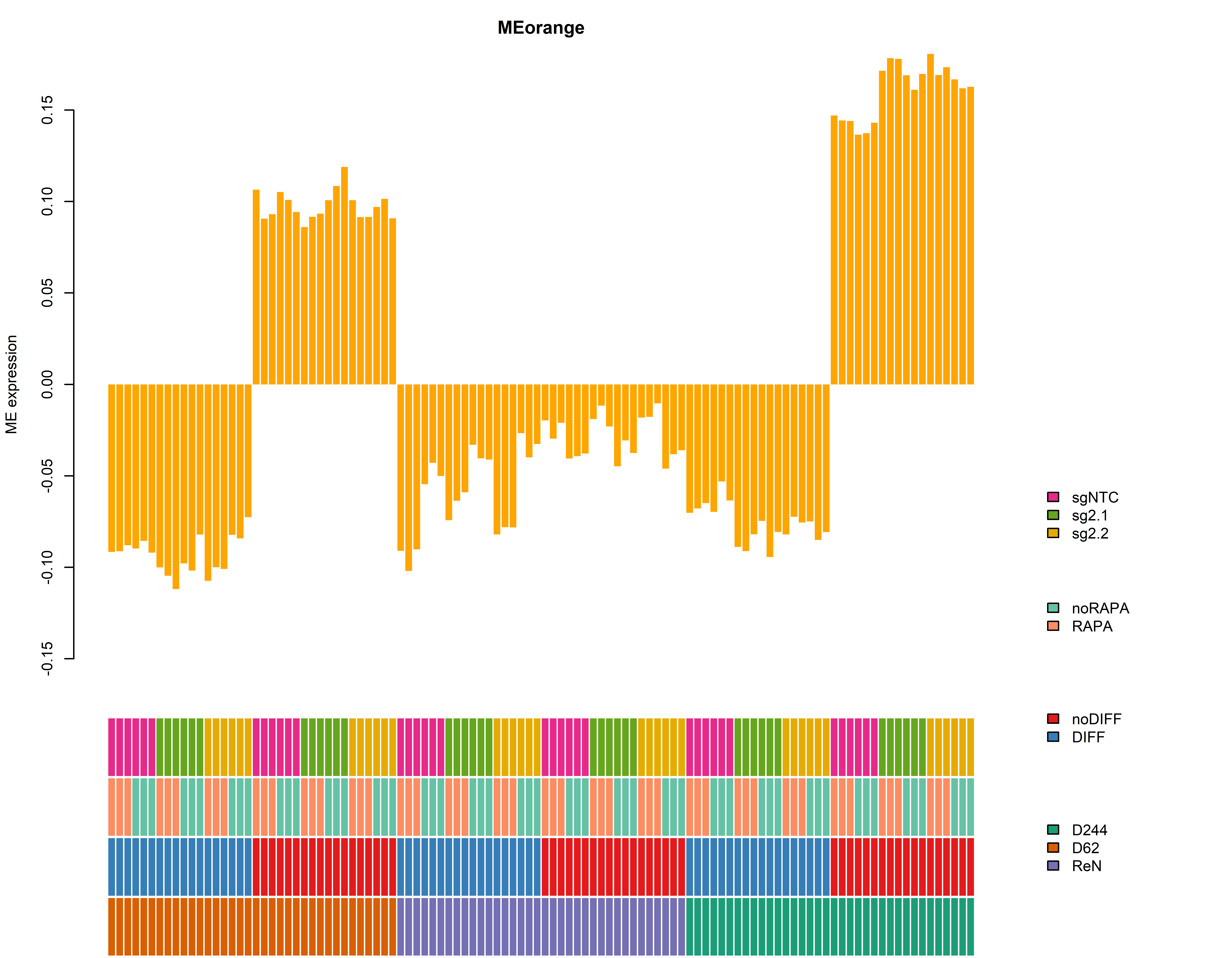
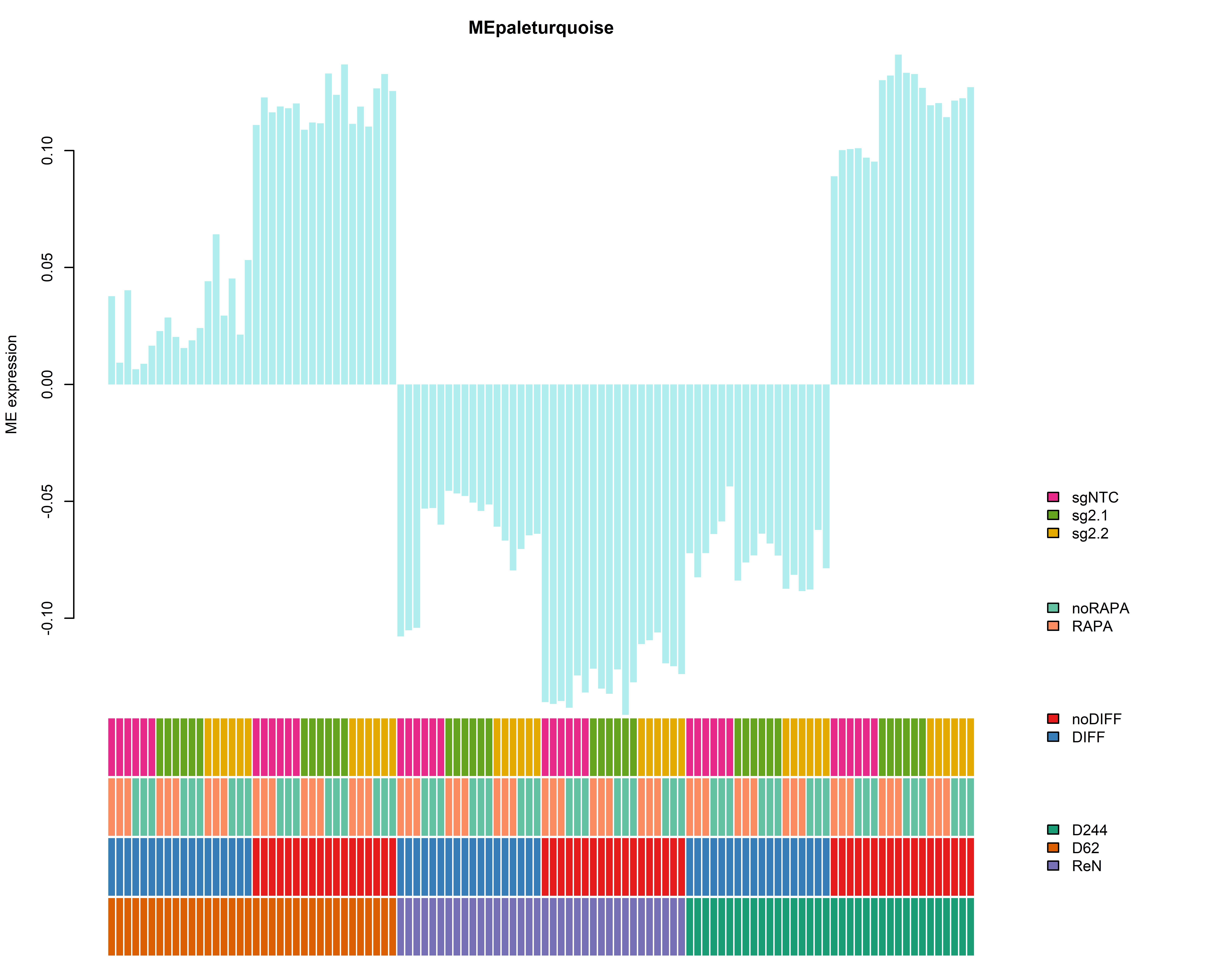
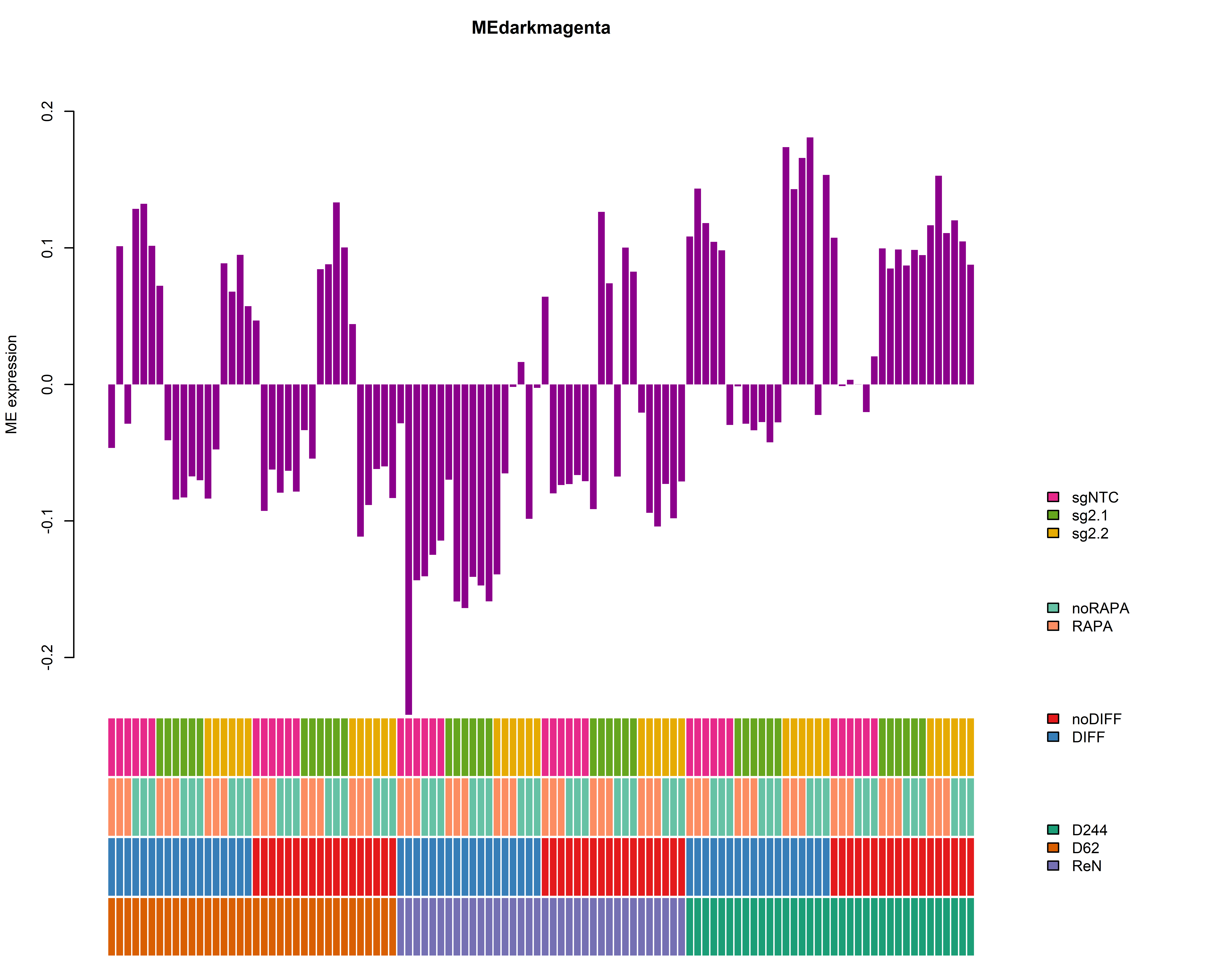
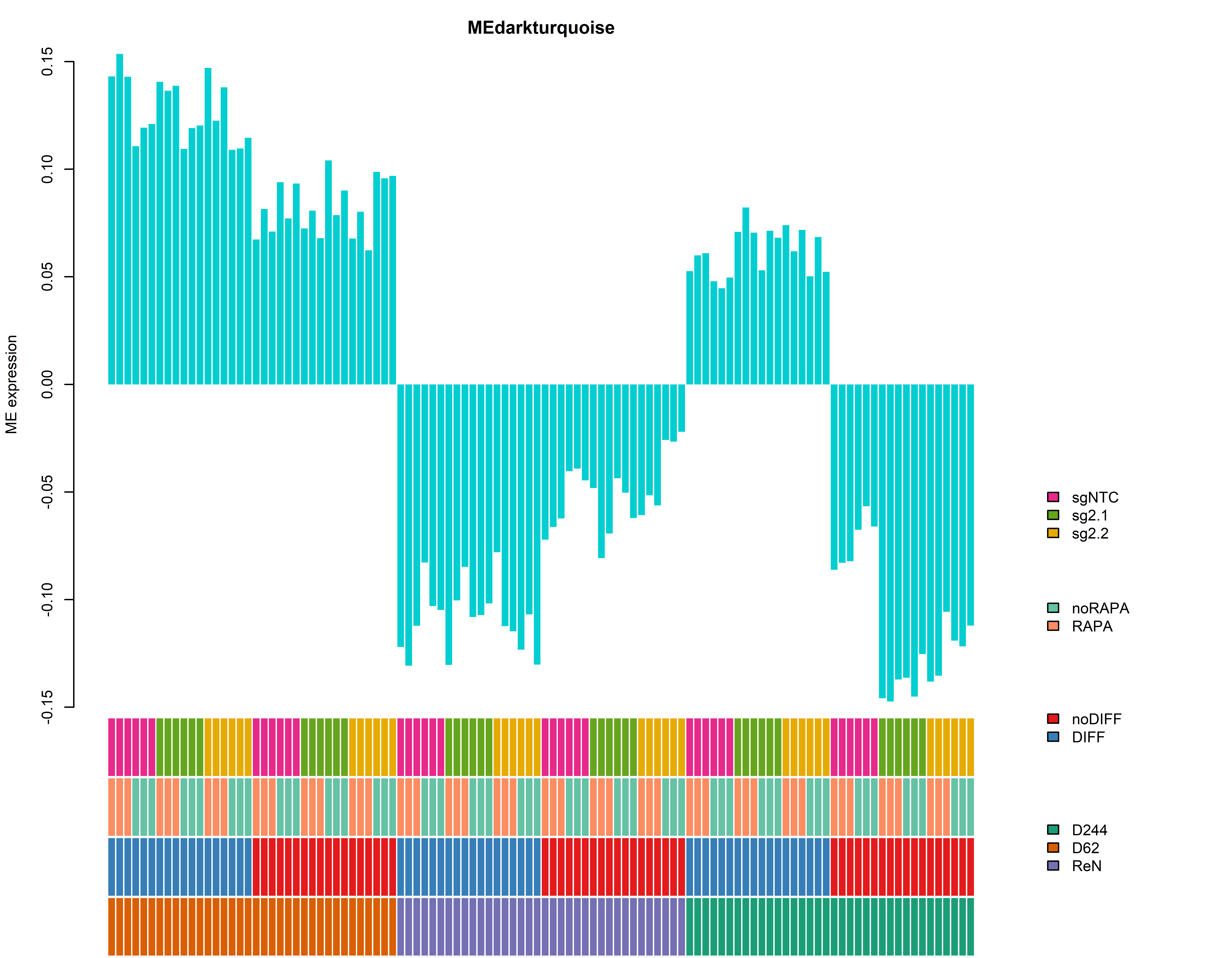
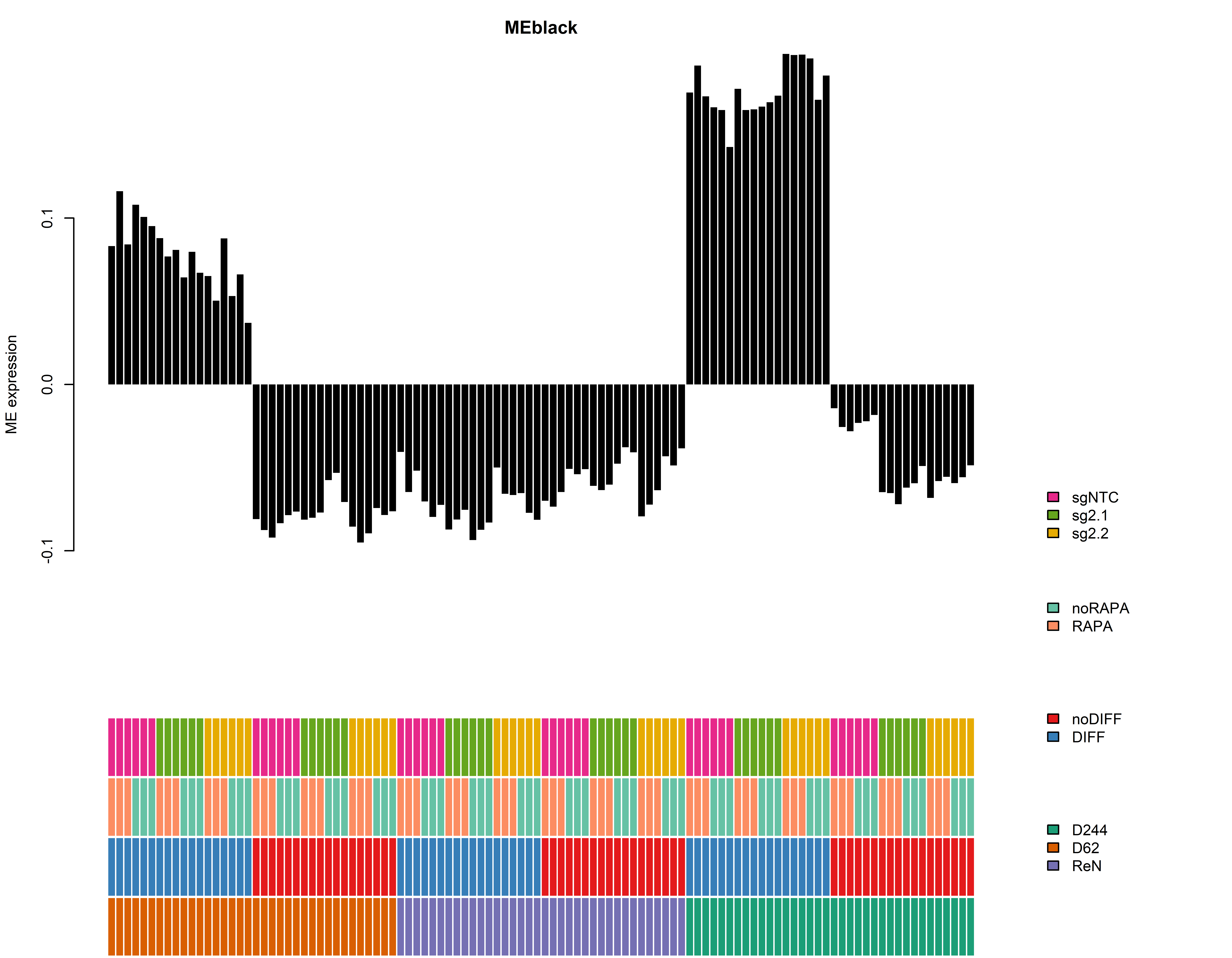
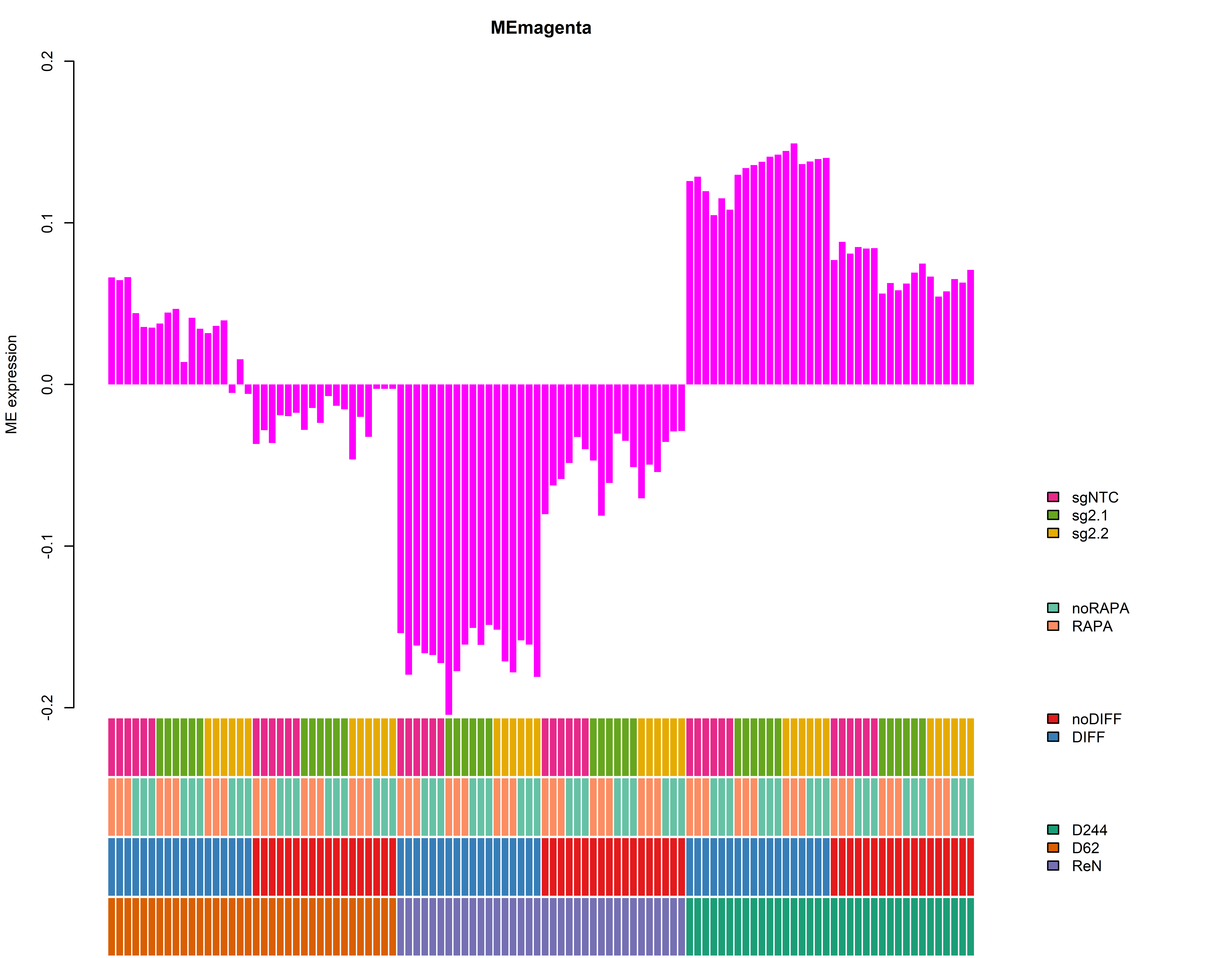
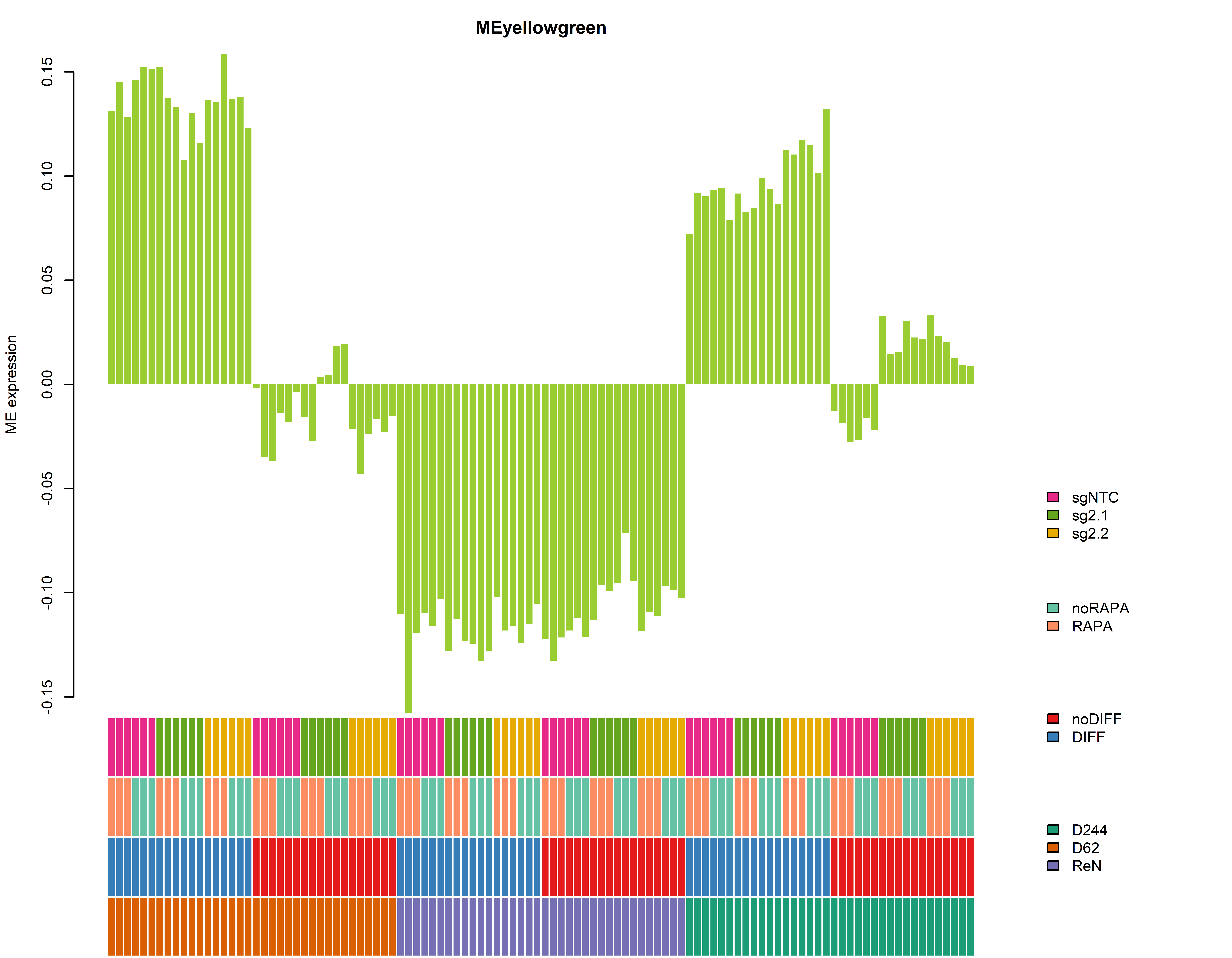
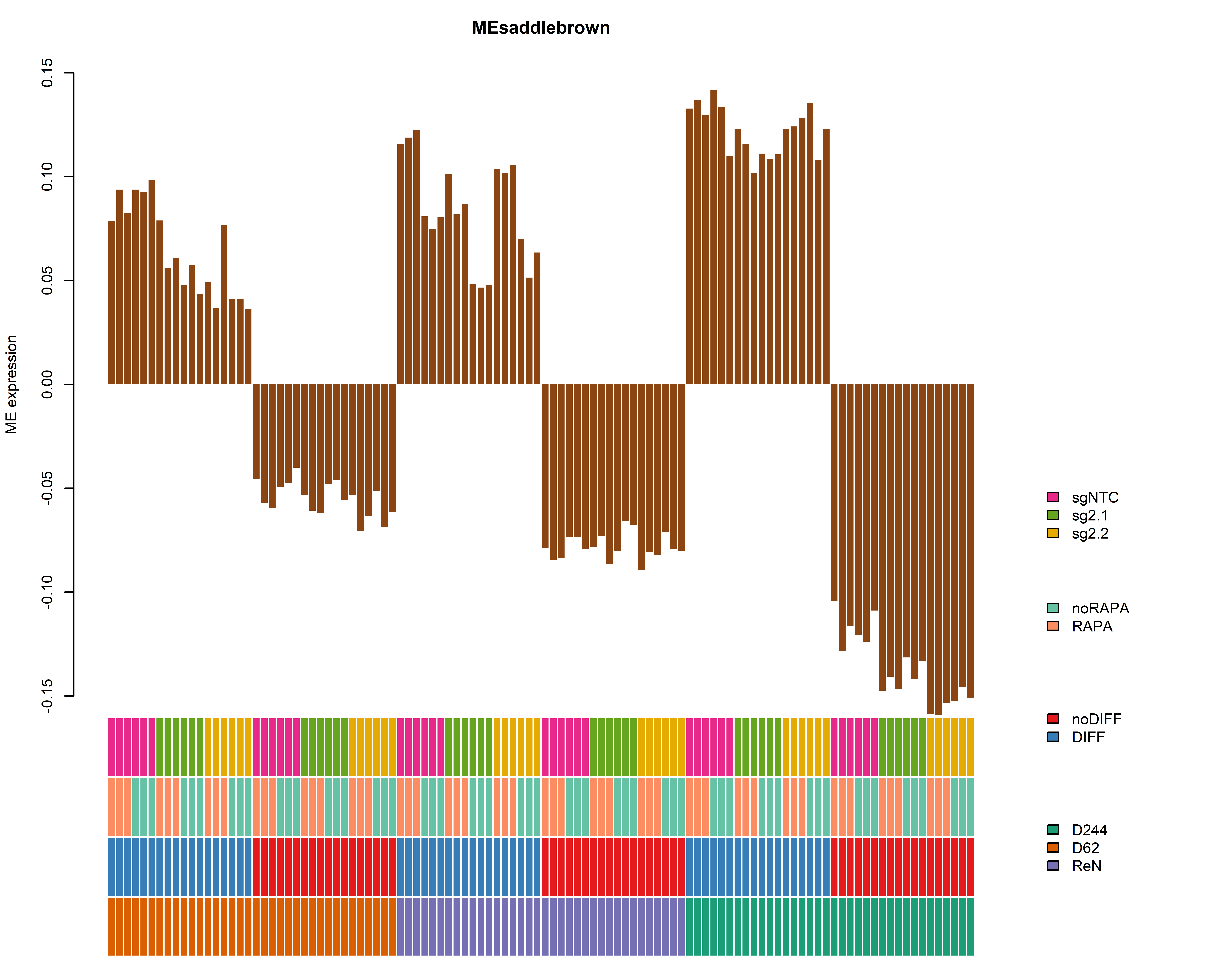
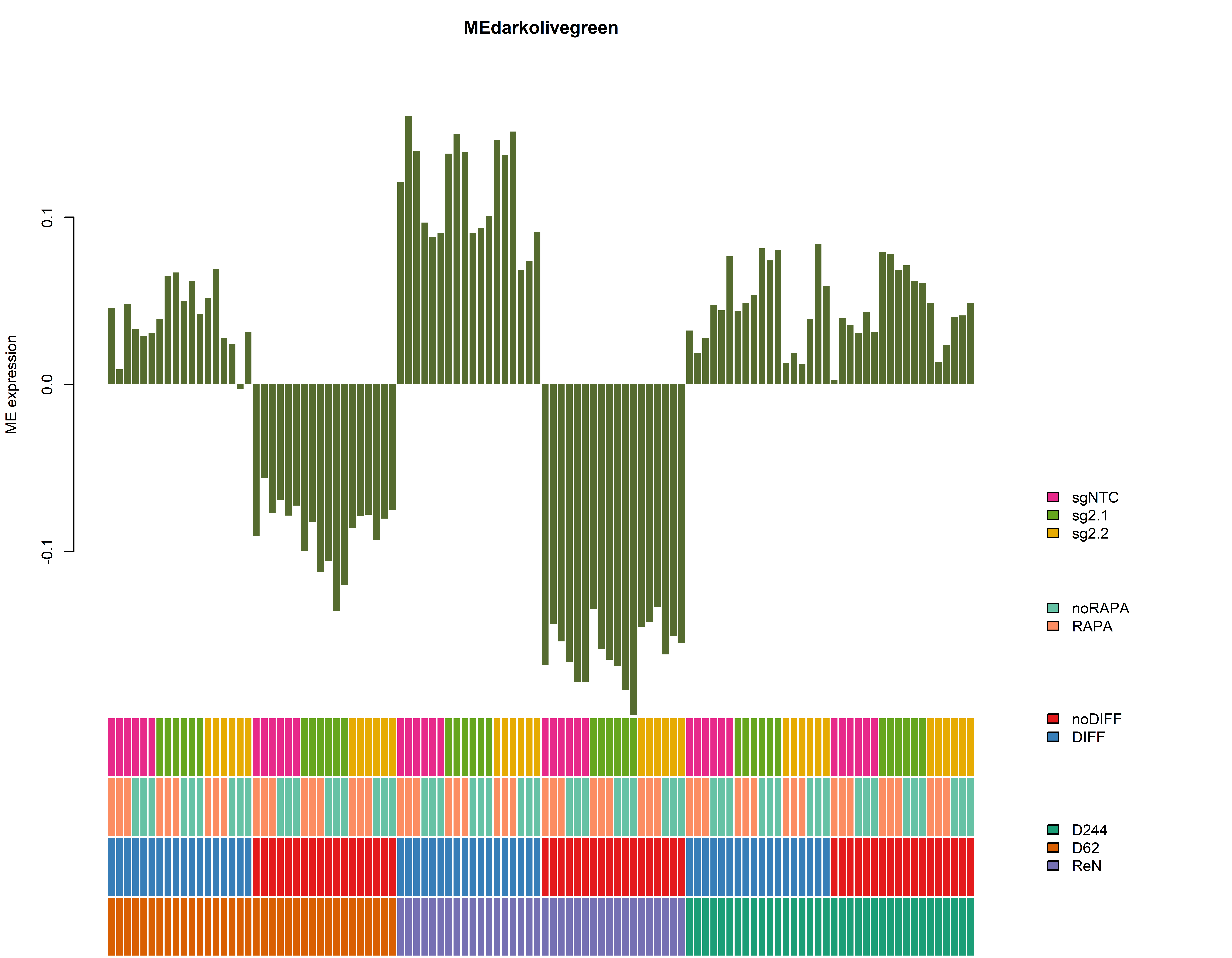
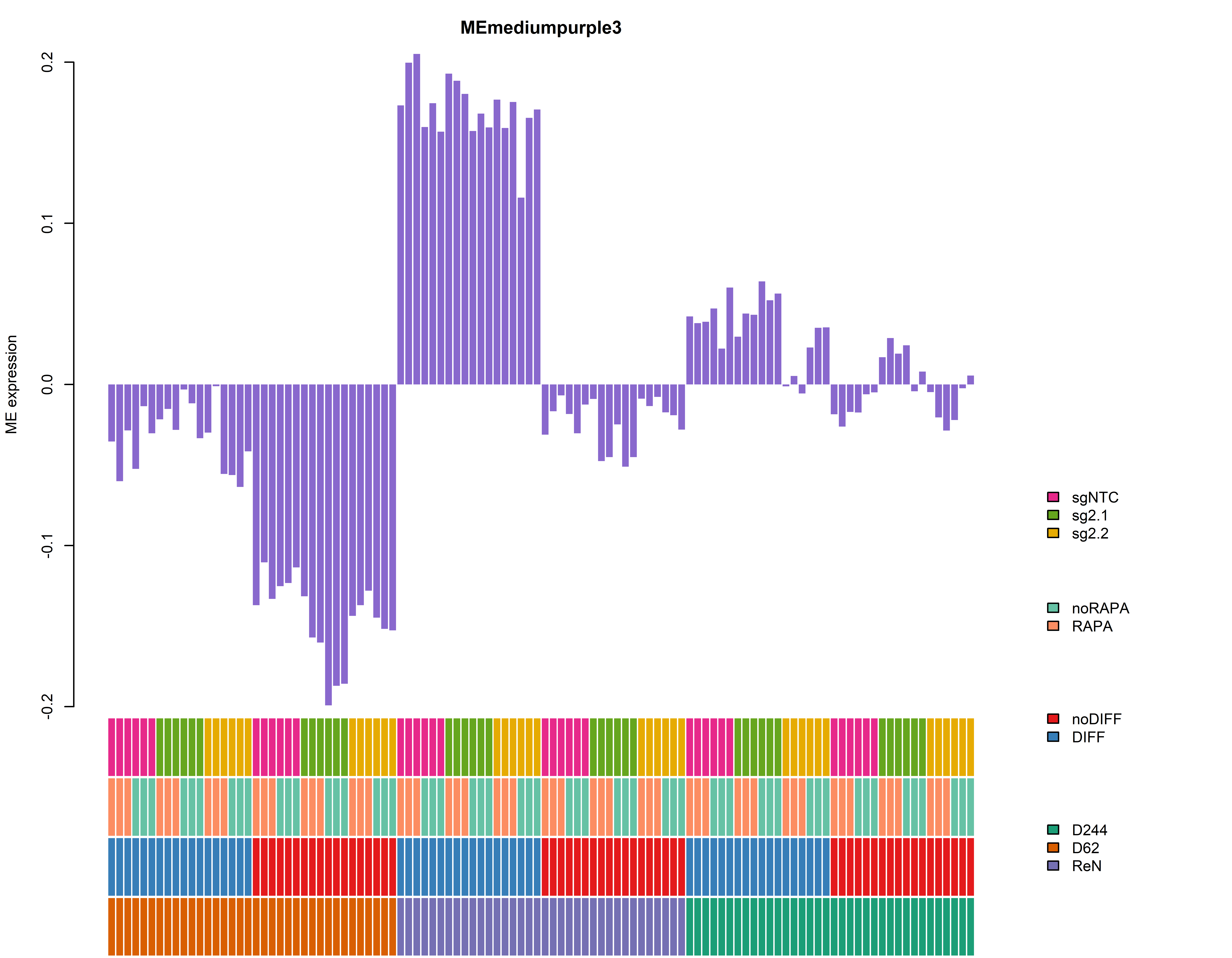
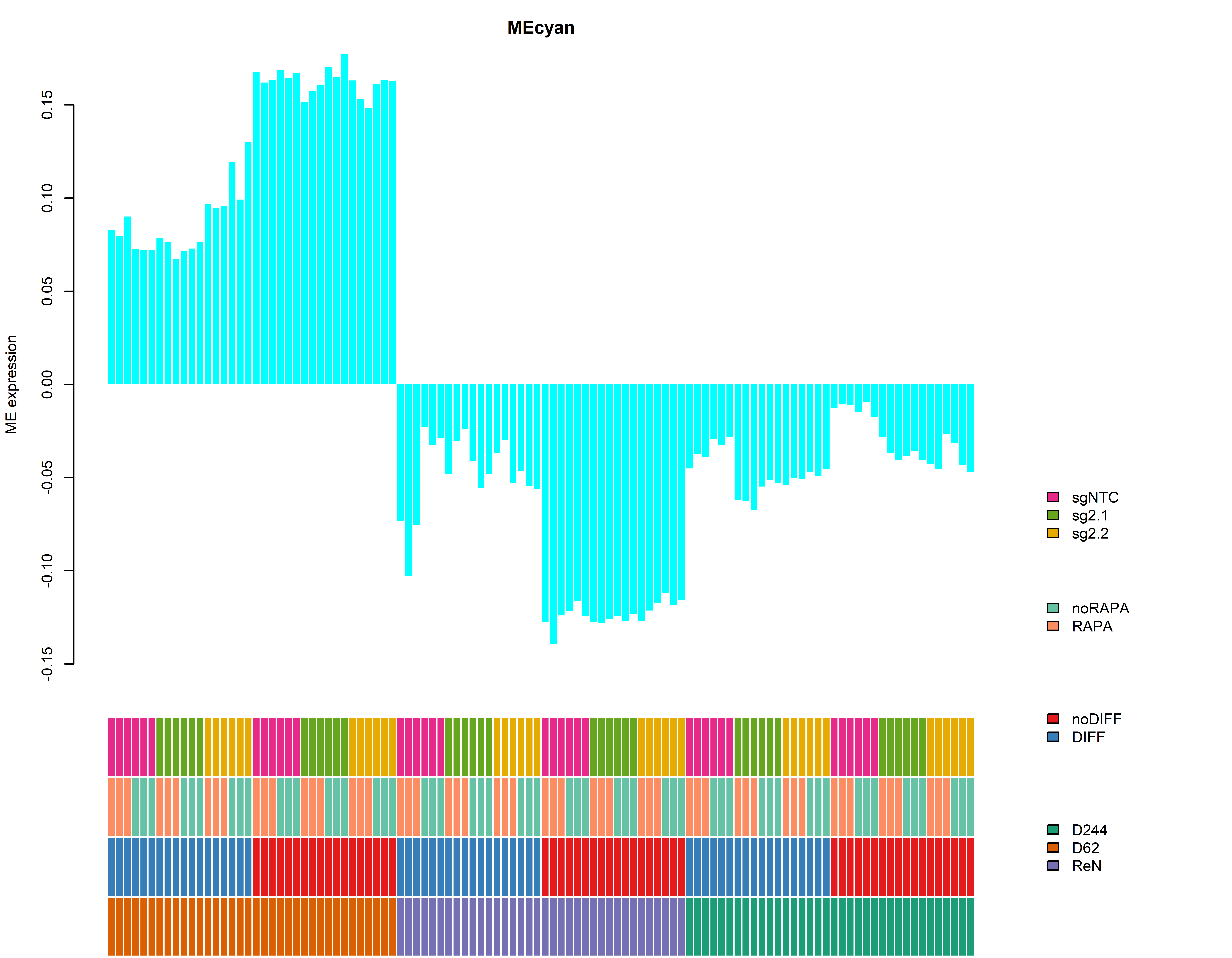
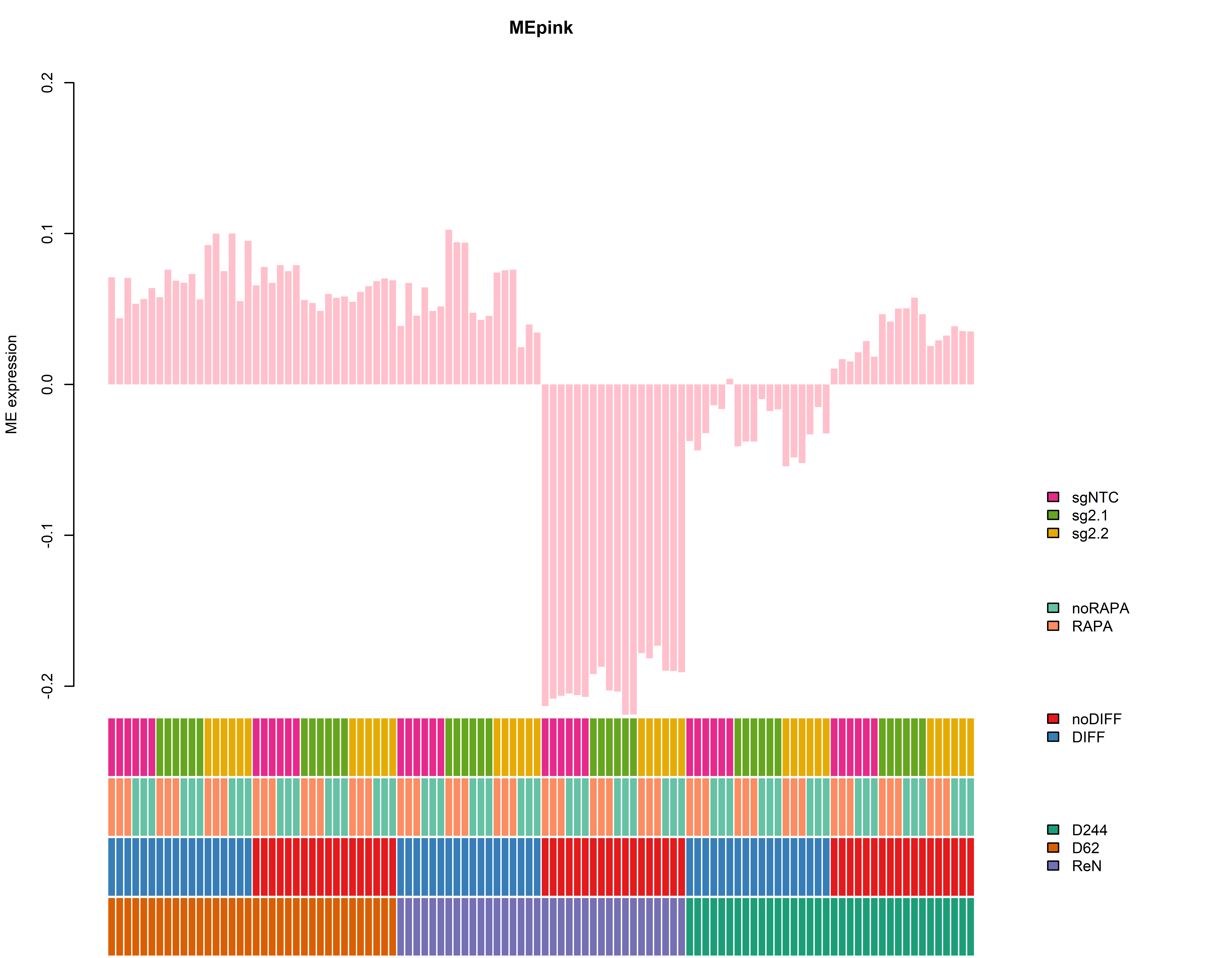
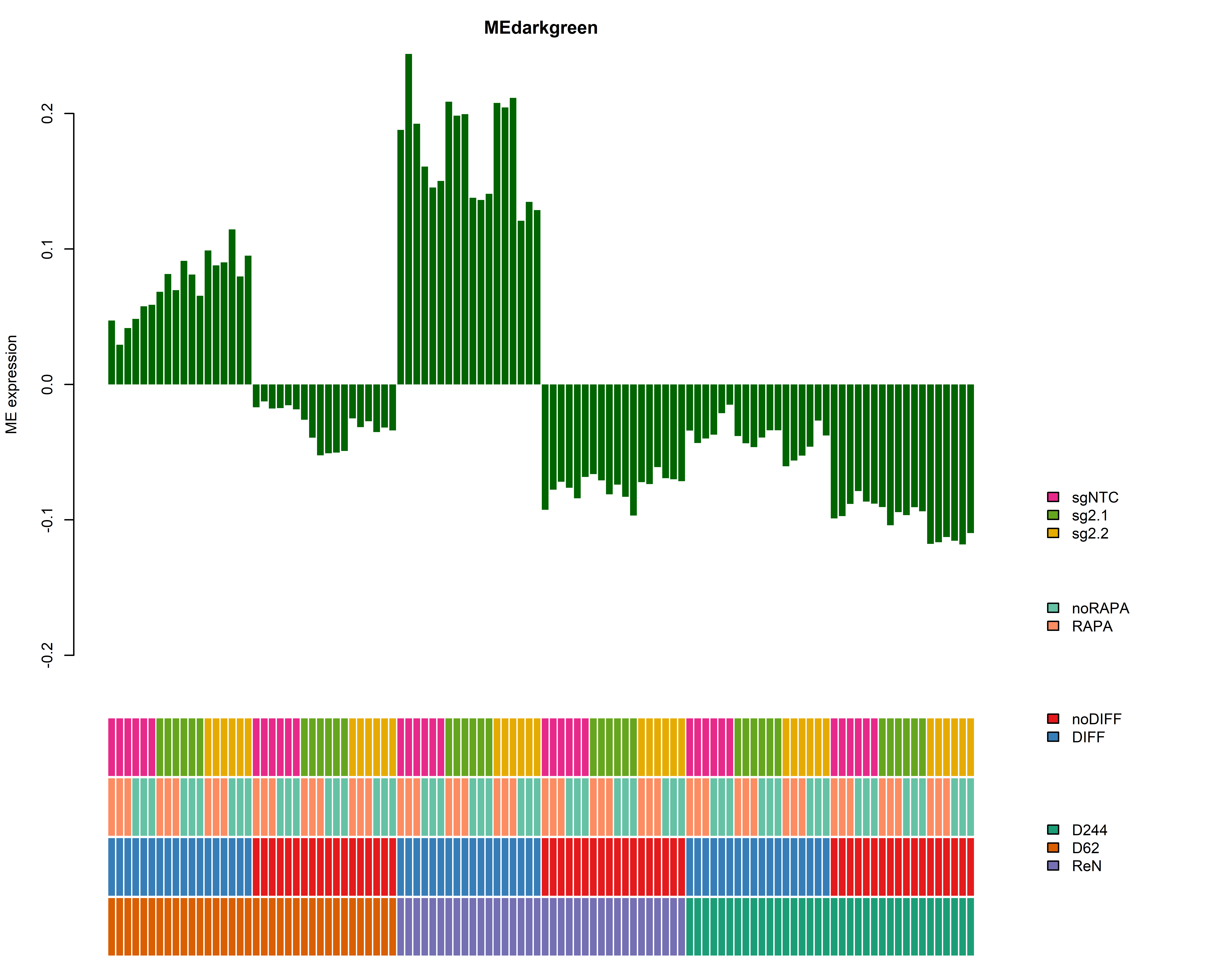
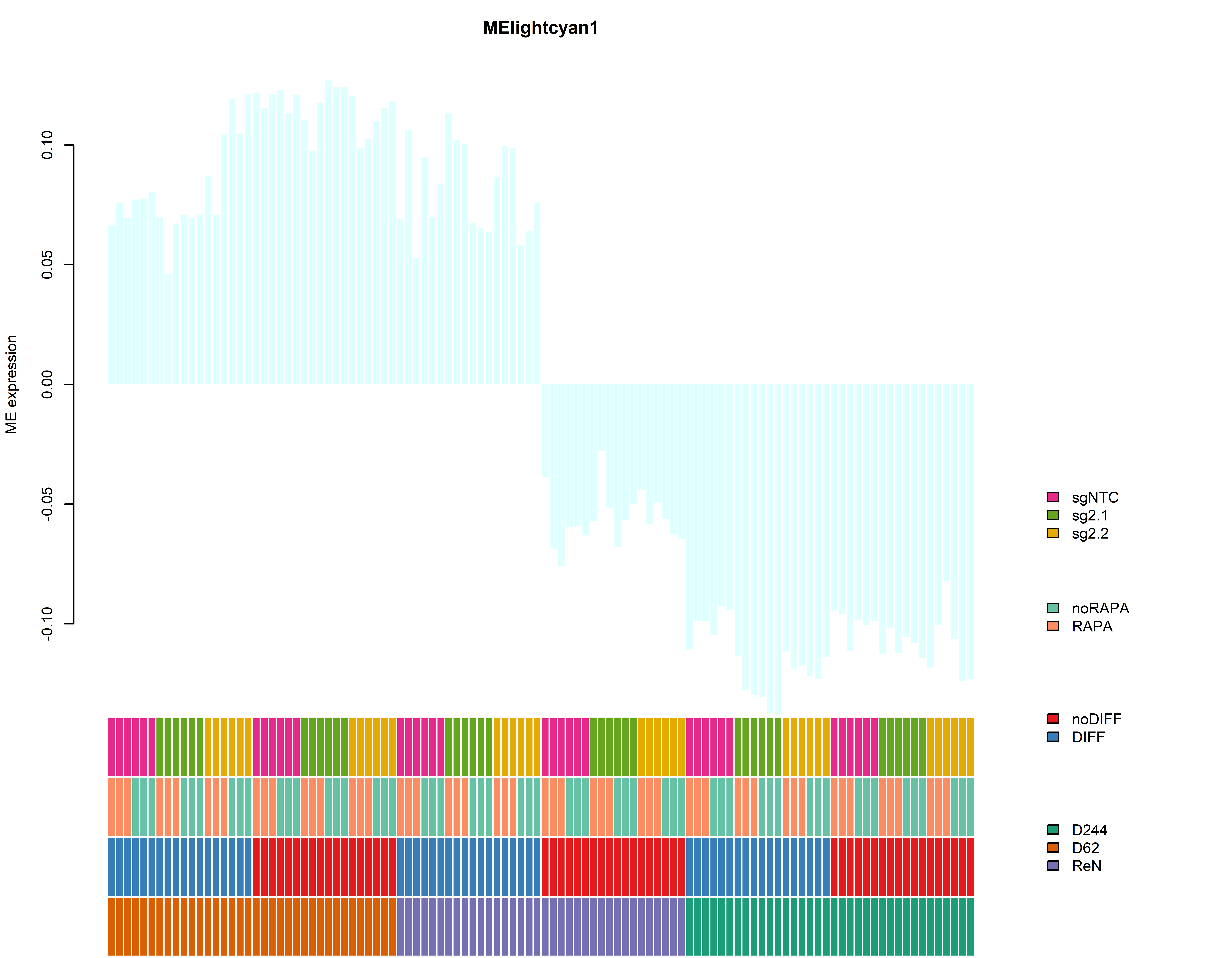
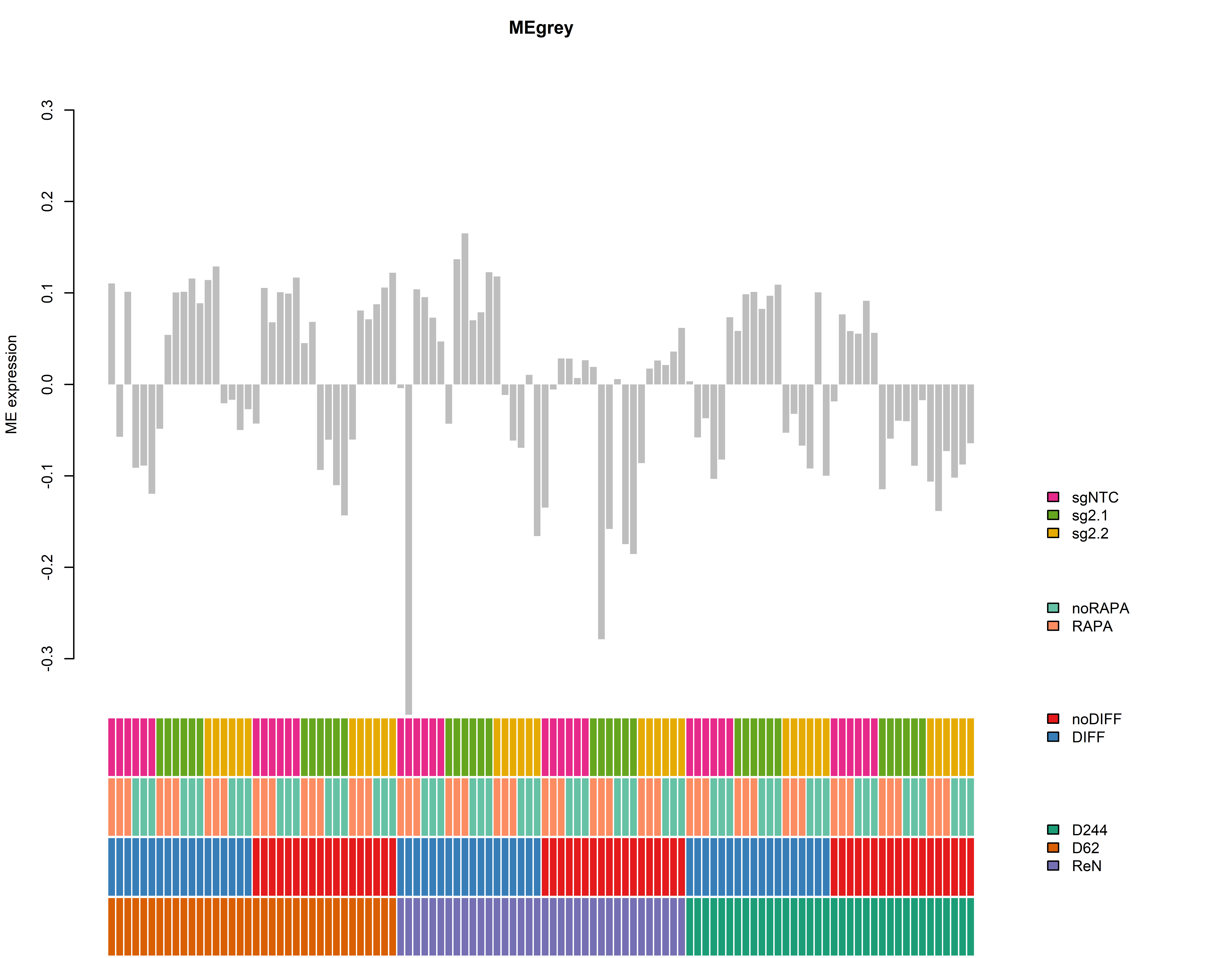
WGCNA module plots only D62
idx=SampleInfo$CellLine=="D62"
EigengenePlot(data=SampleInfo[idx,grep("ME", colnames(SampleInfo))],
Sampledata = SampleInfo[idx,],
samplesincl=idx)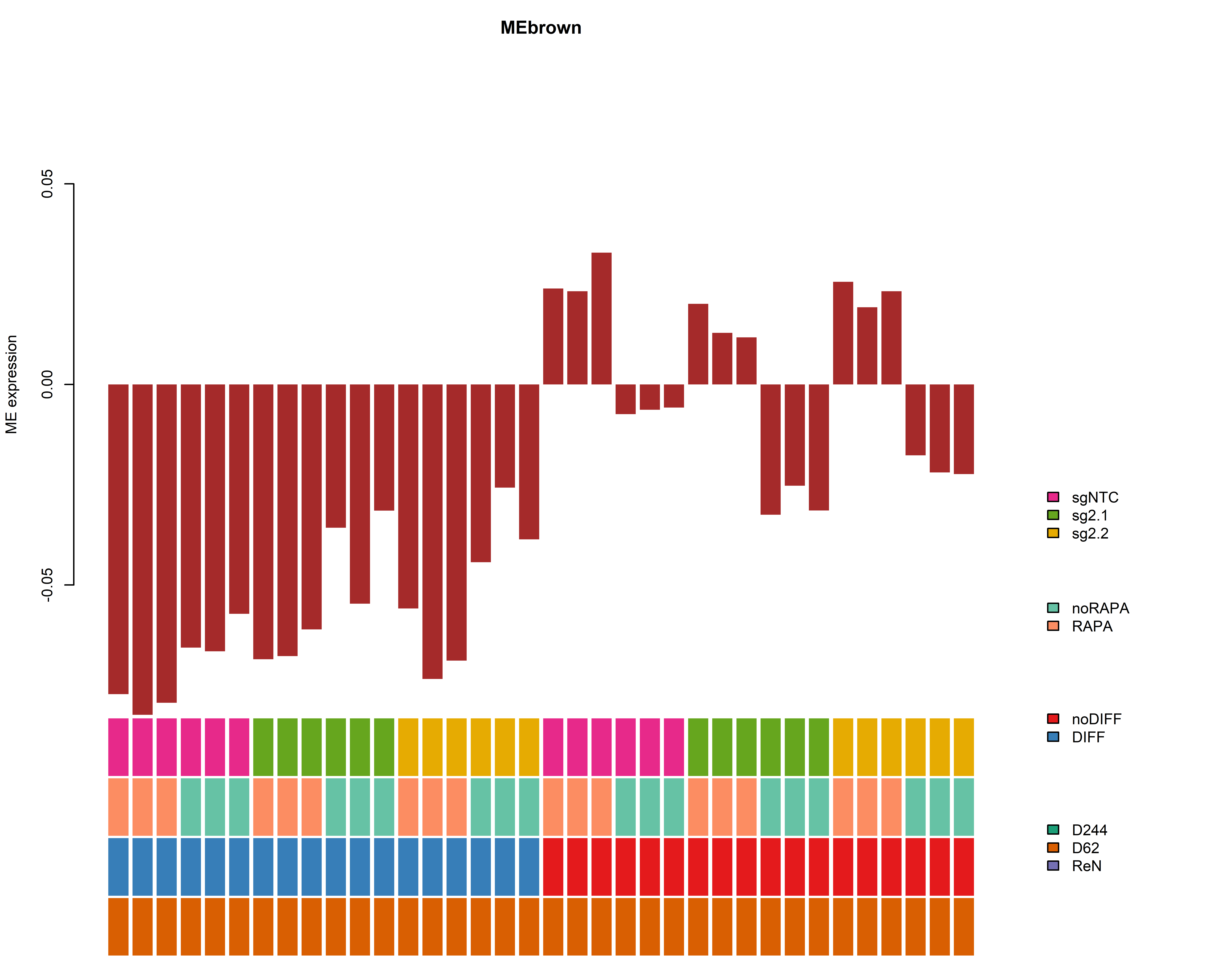
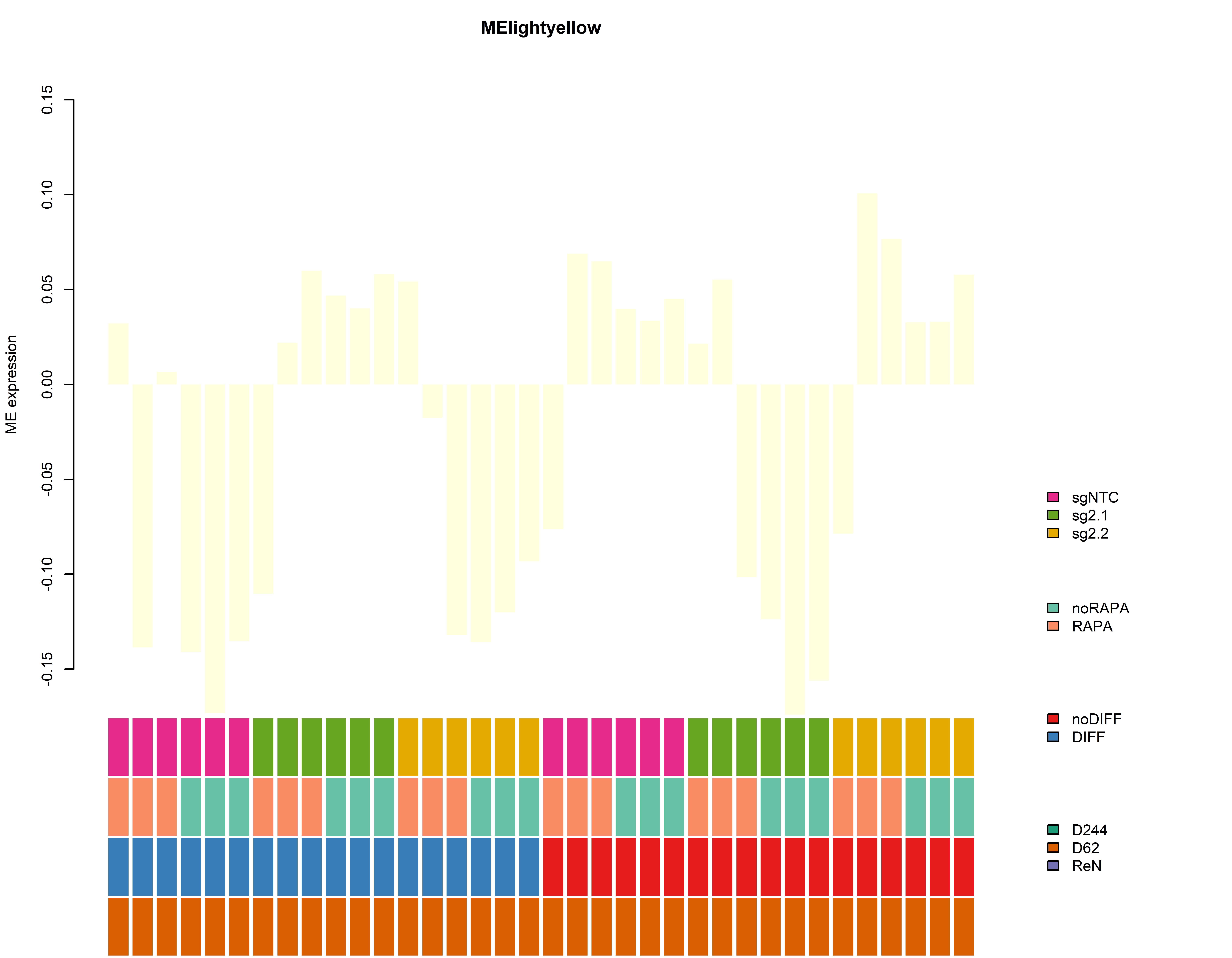
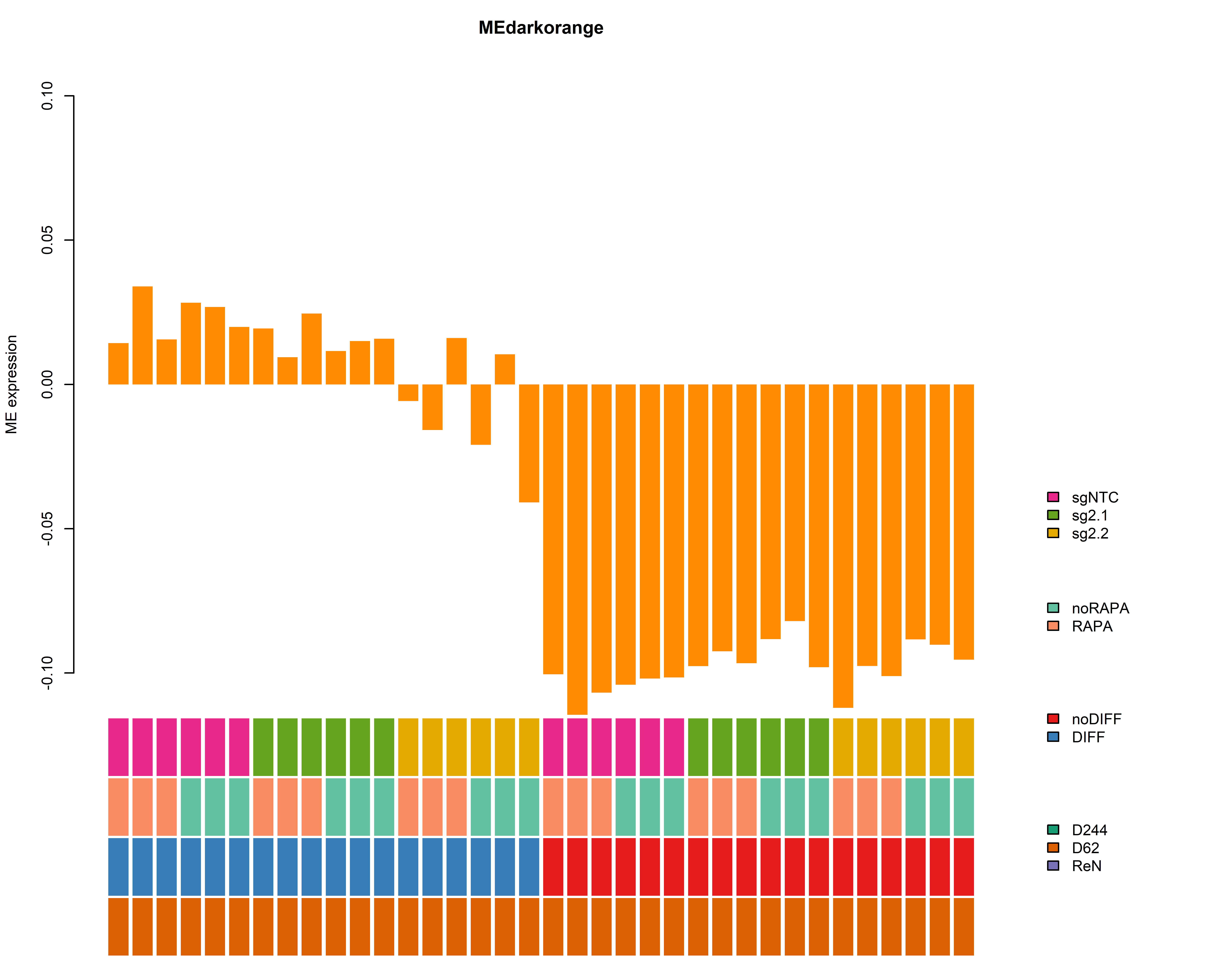
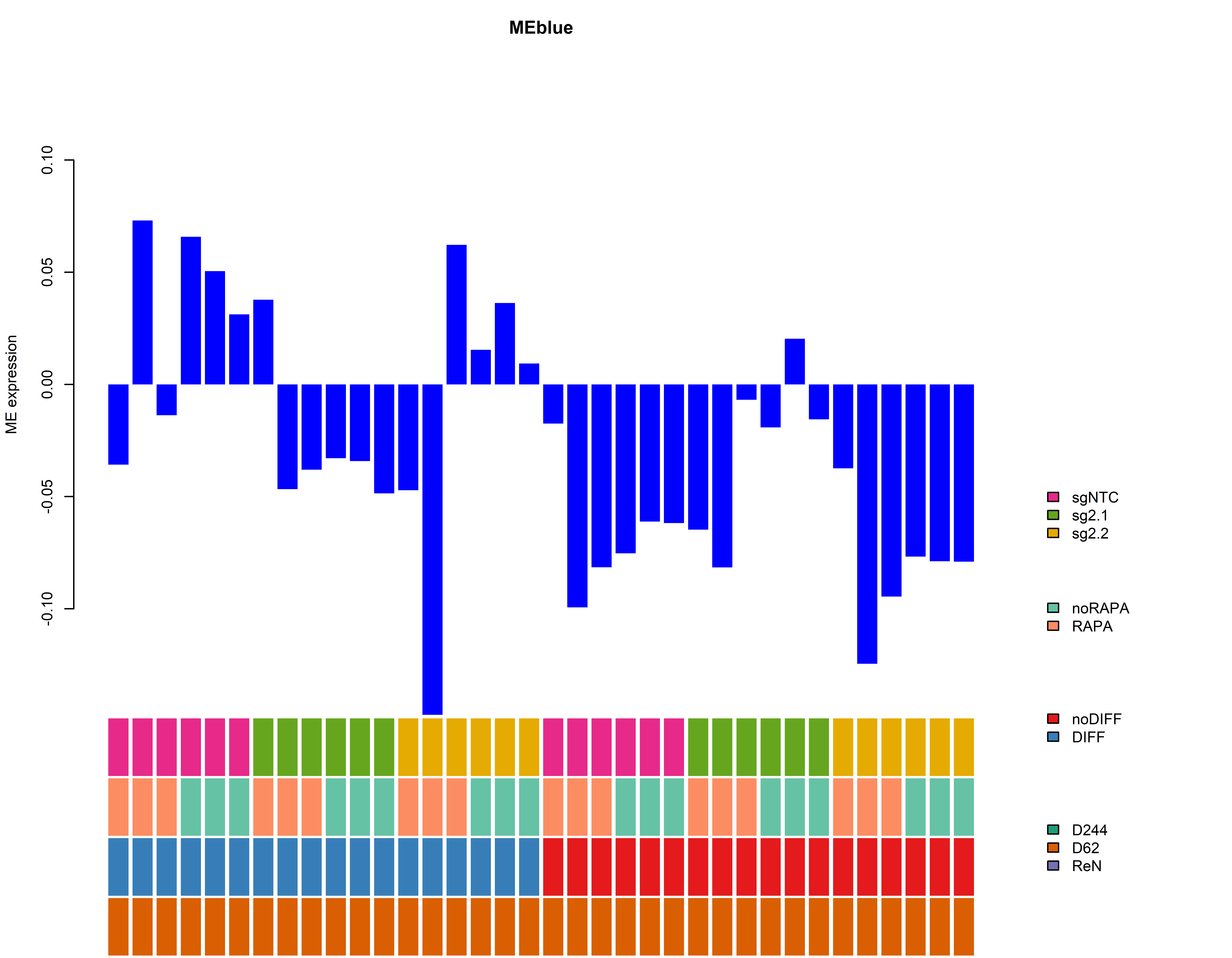
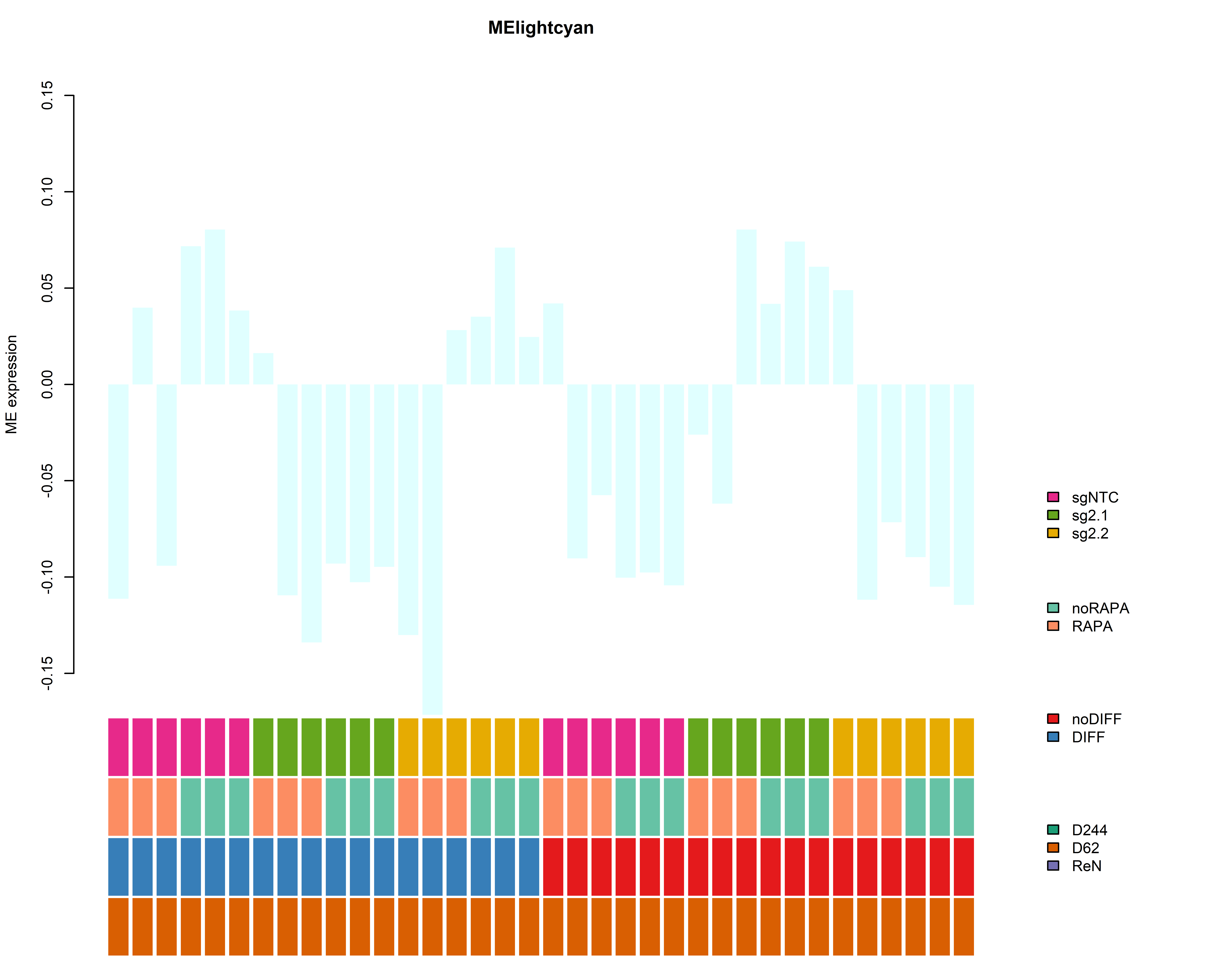
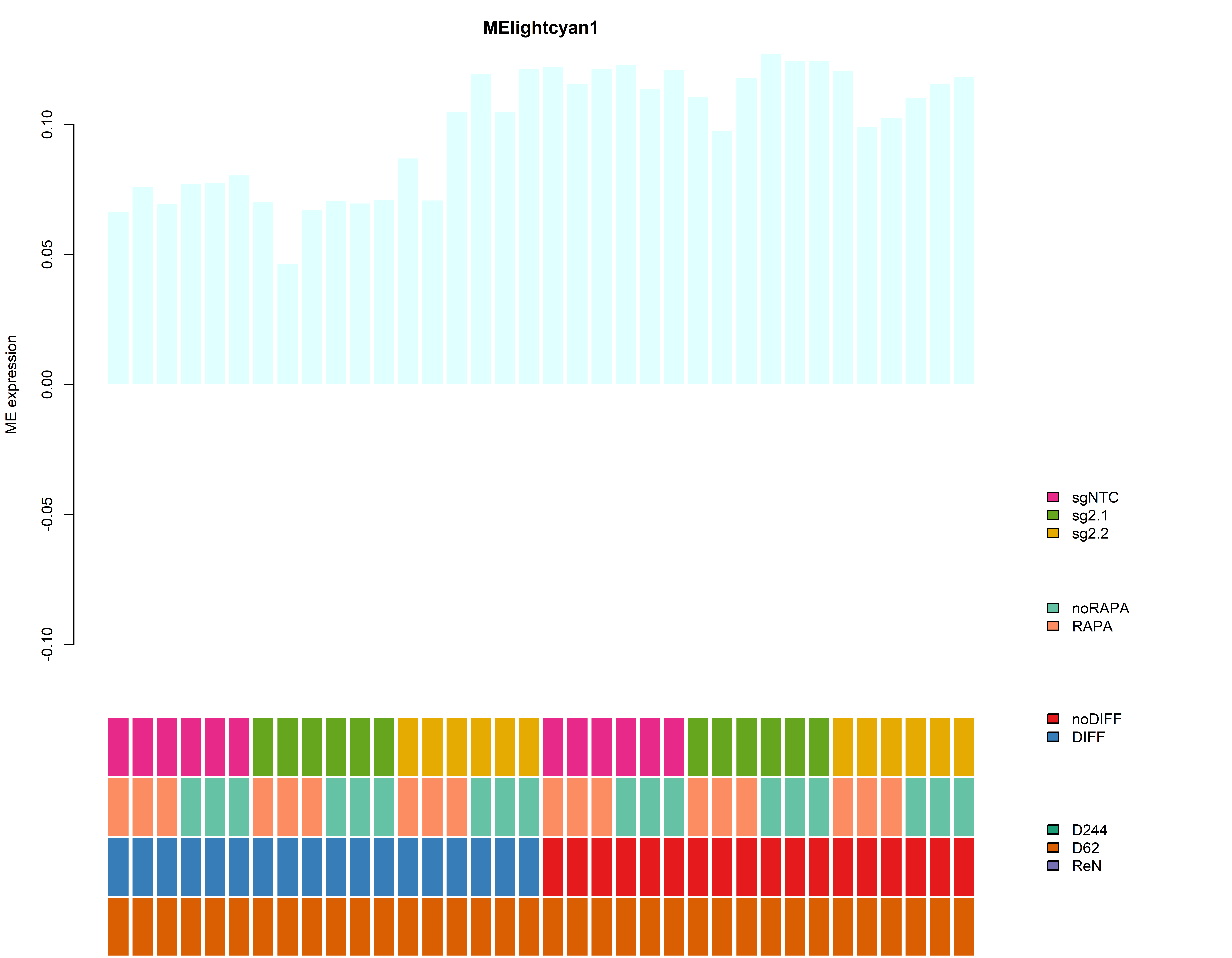
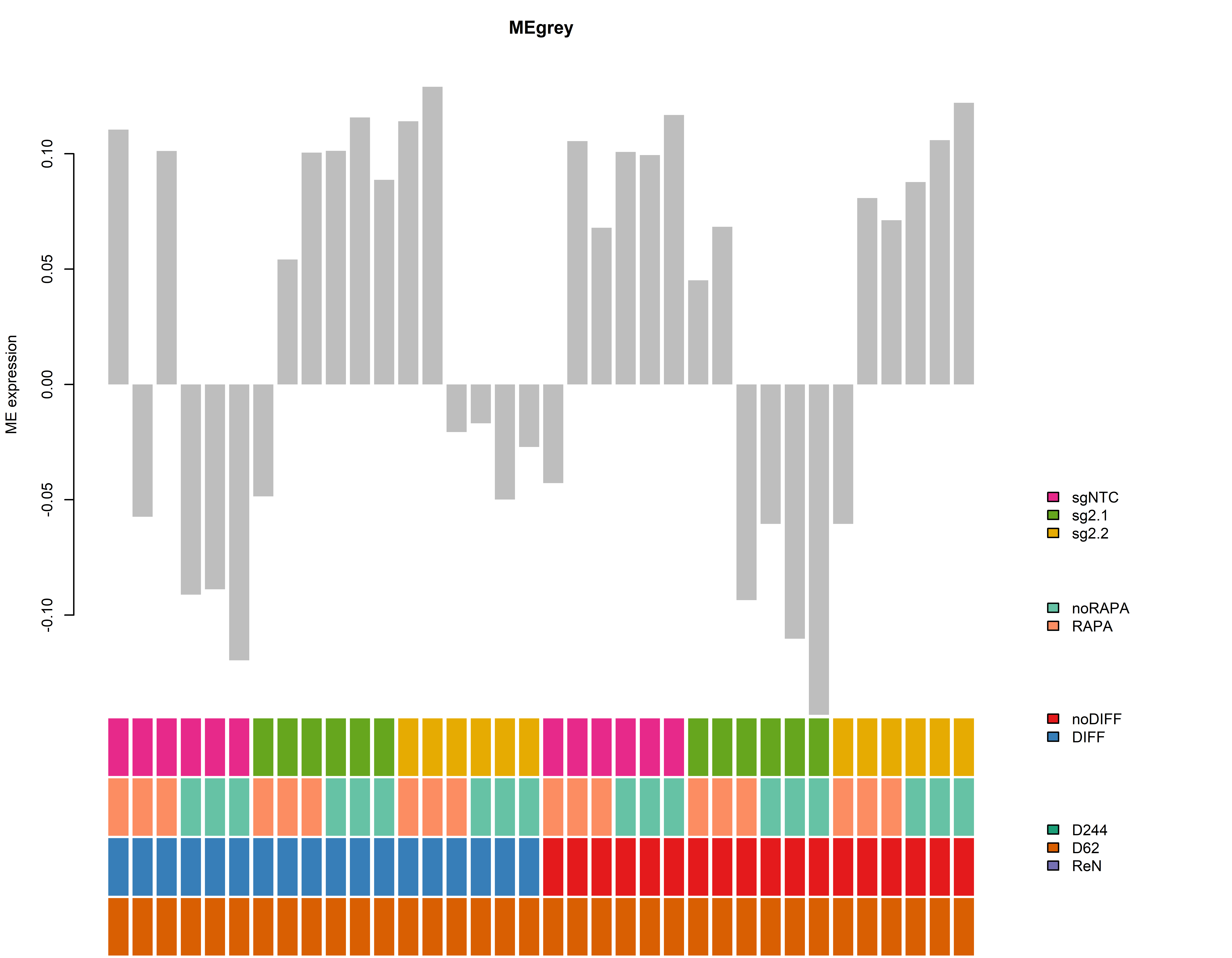
KO effect in noDIFF noRAPA
D62
getWGCNAoutputs(targetline = "D62",targetdiff = "noDIFF",targetrapa = "noRAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown -0.023239447 0.0127042451 -1.416357e-02 0.0195357308
2 MElightyellow -0.190849636 0.0051708995 1.717759e-03 1.0000000000
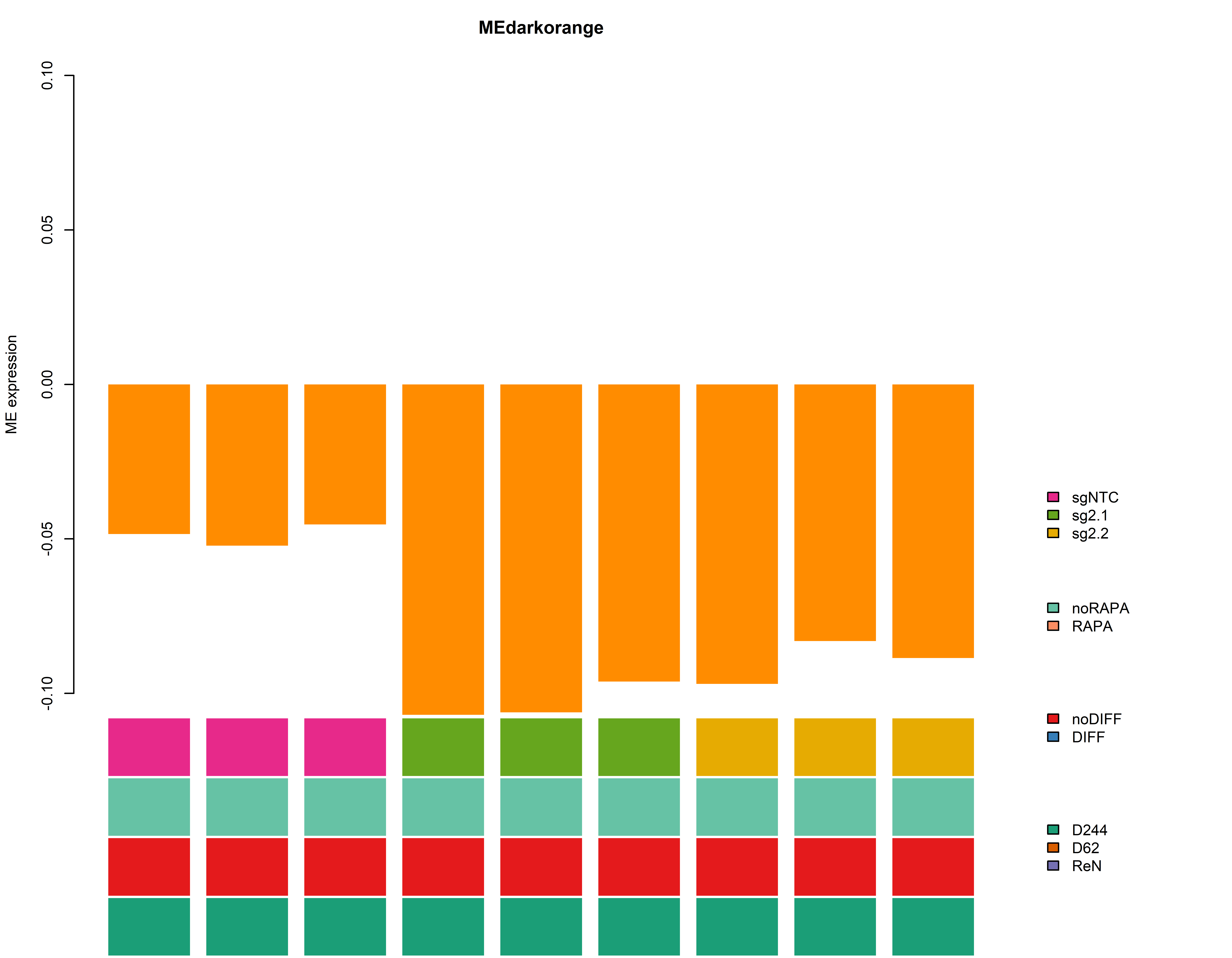
3 MEdarkorange 0.013098845 1.0000000000 1.118628e-02 0.1743265891
4 MEblue 0.061269207 0.2376017163 -1.213026e-02 1.0000000000
5 MElightcyan 0.159775907 0.0017435590 -2.306704e-03 1.0000000000
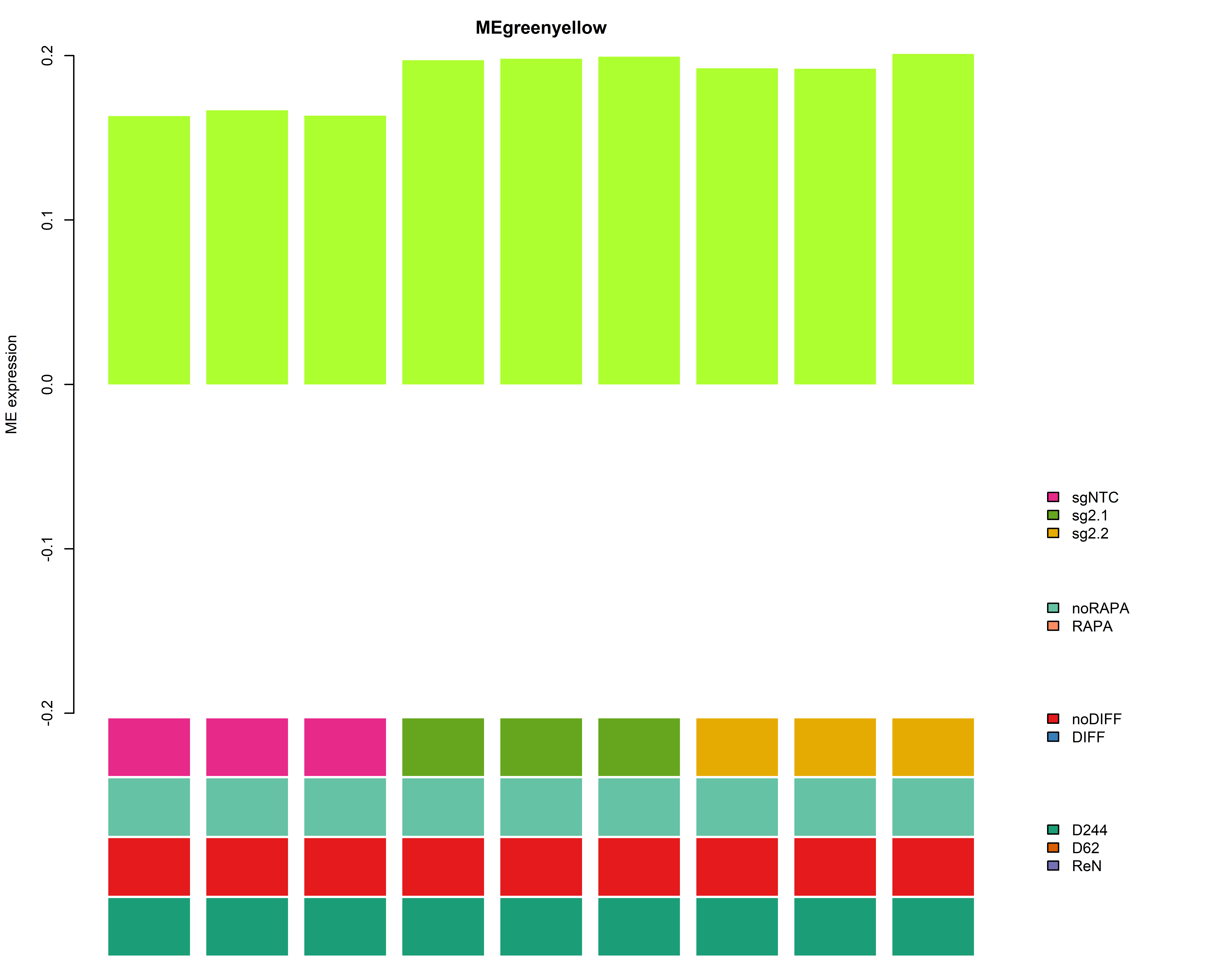
6 MEgreenyellow -0.010037776 1.0000000000 4.515721e-03 1.0000000000
7 MEplum1 -0.029930170 0.1325555586 -2.212925e-03 1.0000000000
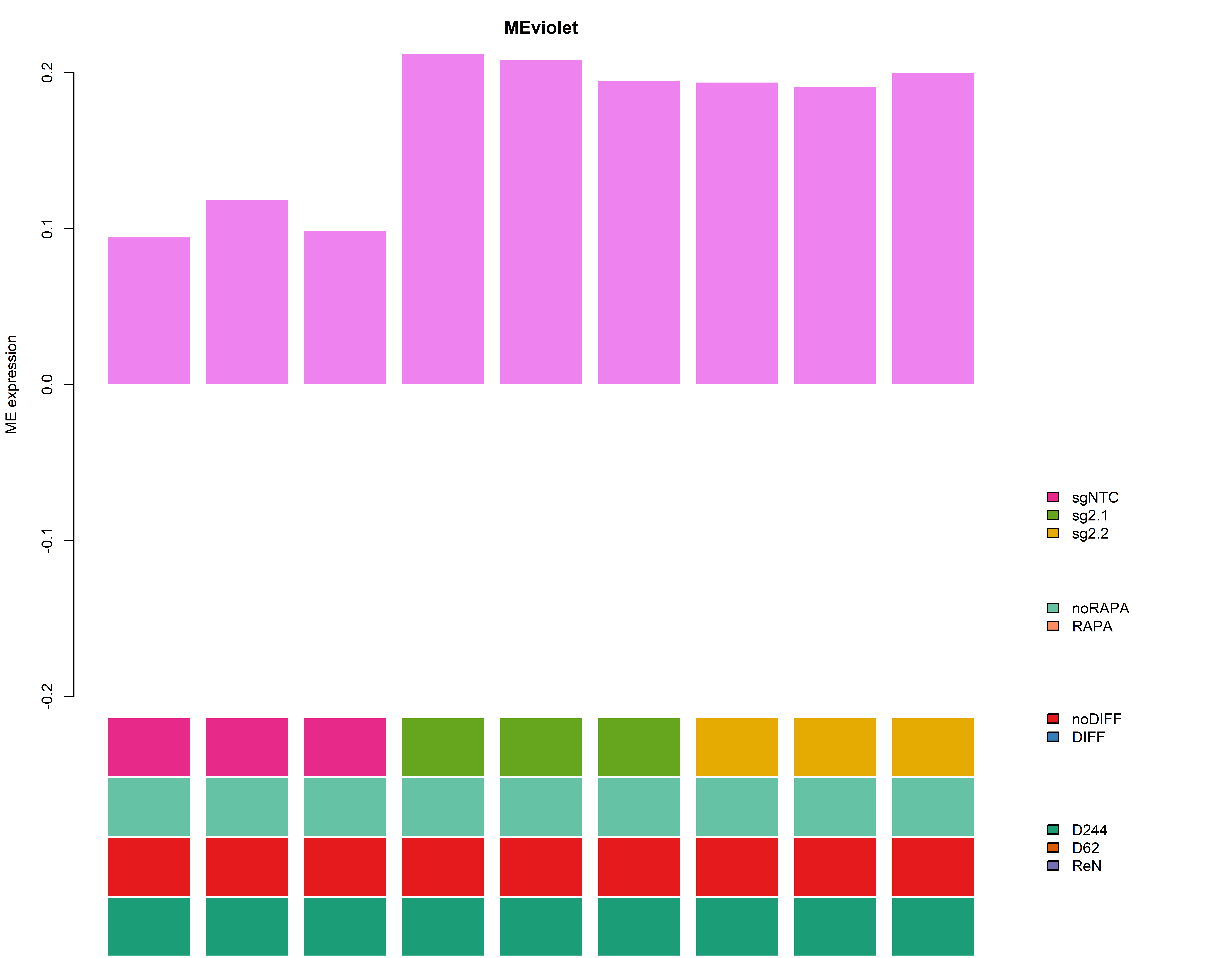
8 MEviolet -0.064701948 0.0434224926 8.744757e-05 1.0000000000
9 MEorange 0.009248861 1.0000000000 -3.642583e-03 1.0000000000
10 MEpaleturquoise 0.012171997 0.8082240748 9.228121e-03 0.3886141490
11 MEdarkmagenta 0.180897873 0.0054821728 5.301102e-03 1.0000000000
12 MEdarkturquoise 0.002797662 1.0000000000 8.977244e-03 1.0000000000
13 MEblack 0.018998851 0.6547218169 3.128855e-03 1.0000000000
14 MEmagenta 0.006818795 1.0000000000 1.604033e-02 0.0003212276
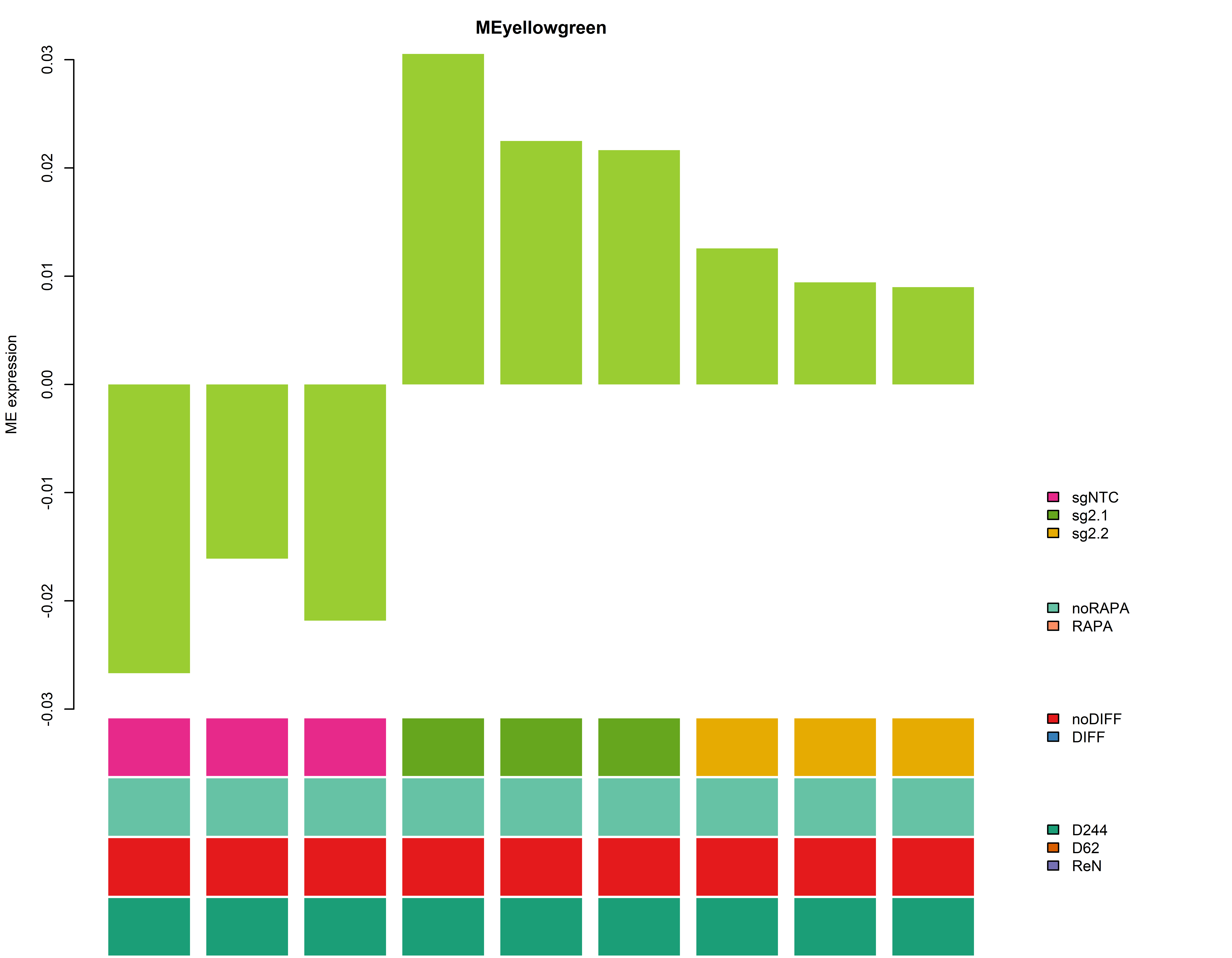
15 MEyellowgreen 0.026093249 0.3460571068 -6.413492e-03 1.0000000000
16 MEsaddlebrown -0.004224174 1.0000000000 -1.489529e-02 1.0000000000
17 MEdarkolivegreen -0.046907471 0.1505047323 -9.376309e-03 1.0000000000
18 MEmediumpurple3 -0.069992409 0.0054047620 -2.904031e-02 0.0619513098
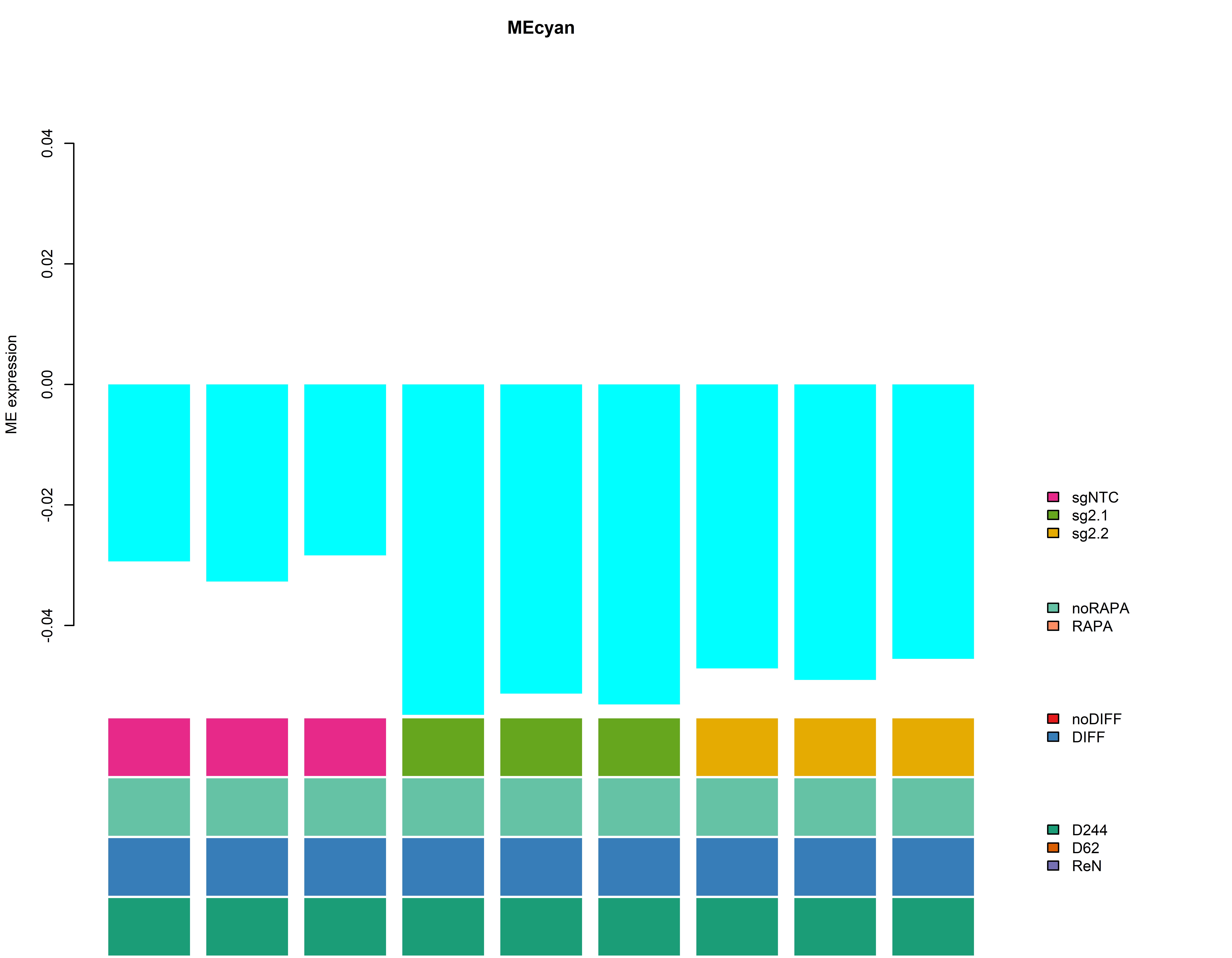
19 MEcyan 0.004429733 1.0000000000 -4.210276e-03 1.0000000000
20 MEpink -0.019163734 0.0054098938 -8.442063e-03 0.0971102878
21 MEdarkgreen -0.032965251 0.0001348384 -1.652470e-02 0.0054817088
22 MElightcyan1 0.006095436 1.0000000000 -4.520734e-03 1.0000000000
23 MEgrey -0.210324590 0.0239687907 -4.419418e-04 1.0000000000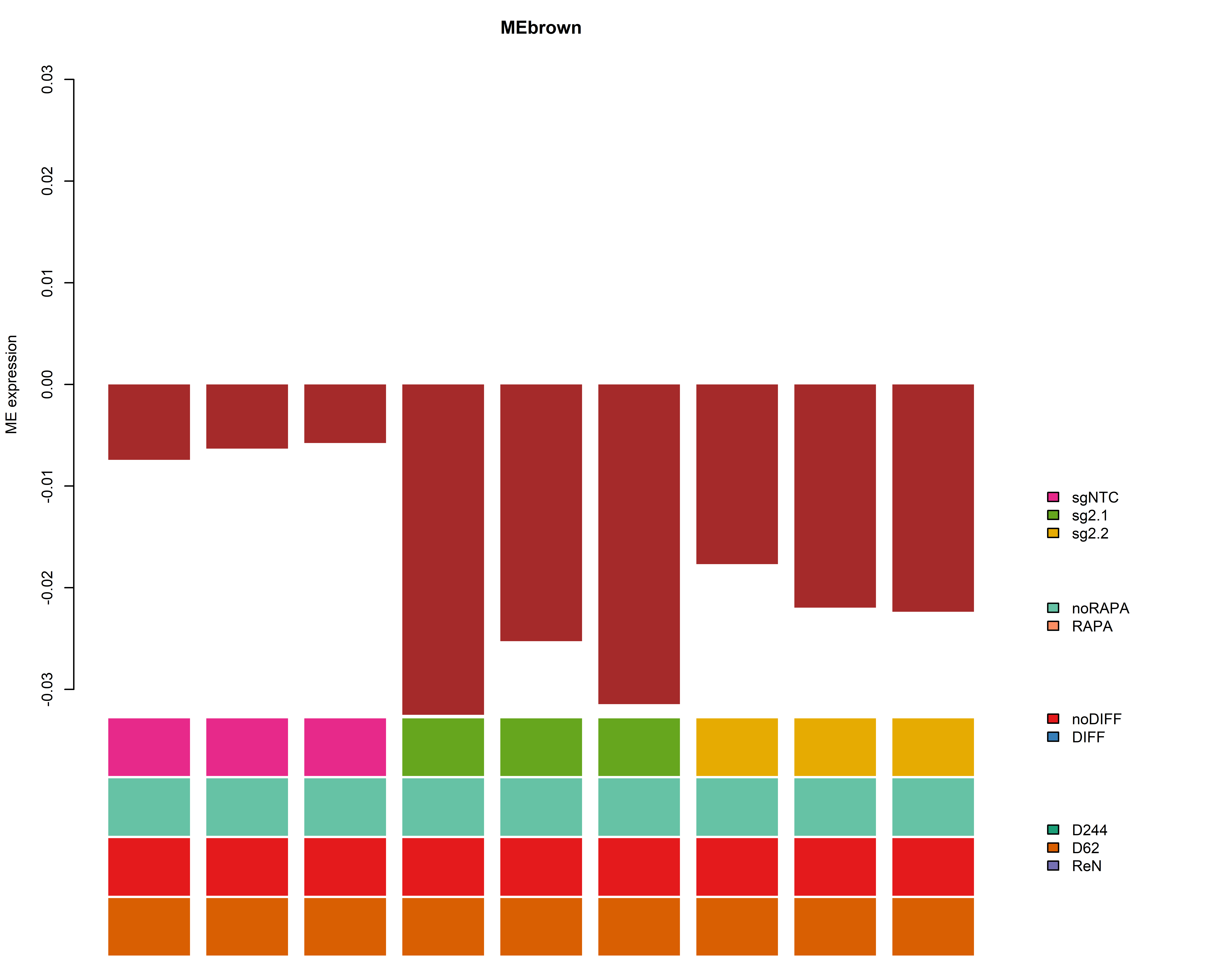
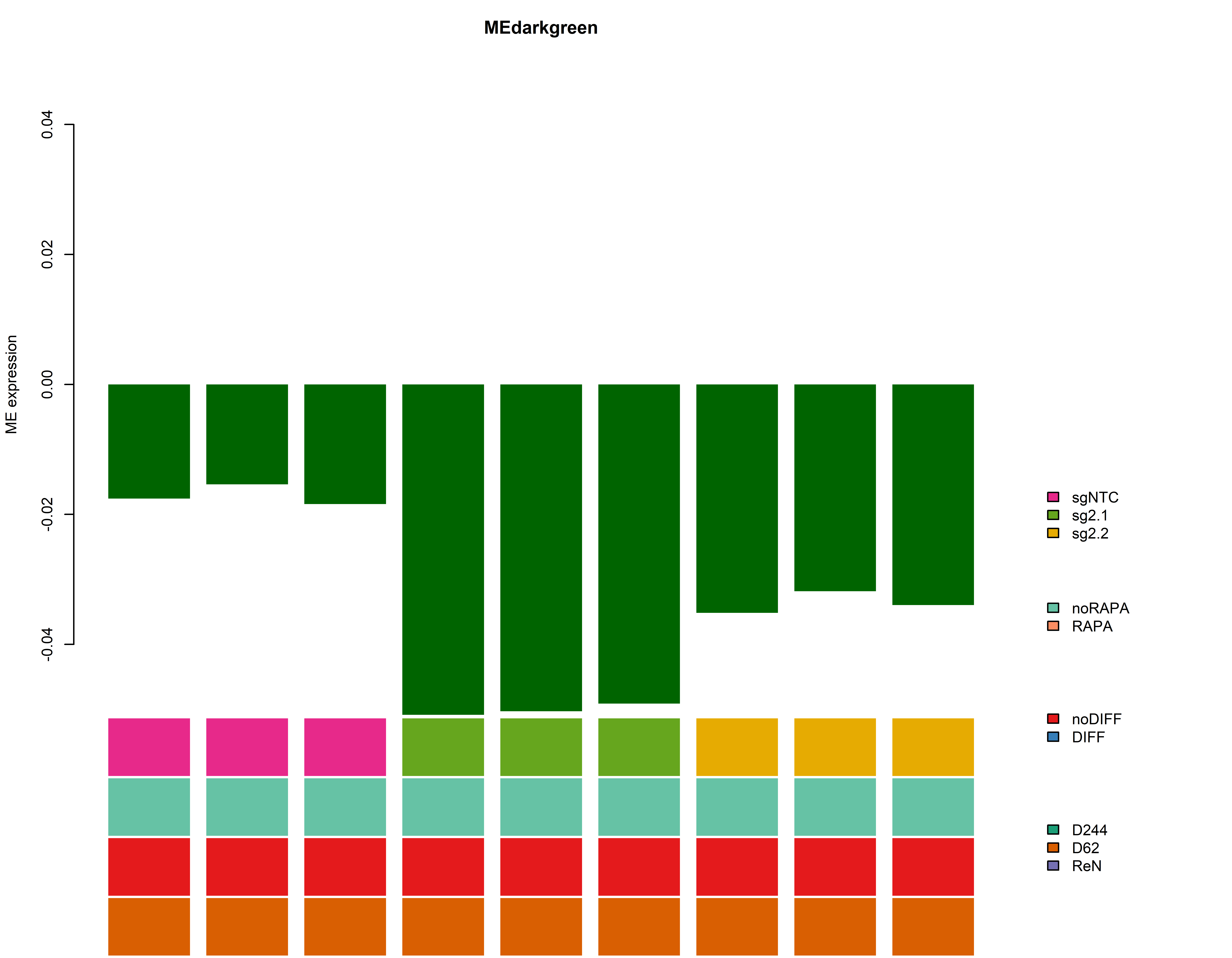
D244
getWGCNAoutputs(targetline = "D244",targetdiff = "noDIFF",targetrapa = "noRAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown -0.002798110 1.0000000000 -0.003667456 1.000000000
2 MElightyellow -0.090171132 0.0400259657 -0.100882319 0.024701422
3 MEdarkorange -0.054447726 0.0038552539 -0.040873767 0.018569865
4 MEblue -0.088020543 0.0443933411 -0.077424493 0.090797845
5 MElightcyan 0.033933023 1.0000000000 0.052732249 1.000000000
6 MEgreenyellow 0.033751421 0.0002892355 0.030607383 0.014873715
7 MEplum1 0.021858869 1.0000000000 0.023214232 0.529748907
8 MEviolet 0.101276808 0.0082691634 0.090883049 0.007229075
9 MEorange 0.027647426 0.0301692338 0.024827734 0.014309459
10 MEpaleturquoise 0.033195217 0.0056374654 0.025857485 0.010603862
11 MEdarkmagenta 0.093302052 0.0365229331 0.104053724 0.052808838
12 MEdarkturquoise -0.072153369 0.0094099720 -0.054223660 0.006054464
13 MEblack -0.035664747 0.0248218214 -0.033397136 0.015285573
14 MEmagenta -0.015682412 0.2773500700 -0.018100978 0.036266602
15 MEyellowgreen 0.046420012 0.0085269285 0.031860045 0.014154846
16 MEsaddlebrown -0.017528313 0.8360736974 -0.031713290 0.074874747
17 MEdarkolivegreen 0.029467812 0.1156677751 0.008279552 1.000000000
18 MEmediumpurple3 0.018830823 1.0000000000 0.003174960 1.000000000
19 MEcyan -0.024446225 0.0194821178 -0.026678650 0.158948336
20 MEpink 0.028654795 0.0693699842 0.013490964 0.338364078
21 MEdarkgreen -0.009195101 1.0000000000 -0.030010563 0.031187354
22 MElightcyan1 -0.010030353 0.4242406563 -0.018501115 0.705893451
23 MEgrey -0.116598929 0.1968748139 -0.152415136 0.015891148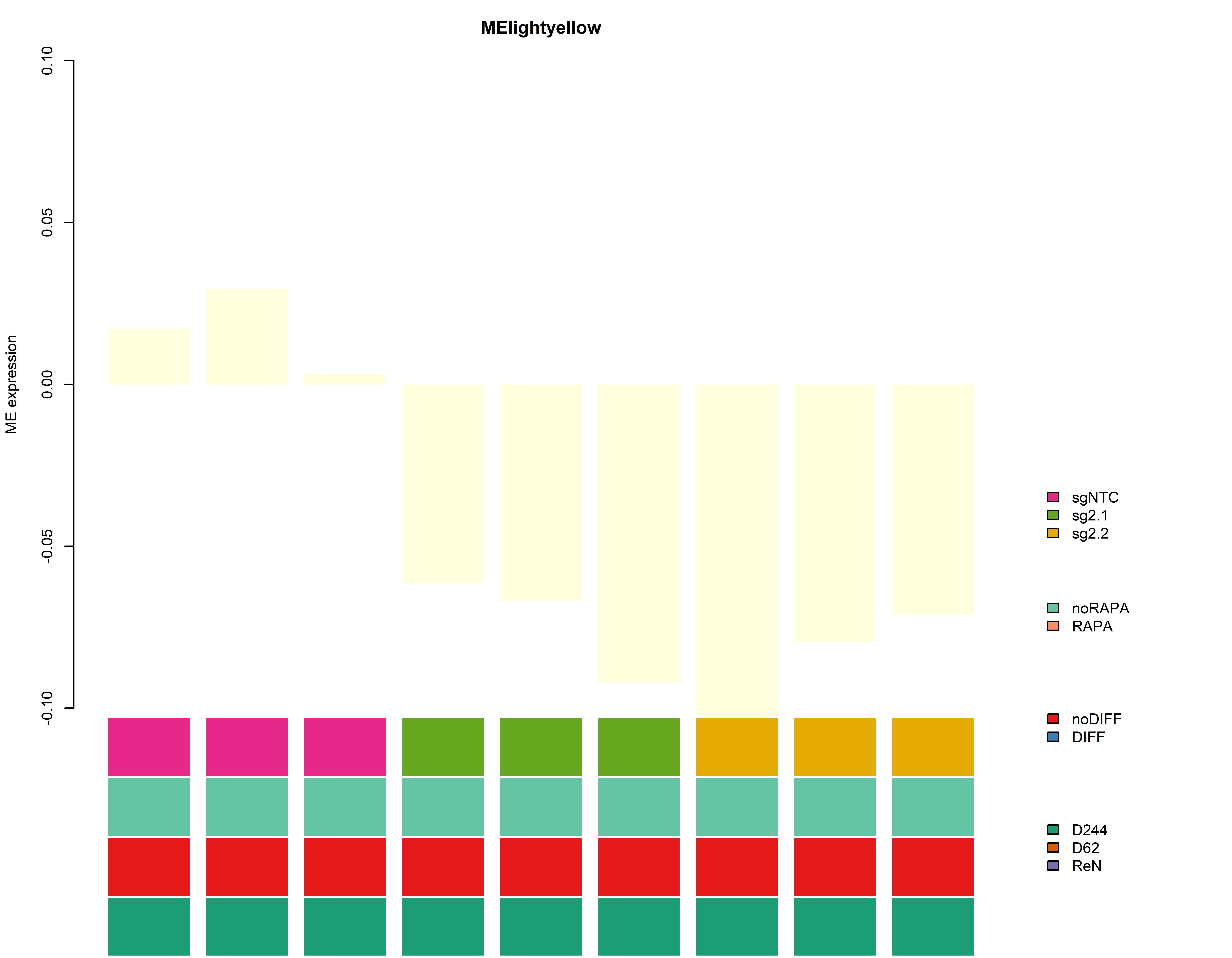
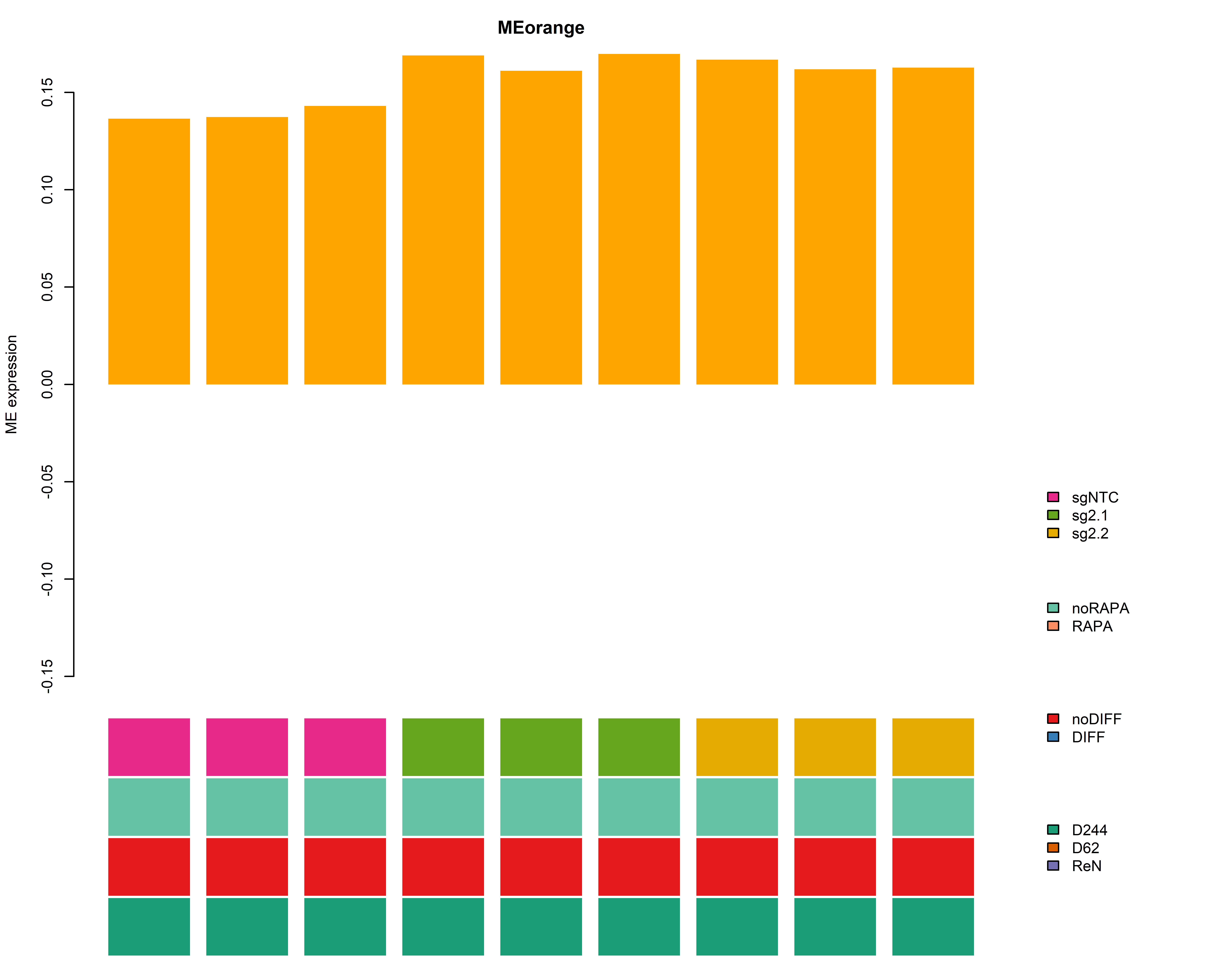
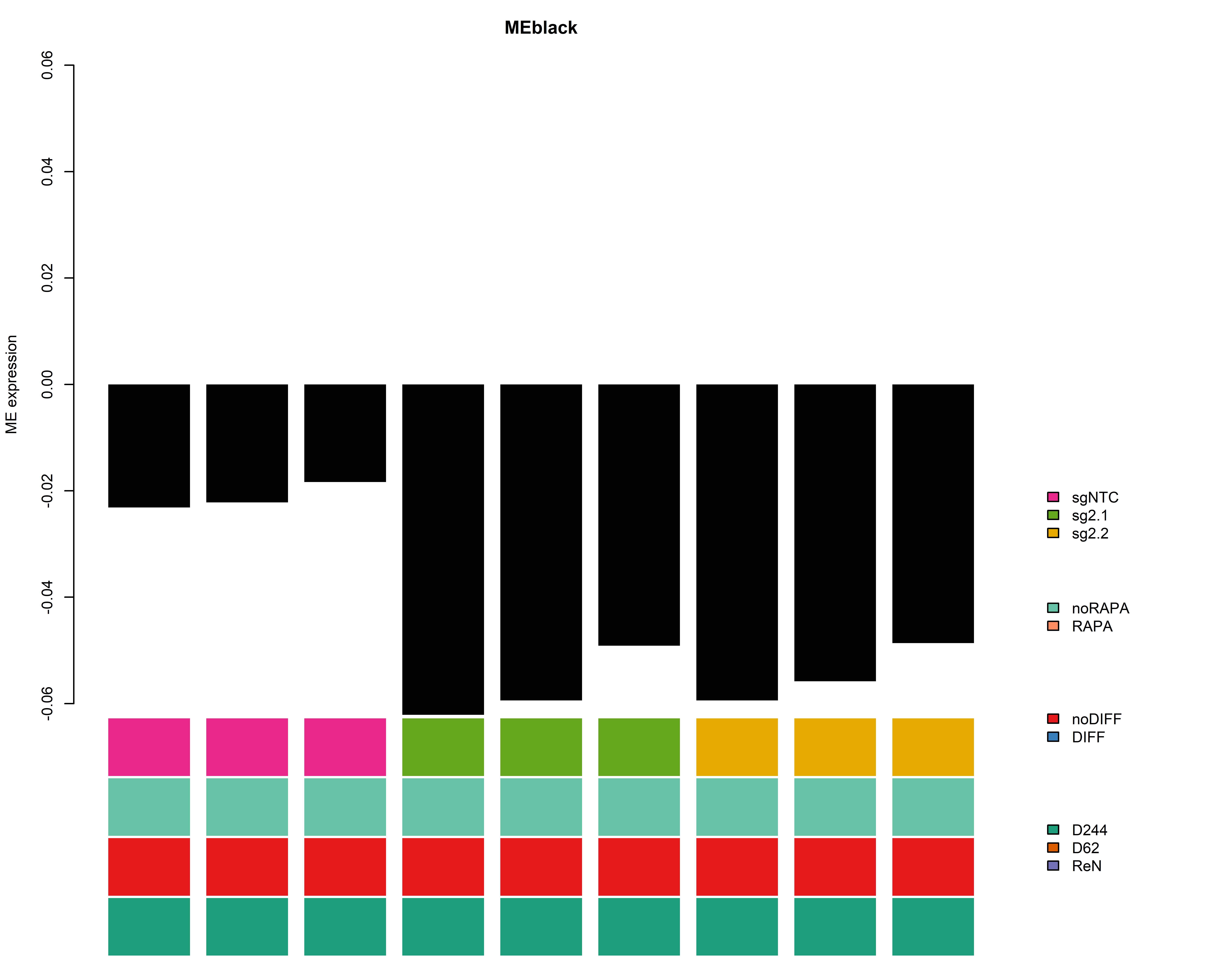
ReN
getWGCNAoutputs(targetline = "ReN",targetdiff = "noDIFF",targetrapa = "noRAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown -0.009230283 0.09867811 -0.0139673799 0.021269272
2 MElightyellow -0.131614381 1.00000000 0.0040438006 1.000000000
3 MEdarkorange 0.006790396 1.00000000 -0.0042673653 1.000000000
4 MEblue 0.052213365 1.00000000 -0.0069273753 1.000000000
5 MElightcyan 0.104914459 1.00000000 -0.0122726078 1.000000000
6 MEgreenyellow -0.006980709 1.00000000 0.0043993489 1.000000000
7 MEplum1 -0.014098948 1.00000000 -0.0011289227 1.000000000
8 MEviolet -0.050062184 1.00000000 0.0010860008 1.000000000
9 MEorange 0.001580815 1.00000000 -0.0008851102 1.000000000
10 MEpaleturquoise 0.001280697 1.00000000 0.0102401318 1.000000000
11 MEdarkmagenta 0.108548120 1.00000000 -0.0105899782 1.000000000
12 MEdarkturquoise -0.010648288 1.00000000 0.0165187198 0.038080500
13 MEblack 0.009804865 0.78354712 0.0084053628 1.000000000
14 MEmagenta 0.001567016 1.00000000 0.0092592207 1.000000000
15 MEyellowgreen 0.030233875 0.51087059 0.0179179545 0.106869489
16 MEsaddlebrown 0.004283410 1.00000000 -0.0012628414 1.000000000
17 MEdarkolivegreen -0.008841033 1.00000000 0.0184842190 0.508100661
18 MEmediumpurple3 -0.019937217 1.00000000 -0.0011249705 1.000000000
19 MEcyan -0.004084179 1.00000000 0.0053022336 1.000000000
20 MEpink -0.007816714 1.00000000 0.0157535973 0.000595805
21 MEdarkgreen -0.008364755 1.00000000 0.0059955085 1.000000000
22 MElightcyan1 0.002487710 1.00000000 -0.0005186695 1.000000000
23 MEgrey -0.138693708 1.00000000 0.0190969329 1.000000000KO effect in DIFF noRAPA
D62
getWGCNAoutputs(targetline = "D62",targetdiff = "DIFF",targetrapa = "noRAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown 0.0225035711 0.999652530 0.026886143 0.2903168939
2 MElightyellow 0.1982307054 0.002445134 0.033446904 1.0000000000
3 MEdarkorange -0.0108935090 0.457066267 -0.042166501 1.0000000000
4 MEblue -0.0877152196 0.033018090 -0.028822097 1.0000000000
5 MElightcyan -0.1602511004 0.006008662 -0.019866890 1.0000000000
6 MEgreenyellow 0.0047726143 1.000000000 -0.018076534 0.1194619707
7 MEplum1 0.0387468649 0.074222126 0.008910281 1.0000000000
8 MEviolet 0.0753197068 0.012739548 0.031349265 1.0000000000
9 MEorange -0.0048567240 1.000000000 0.009381652 1.0000000000
10 MEpaleturquoise 0.0088555991 1.000000000 0.029307968 1.0000000000
11 MEdarkmagenta -0.1942675943 0.001283925 -0.047402603 0.7553378741
12 MEdarkturquoise -0.0007406043 1.000000000 -0.005940129 1.0000000000
13 MEblack -0.0309135734 0.155314925 -0.049188680 0.1359655611
14 MEmagenta -0.0083998343 1.000000000 -0.036779851 0.1942232083
15 MEyellowgreen -0.0320484455 0.217164123 -0.017281860 0.6537591418
16 MEsaddlebrown -0.0453145633 0.012726864 -0.055485602 0.0004265345
17 MEdarkolivegreen 0.0204156645 0.579661232 -0.013303048 1.0000000000
18 MEmediumpurple3 0.0159838406 1.000000000 -0.021734359 1.0000000000
19 MEcyan 0.0014477274 1.000000000 0.044011556 0.1908950418
20 MEpink 0.0077098818 1.000000000 0.025553136 1.0000000000
21 MEdarkgreen 0.0243045485 0.951234615 0.041448961 0.3960896855
22 MElightcyan1 -0.0079963922 0.041096655 0.036790286 0.0508016060
23 MEgrey 0.2017575899 0.002051627 0.068567193 0.1806038989D244
getWGCNAoutputs(targetline = "D244",targetdiff = "DIFF",targetrapa = "noRAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown 0.003976069 1.00000000 -0.003148582 1.00000000
2 MElightyellow 0.115445385 1.00000000 -0.034282459 1.00000000
3 MEdarkorange 0.029036040 0.39936338 0.036229820 0.29597140
4 MEblue -0.080012533 1.00000000 0.004099908 1.00000000
5 MElightcyan -0.105986910 1.00000000 0.026367643 1.00000000
6 MEgreenyellow 0.020129449 0.38662854 0.001877617 1.00000000
7 MEplum1 0.022832164 1.00000000 0.003130540 1.00000000
8 MEviolet 0.043991398 1.00000000 -0.013092841 1.00000000
9 MEorange -0.021132506 1.00000000 -0.018158020 0.75472113
10 MEpaleturquoise -0.012957666 1.00000000 -0.020792373 1.00000000
11 MEdarkmagenta -0.090252741 1.00000000 0.046361629 1.00000000
12 MEdarkturquoise 0.016727215 1.00000000 0.009567142 1.00000000
13 MEblack 0.011964705 1.00000000 0.026135472 1.00000000
14 MEmagenta 0.030897859 0.01757824 0.029801716 0.01592759
15 MEyellowgreen 0.004225403 1.00000000 0.027341795 1.00000000
16 MEsaddlebrown -0.018298002 1.00000000 -0.006282741 1.00000000
17 MEdarkolivegreen 0.022580382 1.00000000 0.004465159 1.00000000
18 MEmediumpurple3 0.014347796 1.00000000 -0.012013171 1.00000000
19 MEcyan -0.022937198 0.00363728 -0.017085518 0.01151321
20 MEpink -0.005843991 1.00000000 -0.018110346 1.00000000
21 MEdarkgreen -0.011226485 1.00000000 -0.012349967 1.00000000
22 MElightcyan1 -0.038089945 0.02459818 -0.022496146 0.21733508
23 MEgrey 0.133533017 1.00000000 0.006997965 1.00000000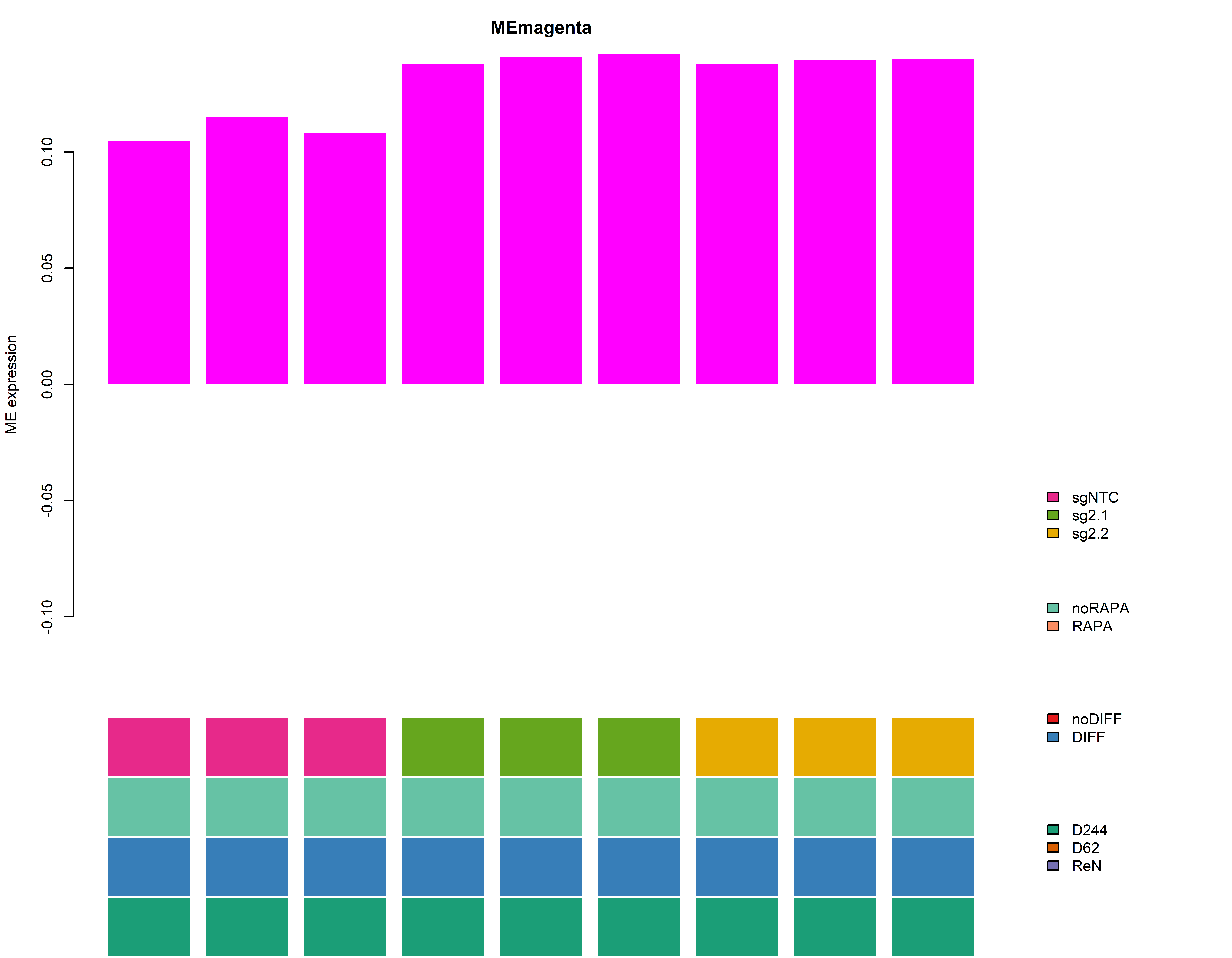
ReN
getWGCNAoutputs(targetline = "ReN",targetdiff = "DIFF",targetrapa = "noRAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown 0.012458755 0.634257663 -0.0040062852 1.00000000
2 MElightyellow 0.046259405 0.010973954 -0.0833120638 1.00000000
3 MEdarkorange 0.003230998 1.000000000 0.0124520954 1.00000000
4 MEblue -0.011071284 1.000000000 0.0571710808 1.00000000
5 MElightcyan -0.021890355 0.670494938 0.1063186139 0.46065169
6 MEgreenyellow 0.015585385 0.185980950 0.0060606535 1.00000000
7 MEplum1 0.046397614 0.075679713 0.0124349480 1.00000000
8 MEviolet 0.019299716 0.993914199 -0.0189530083 1.00000000
9 MEorange 0.011000555 1.000000000 0.0161137538 0.79058022
10 MEpaleturquoise 0.003288921 1.000000000 -0.0109531442 0.56121694
11 MEdarkmagenta -0.022412520 1.000000000 0.0984067722 1.00000000
12 MEdarkturquoise -0.008821813 1.000000000 -0.0232555755 1.00000000
13 MEblack -0.013850997 0.664602362 -0.0005153248 1.00000000
14 MEmagenta 0.015150659 0.547249302 0.0019063586 1.00000000
15 MEyellowgreen -0.018758243 0.311989085 -0.0052474413 1.00000000
16 MEsaddlebrown -0.031061316 0.002377907 -0.0170271245 0.97673308
17 MEdarkolivegreen 0.003056362 1.000000000 -0.0139317747 1.00000000
18 MEmediumpurple3 -0.002105930 1.000000000 -0.0130717728 1.00000000
19 MEcyan -0.020128373 0.361574572 -0.0242590606 0.09416076
20 MEpink -0.009631877 1.000000000 -0.0219500441 0.63901504
21 MEdarkgreen -0.013919546 0.976670791 -0.0240111468 0.38848306
22 MElightcyan1 -0.017344461 1.000000000 -0.0169127515 1.00000000
23 MEgrey 0.018743598 1.000000000 -0.1466959778 1.00000000KO effect in noDIFF RAPA
D62
getWGCNAoutputs(targetline = "D62",targetdiff = "noDIFF",targetrapa = "RAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown -0.0117656325 1.0000000 -3.965234e-03 1.0000000
2 MElightyellow -0.0274081344 1.0000000 1.379870e-02 1.0000000
3 MEdarkorange 0.0116804550 1.0000000 3.681162e-03 1.0000000
4 MEblue 0.0150719308 1.0000000 -1.935969e-02 1.0000000
5 MElightcyan 0.0327328781 1.0000000 -9.540489e-03 1.0000000
6 MEgreenyellow -0.0036025778 1.0000000 1.455638e-05 1.0000000
7 MEplum1 -0.0173449164 1.0000000 5.289826e-04 1.0000000
8 MEviolet -0.0159293134 1.0000000 1.486866e-02 1.0000000
9 MEorange -0.0063638273 1.0000000 -2.109614e-03 1.0000000
10 MEpaleturquoise -0.0058321690 1.0000000 -3.201509e-03 1.0000000
11 MEdarkmagenta 0.0349148946 1.0000000 -1.584140e-02 1.0000000
12 MEdarkturquoise 0.0004216442 1.0000000 -3.178335e-03 1.0000000
13 MEblack 0.0074168821 1.0000000 -3.161364e-03 1.0000000
14 MEmagenta 0.0116513081 1.0000000 8.337287e-04 1.0000000
15 MEyellowgreen 0.0115432617 1.0000000 -4.830650e-03 1.0000000
16 MEsaddlebrown -0.0047834406 1.0000000 -8.579157e-03 1.0000000
17 MEdarkolivegreen -0.0234906263 1.0000000 -6.249704e-03 1.0000000
18 MEmediumpurple3 -0.0227320066 1.0000000 -9.378458e-03 1.0000000
19 MEcyan -0.0078941624 1.0000000 -9.623402e-03 1.0000000
20 MEpink -0.0173857310 0.3861186 -9.907869e-03 1.0000000
21 MEdarkgreen -0.0235550161 0.8894477 -1.222265e-02 0.1847832
22 MElightcyan1 -0.0109625697 1.0000000 -1.225391e-02 1.0000000
23 MEgrey -0.0368871049 1.0000000 -1.302151e-02 1.0000000D244
getWGCNAoutputs(targetline = "D244",targetdiff = "noDIFF",targetrapa = "RAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown 0.0060974279 4.661505e-01 5.717870e-03 6.398163e-02
2 MElightyellow -0.0561255532 1.000000e+00 -9.289121e-02 1.000000e+00
3 MEdarkorange -0.0579633620 7.128329e-05 -4.775148e-02 3.182472e-05
4 MEblue -0.1145844851 2.050760e-01 -8.379959e-02 5.587550e-01
5 MElightcyan 0.0007179999 1.000000e+00 5.435302e-02 1.000000e+00
6 MEgreenyellow 0.0302255551 6.526536e-02 1.919281e-02 1.000000e+00
7 MEplum1 0.0161065787 1.000000e+00 2.156455e-02 1.000000e+00
8 MEviolet 0.1114343571 5.863009e-02 6.588085e-02 3.400159e-01
9 MEorange 0.0308379484 5.118646e-03 2.931812e-02 2.533413e-02
10 MEpaleturquoise 0.0378145958 3.986492e-02 2.138194e-02 1.672942e-01
11 MEdarkmagenta 0.0579085517 1.000000e+00 9.021368e-02 1.000000e+00
12 MEdarkturquoise -0.0597221400 1.436239e-03 -4.264068e-02 3.467685e-01
13 MEblack -0.0446919444 1.763903e-02 -3.791056e-02 6.263752e-02
14 MEmagenta -0.0229421309 8.976510e-02 -2.245567e-02 2.416616e-01
15 MEyellowgreen 0.0407198466 1.170128e-01 4.544839e-02 3.259647e-02
16 MEsaddlebrown -0.0285972867 3.791456e-01 -4.072937e-02 1.051042e-01
17 MEdarkolivegreen 0.0491562287 3.561212e-01 2.722072e-03 1.000000e+00
18 MEmediumpurple3 0.0421795899 1.821074e-02 2.619224e-03 1.000000e+00
19 MEcyan -0.0237708493 7.620669e-02 -2.658402e-02 2.502805e-01
20 MEpink 0.0319605915 1.203874e-02 1.485947e-02 1.293239e-01
21 MEdarkgreen -0.0014834010 1.000000e+00 -2.084226e-02 1.059937e-01
22 MElightcyan1 -0.0083398613 1.000000e+00 3.404348e-05 1.000000e+00
23 MEgrey -0.1099971411 9.289151e-01 -1.447057e-01 3.247767e-01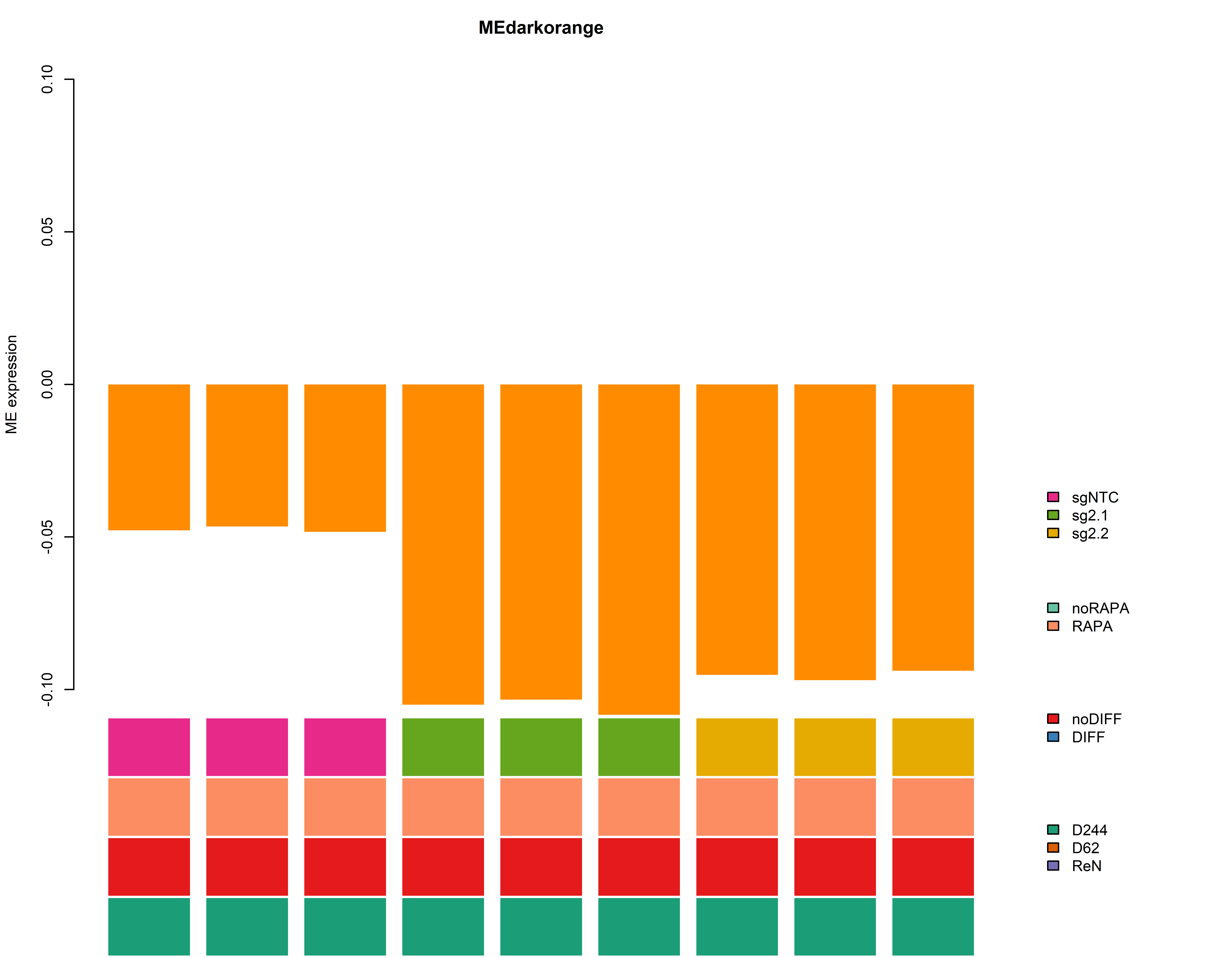
ReN
getWGCNAoutputs(targetline = "ReN",targetdiff = "noDIFF",targetrapa = "RAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown -0.0090871925 1.0000000 -0.011111868 0.792985988
2 MElightyellow -0.0850335347 1.0000000 0.047555696 1.000000000
3 MEdarkorange -0.0119233605 1.0000000 -0.017606700 0.051331278
4 MEblue 0.0320349984 1.0000000 -0.026132195 1.000000000
5 MElightcyan 0.0804284195 1.0000000 -0.047271888 1.000000000
6 MEgreenyellow 0.0012294398 1.0000000 0.009222012 1.000000000
7 MEplum1 -0.0120700372 1.0000000 0.011037606 1.000000000
8 MEviolet -0.0265077982 1.0000000 0.021942972 1.000000000
9 MEorange 0.0055902925 1.0000000 0.008040177 1.000000000
10 MEpaleturquoise 0.0079552382 1.0000000 0.027098905 0.001292973
11 MEdarkmagenta 0.0661012671 1.0000000 -0.043173764 1.000000000
12 MEdarkturquoise 0.0008276224 1.0000000 0.010699929 1.000000000
13 MEblack 0.0077635068 1.0000000 -0.002363520 1.000000000
14 MEmagenta 0.0040205209 1.0000000 0.009003270 1.000000000
15 MEyellowgreen 0.0225268657 0.5497912 0.012395819 1.000000000
16 MEsaddlebrown 0.0030681824 1.0000000 -0.001658637 1.000000000
17 MEdarkolivegreen 0.0026763486 1.0000000 0.014893367 1.000000000
18 MEmediumpurple3 -0.0156812163 1.0000000 0.008185163 1.000000000
19 MEcyan 0.0033332148 1.0000000 0.008418724 1.000000000
20 MEpink 0.0152837840 0.8998573 0.031665436 0.013021131
21 MEdarkgreen 0.0079320212 1.0000000 0.011746614 1.000000000
22 MElightcyan1 0.0154933054 1.0000000 0.010454147 1.000000000
23 MEgrey -0.1018878801 1.0000000 0.023124576 1.000000000KO effect in DIFF RAPA
D62
getWGCNAoutputs(targetline = "D62",targetdiff = "DIFF",targetrapa = "RAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown 0.013887384 0.17453700 0.0136135880 1.000000000
2 MElightyellow 0.023820286 1.00000000 0.0014751944 1.000000000
3 MEdarkorange -0.003499460 1.00000000 -0.0231024728 1.000000000
4 MEblue -0.023527791 1.00000000 -0.0519286668 1.000000000
5 MElightcyan -0.020543251 1.00000000 -0.0359458281 1.000000000
6 MEgreenyellow -0.008773074 1.00000000 -0.0092754110 1.000000000
7 MEplum1 0.014433339 1.00000000 -0.0003704416 1.000000000
8 MEviolet 0.027817441 1.00000000 0.0556621524 1.000000000
9 MEorange -0.015272415 0.31224335 -0.0125009070 0.201899549
10 MEpaleturquoise -0.005178901 1.00000000 0.0168124882 1.000000000
11 MEdarkmagenta -0.026307862 1.00000000 -0.0228079695 1.000000000
12 MEdarkturquoise -0.007963201 1.00000000 -0.0106900312 1.000000000
13 MEblack -0.012575234 1.00000000 -0.0267312211 1.000000000
14 MEmagenta -0.022752893 0.02819868 -0.0298114138 0.005325429
15 MEyellowgreen 0.006086539 1.00000000 0.0085618428 1.000000000
16 MEsaddlebrown -0.019708690 1.00000000 -0.0307426934 1.000000000
17 MEdarkolivegreen 0.022663715 1.00000000 0.0150837217 1.000000000
18 MEmediumpurple3 0.019640624 1.00000000 0.0124781773 1.000000000
19 MEcyan -0.010004609 1.00000000 0.0114898220 0.496968199
20 MEpink 0.005663475 1.00000000 0.0273611313 1.000000000
21 MEdarkgreen 0.033816222 0.17047811 0.0529412838 0.024631909
22 MElightcyan1 -0.009412863 1.00000000 0.0168589735 1.000000000
23 MEgrey -0.016082514 1.00000000 0.0227220527 1.000000000D244
getWGCNAoutputs(targetline = "D244",targetdiff = "DIFF",targetrapa = "RAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown 0.0128465458 0.04190437 0.001914271 1.000000000
2 MElightyellow 0.1643135893 0.02437463 -0.042039712 1.000000000
3 MEdarkorange 0.0084506899 1.00000000 0.022544240 0.209499522
4 MEblue -0.1016329669 0.03634386 0.004812568 1.000000000
5 MElightcyan -0.1480205503 0.03854498 0.027502256 1.000000000
6 MEgreenyellow 0.0124251420 0.71772804 -0.003813493 1.000000000
7 MEplum1 0.0183597335 1.00000000 -0.001360435 1.000000000
8 MEviolet 0.0531857173 0.31339970 -0.027682392 1.000000000
9 MEorange -0.0196693770 0.07740634 -0.008998253 1.000000000
10 MEpaleturquoise -0.0021294830 1.00000000 -0.010137578 1.000000000
11 MEdarkmagenta -0.1446088710 0.01302601 0.037591287 1.000000000
12 MEdarkturquoise 0.0167069754 0.52885831 0.011375506 1.000000000
13 MEblack -0.0107987453 1.00000000 0.018223317 0.813295793
14 MEmagenta 0.0084556604 1.00000000 0.018670826 0.348747119
15 MEyellowgreen 0.0015653762 1.00000000 0.028725222 0.282027523
16 MEsaddlebrown -0.0197081211 0.93813000 -0.007947650 0.900010599
17 MEdarkolivegreen 0.0224396308 0.23006103 -0.011662325 1.000000000
18 MEmediumpurple3 -0.0007637968 1.00000000 -0.040247009 0.006611766
19 MEcyan -0.0235301806 0.02858187 -0.011250277 0.278267783
20 MEpink -0.0010790312 1.00000000 -0.013717300 0.491084865
21 MEdarkgreen -0.0035777057 1.00000000 -0.017299169 0.183621568
22 MElightcyan1 -0.0208459185 0.75922869 -0.013163636 1.000000000
23 MEgrey 0.1166265945 0.15619231 -0.020145055 1.000000000ReN
getWGCNAoutputs(targetline = "ReN",targetdiff = "DIFF",targetrapa = "RAPA") Module beta_2.1 bonferroni_2.1 beta_2.2 bonferroni_2.2
1 MEbrown 0.0001217972 1.0000000000 -0.0041060278 1.00000000
2 MElightyellow 0.0083568224 1.0000000000 -0.0311110392 1.00000000
3 MEdarkorange -0.0500618156 0.1191677862 -0.0247297482 1.00000000
4 MEblue -0.0268251917 1.0000000000 0.0210699282 1.00000000
5 MElightcyan -0.0280486182 1.0000000000 0.0428903689 1.00000000
6 MEgreenyellow 0.0124775663 1.0000000000 0.0088206527 1.00000000
7 MEplum1 0.0178124544 1.0000000000 0.0010244916 1.00000000
8 MEviolet 0.0382540586 1.0000000000 0.0144621737 1.00000000
9 MEorange 0.0288011585 0.1866954123 0.0149657431 0.46665071
10 MEpaleturquoise 0.0590600342 0.0000295626 0.0365856044 0.06650126
11 MEdarkmagenta 0.0071301350 1.0000000000 0.0692839398 1.00000000
12 MEdarkturquoise 0.0164730419 1.0000000000 0.0199323078 1.00000000
13 MEblack -0.0289122219 0.4684872877 -0.0083590351 1.00000000
14 MEmagenta -0.0159441485 1.0000000000 -0.0021096027 1.00000000
15 MEyellowgreen 0.0079389079 1.0000000000 0.0171041277 1.00000000
16 MEsaddlebrown -0.0289063972 0.2116105363 -0.0153159662 0.05172409
17 MEdarkolivegreen 0.0017994209 1.0000000000 0.0044915223 1.00000000
18 MEmediumpurple3 -0.0054450917 1.0000000000 -0.0222543712 1.00000000
19 MEcyan 0.0498334965 0.3114563737 0.0441161786 0.44645182
20 MEpink 0.0463666938 0.1559934224 0.0247604406 1.00000000
21 MEdarkgreen -0.0058971338 1.0000000000 -0.0001507283 1.00000000
22 MElightcyan1 0.0293156997 1.0000000000 0.0188974124 1.00000000
23 MEgrey 0.1734879045 1.0000000000 0.1021618845 1.00000000gene_univers = rownames(ddsMat)
Genes_of_interset = split(rownames(ddsMat), mcols(ddsMat)$cluster)
gostres = getGOresults(Genes_of_interset, gene_univers)
toptab = gostres$result
write.xlsx2(toptab, file = paste0(output,"/GOresWGCNA.xlsx"), sheetName = "GO_enrichment")
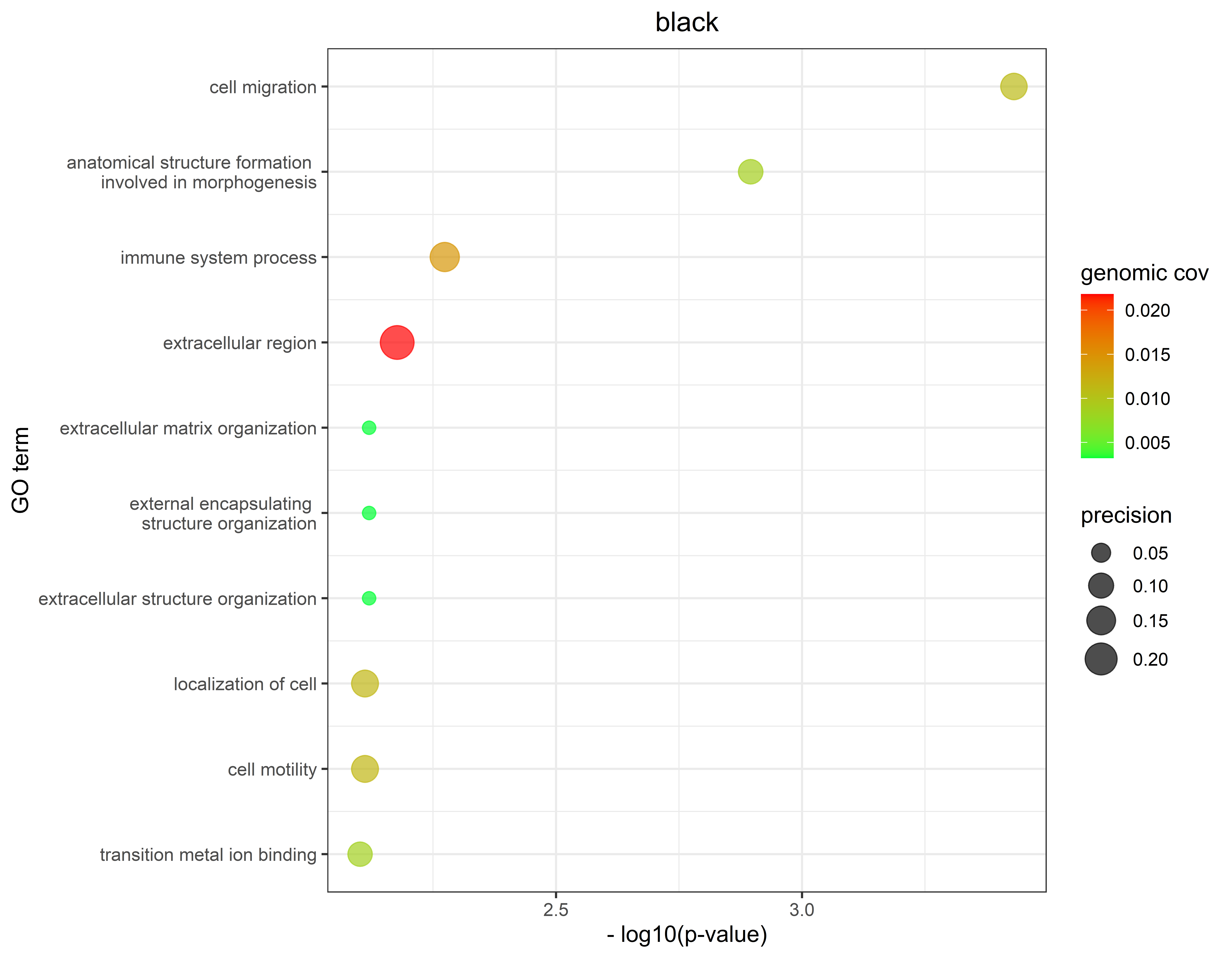
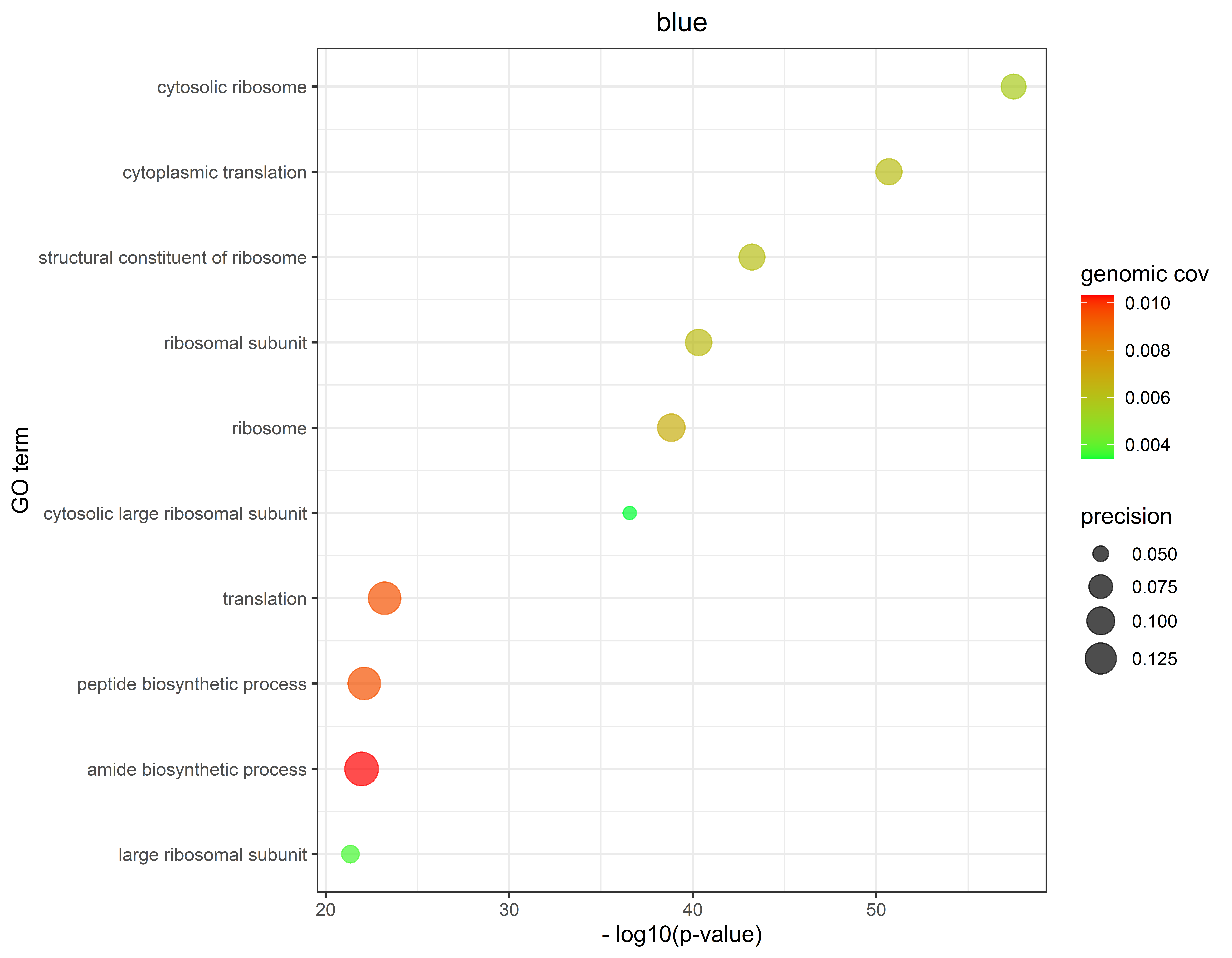
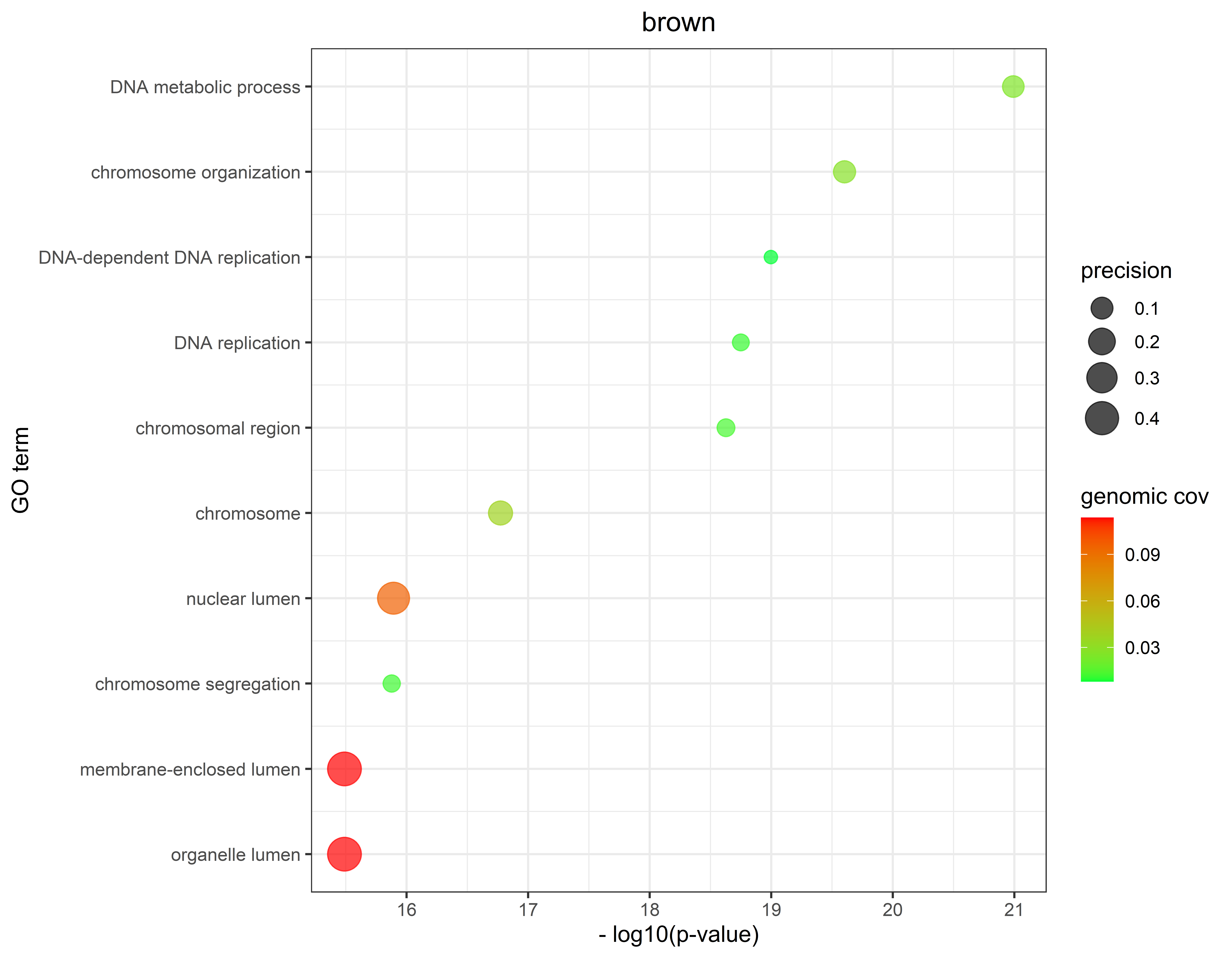
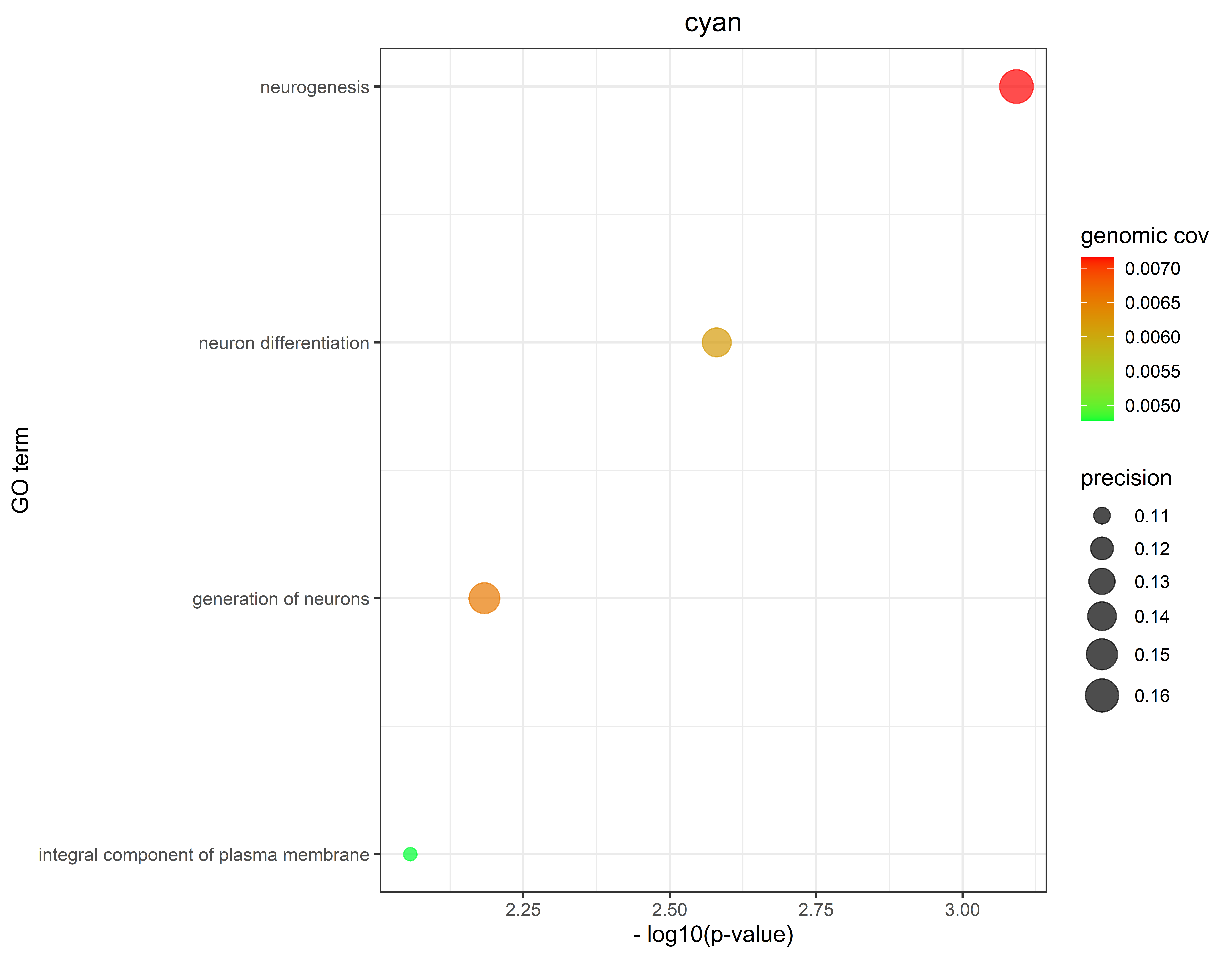
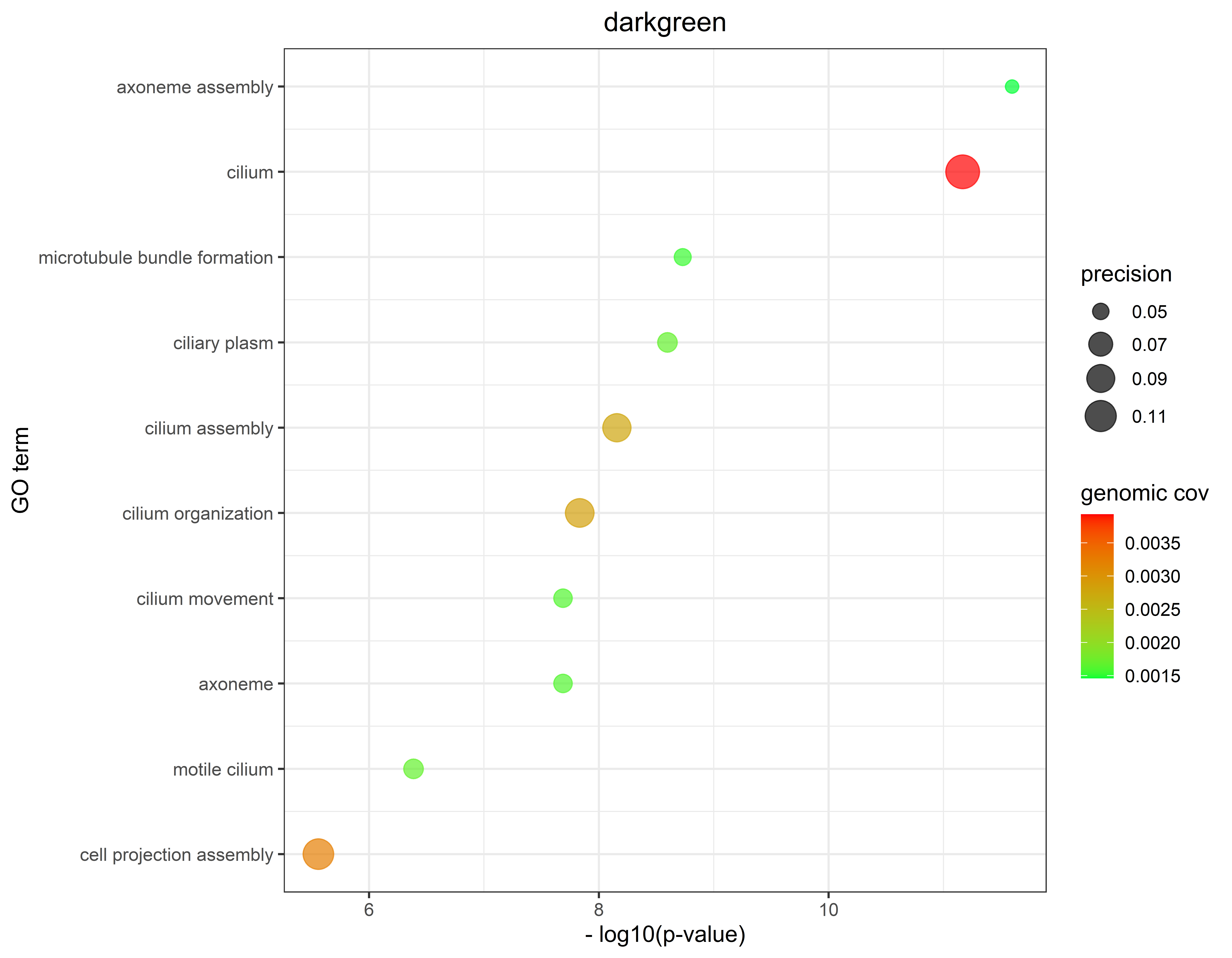
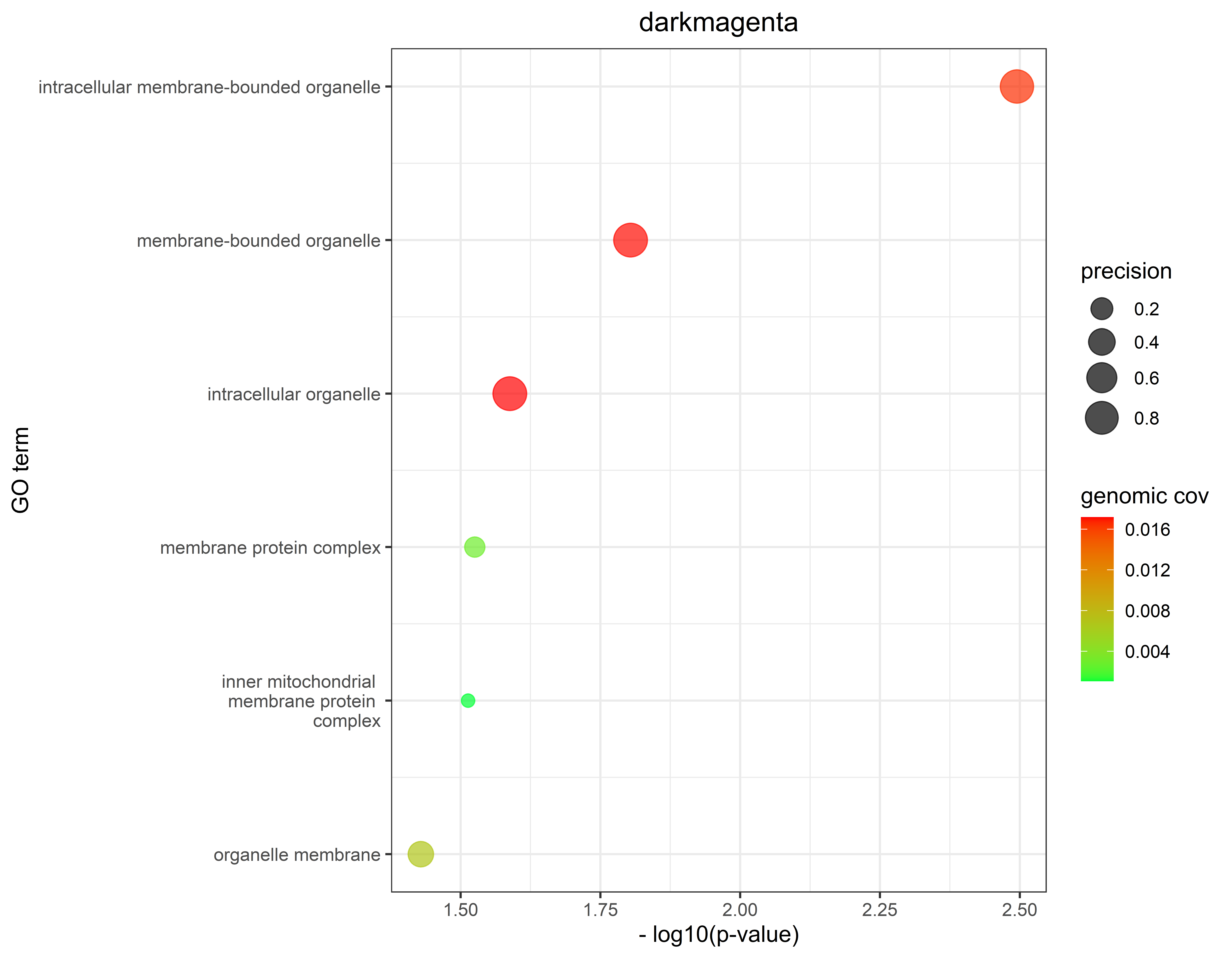
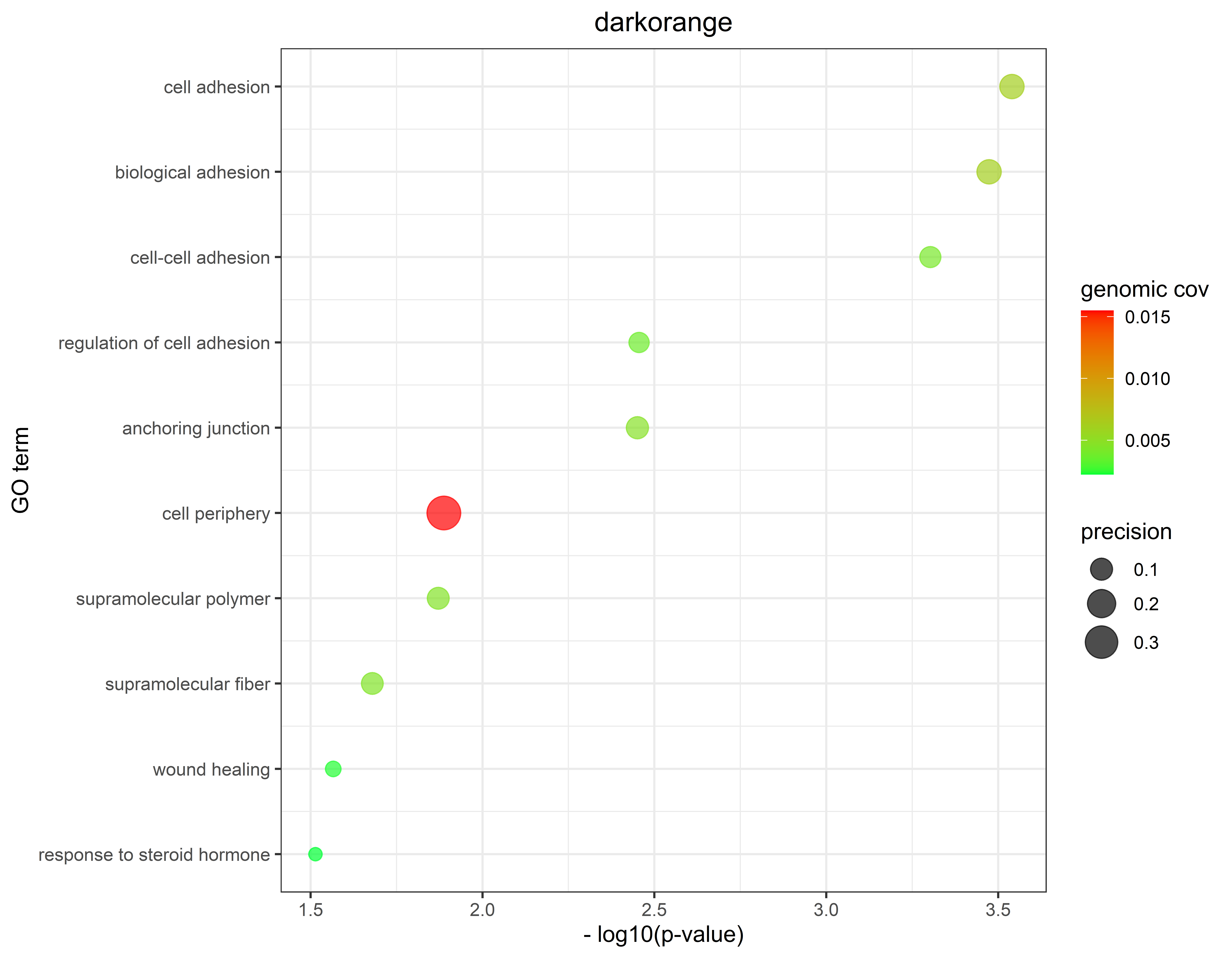
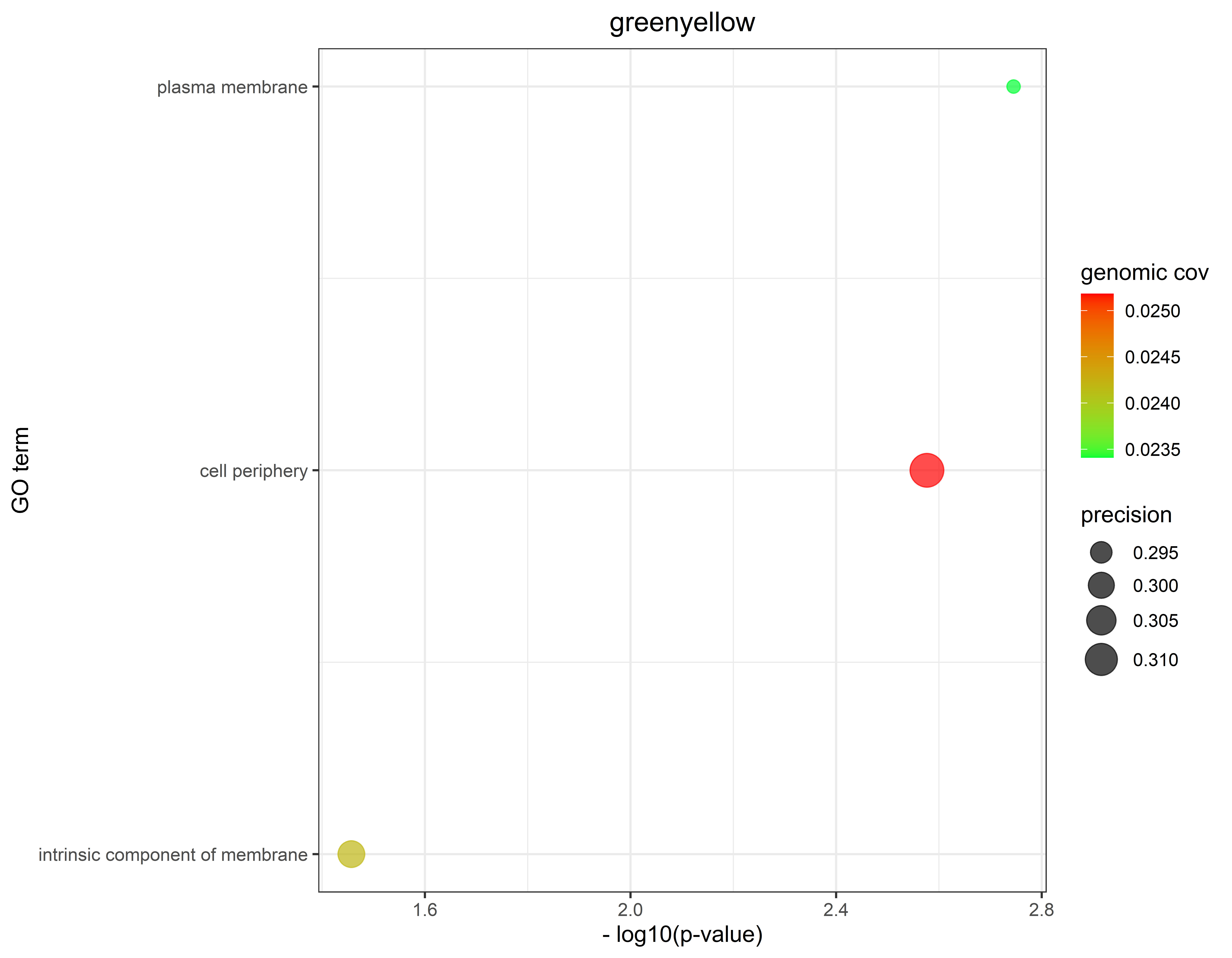
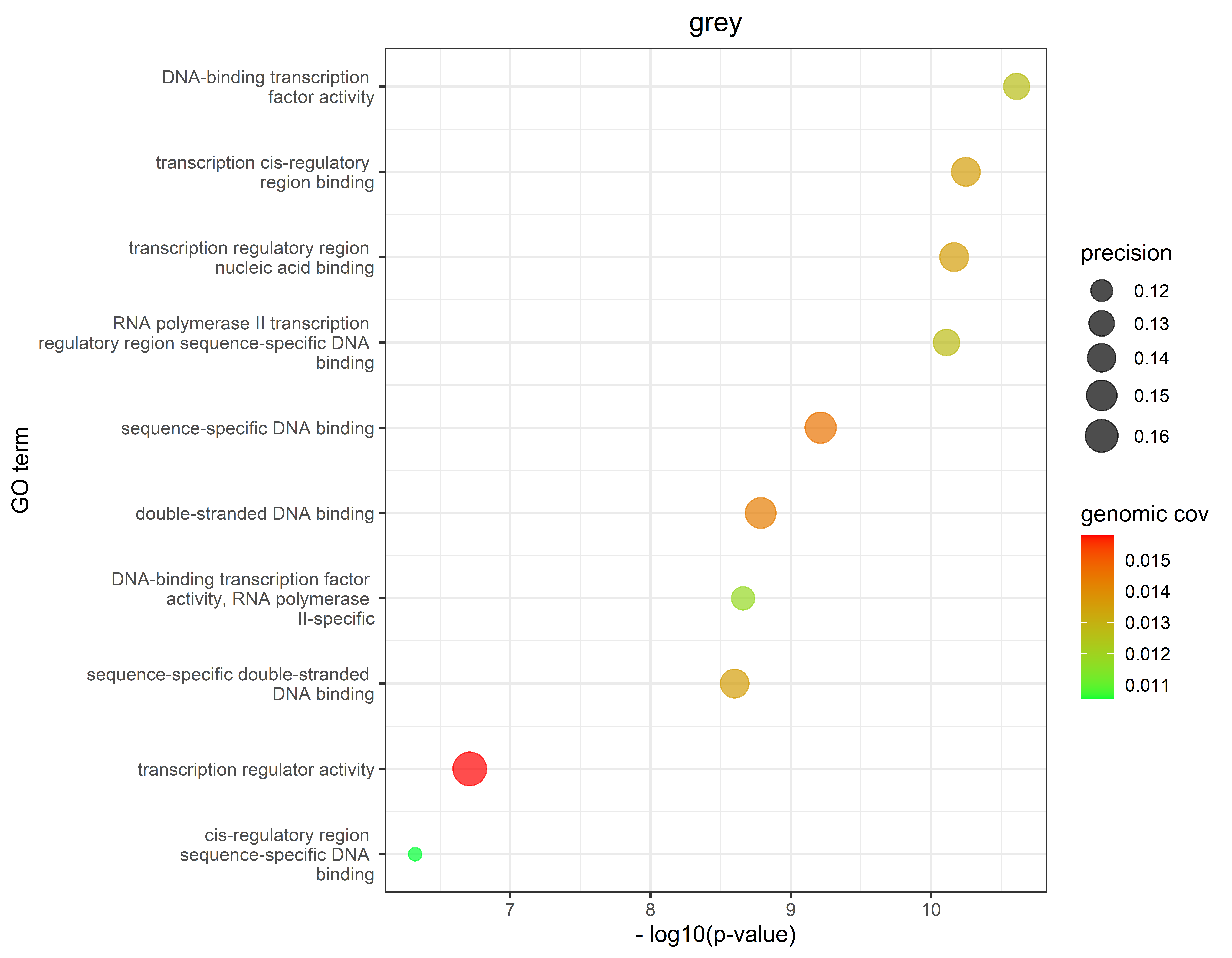
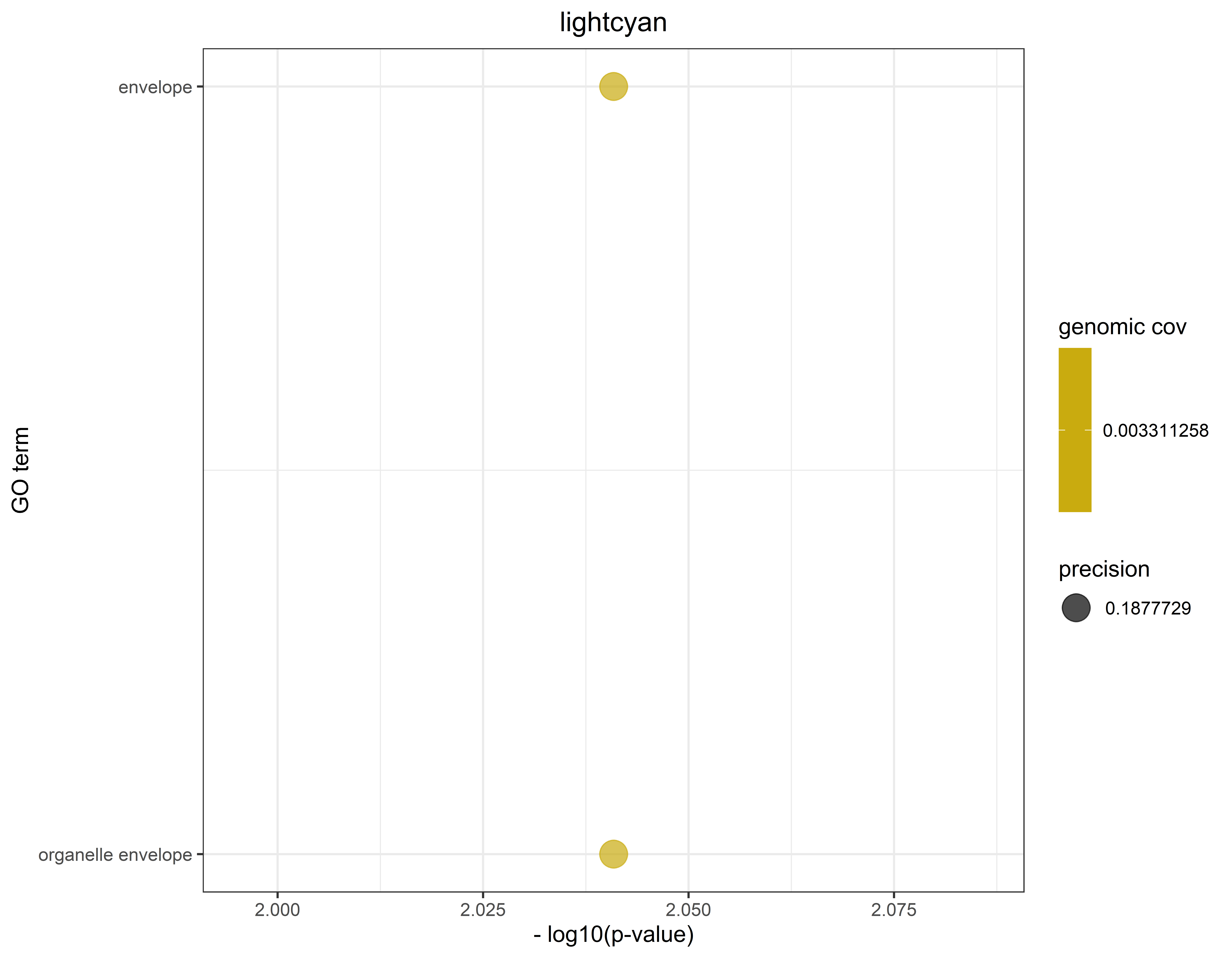
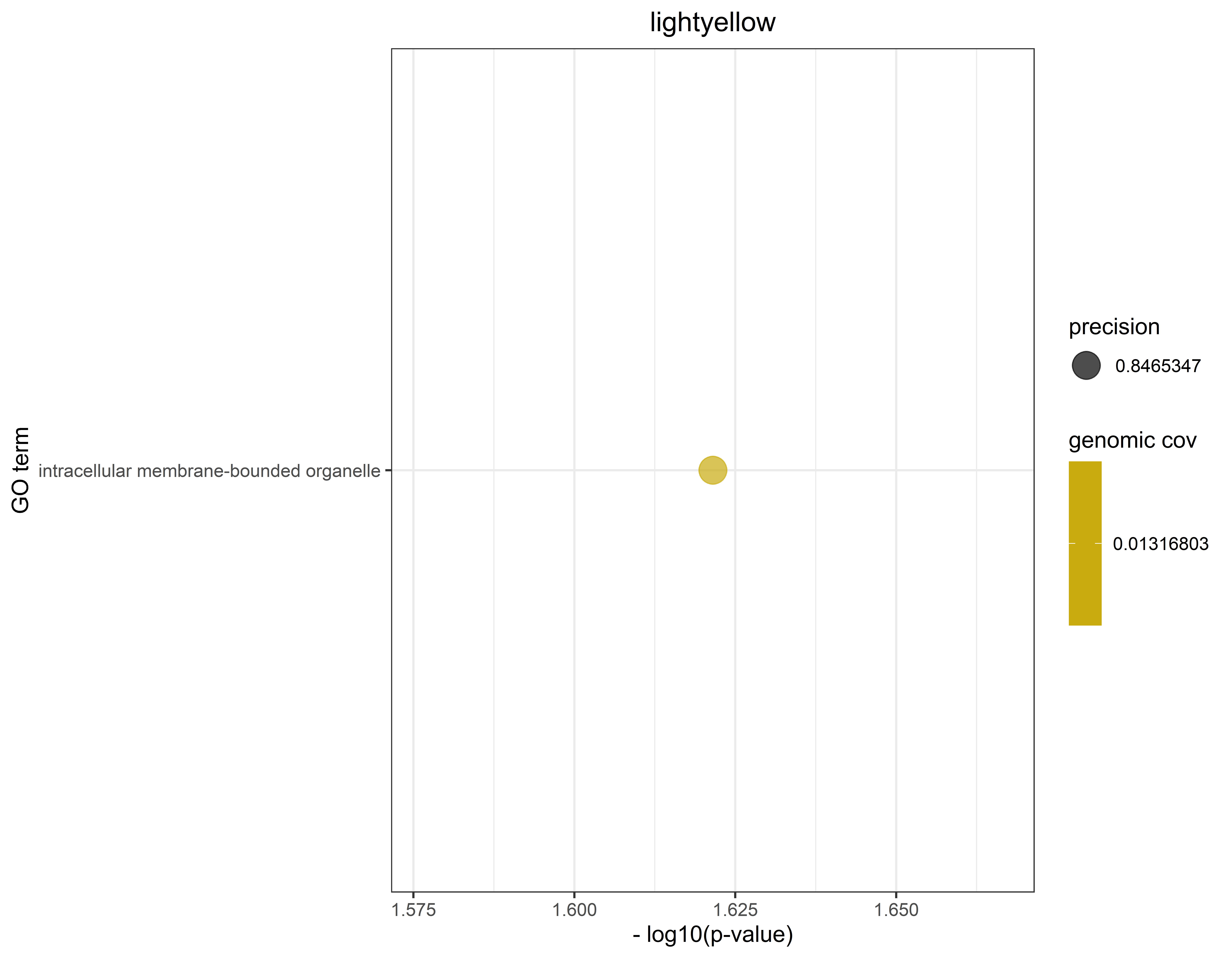
for (module in names(Genes_of_interset)){
idx = toptab$query==module & grepl("GO", toptab$source)
if(!any(idx)){
p = ggplot() + annotate("text", x = 4, y = 25, size=4,
label = "no significant GO term") +
ggtitle(module)+theme_void()+
theme(plot.title = element_text(hjust = 0.5))
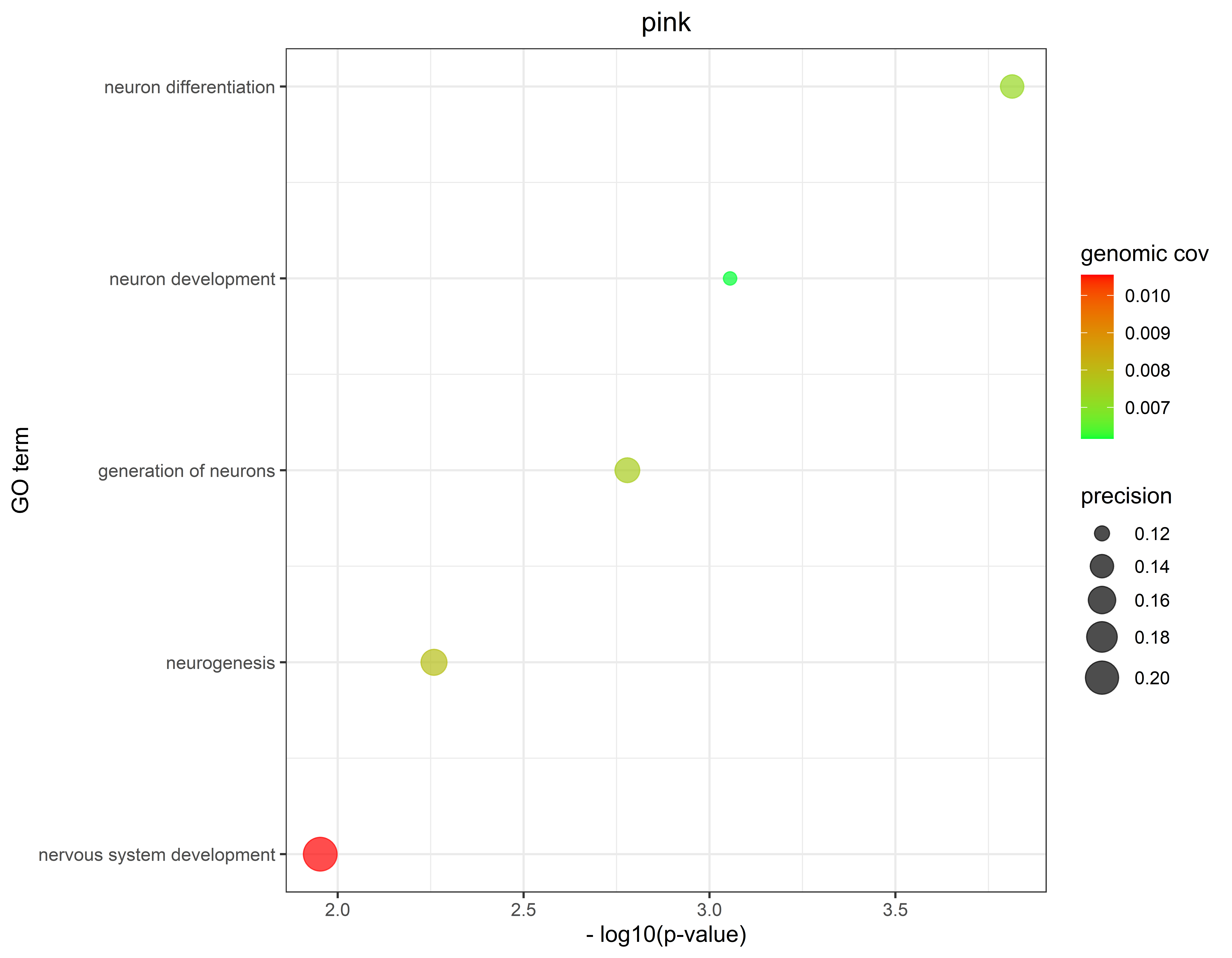
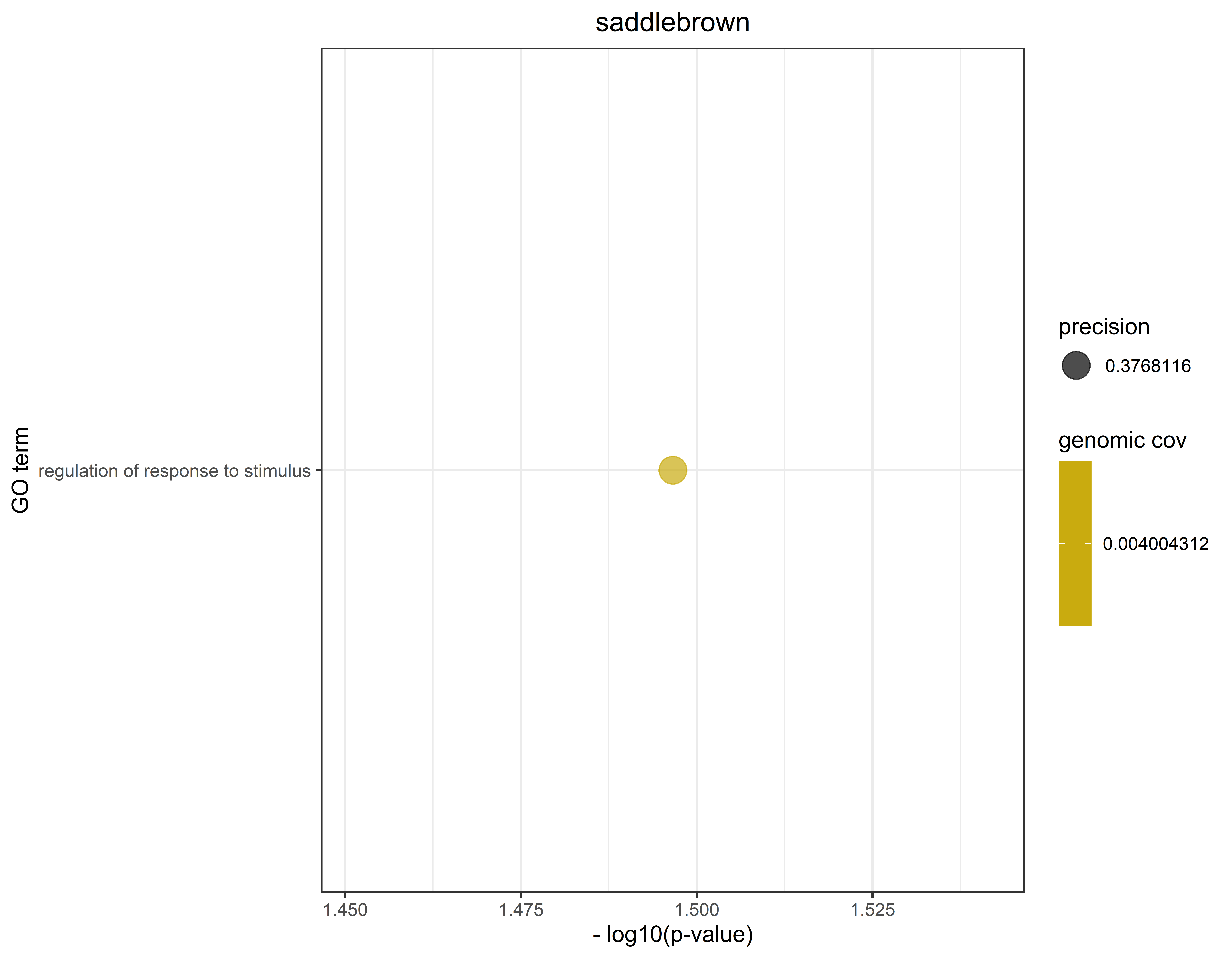
} else {
p=GOplot(toptab[idx, ], 10, Title =module)
}
print(p)
}









sessionInfo()R version 4.1.2 (2021-11-01)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 18363)
Matrix products: default
locale:
[1] LC_COLLATE=German_Germany.1252 LC_CTYPE=German_Germany.1252
[3] LC_MONETARY=German_Germany.1252 LC_NUMERIC=C
[5] LC_TIME=German_Germany.1252
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] xlsx_0.6.5 gprofiler2_0.2.1
[3] flashClust_1.01-2 WGCNA_1.70-3
[5] fastcluster_1.2.3 dynamicTreeCut_1.63-1
[7] knitr_1.37 DESeq2_1.34.0
[9] SummarizedExperiment_1.24.0 Biobase_2.54.0
[11] MatrixGenerics_1.6.0 matrixStats_0.61.0
[13] GenomicRanges_1.46.1 GenomeInfoDb_1.30.1
[15] IRanges_2.28.0 S4Vectors_0.32.3
[17] BiocGenerics_0.40.0 pheatmap_1.0.12
[19] RColorBrewer_1.1-2 compareGroups_4.5.1
[21] forcats_0.5.1 stringr_1.4.0
[23] dplyr_1.0.7 purrr_0.3.4
[25] readr_2.1.2 tidyr_1.1.4
[27] tibble_3.1.6 ggplot2_3.3.5
[29] tidyverse_1.3.1 kableExtra_1.3.4
[31] workflowr_1.7.0
loaded via a namespace (and not attached):
[1] utf8_1.2.2 tidyselect_1.1.1 RSQLite_2.2.9
[4] AnnotationDbi_1.56.2 htmlwidgets_1.5.4 grid_4.1.2
[7] BiocParallel_1.28.3 munsell_0.5.0 codetools_0.2-18
[10] preprocessCore_1.56.0 chron_2.3-56 withr_2.4.3
[13] colorspace_2.0-2 highr_0.9 uuid_1.0-3
[16] rstudioapi_0.13 officer_0.4.1 rJava_1.0-6
[19] labeling_0.4.2 git2r_0.29.0 GenomeInfoDbData_1.2.7
[22] bit64_4.0.5 farver_2.1.0 rprojroot_2.0.2
[25] vctrs_0.3.8 generics_0.1.2 xfun_0.29
[28] R6_2.5.1 doParallel_1.0.16 locfit_1.5-9.4
[31] bitops_1.0-7 cachem_1.0.6 DelayedArray_0.20.0
[34] assertthat_0.2.1 promises_1.2.0.1 scales_1.1.1
[37] nnet_7.3-17 gtable_0.3.0 processx_3.5.2
[40] rlang_1.0.0 genefilter_1.76.0 systemfonts_1.0.3
[43] splines_4.1.2 lazyeval_0.2.2 impute_1.68.0
[46] broom_0.7.12 checkmate_2.0.0 yaml_2.2.2
[49] modelr_0.1.8 backports_1.4.1 httpuv_1.6.5
[52] HardyWeinberg_1.7.4 Hmisc_4.6-0 tools_4.1.2
[55] ellipsis_0.3.2 gplots_3.1.1 jquerylib_0.1.4
[58] Rsolnp_1.16 Rcpp_1.0.8 base64enc_0.1-3
[61] zlibbioc_1.40.0 RCurl_1.98-1.5 ps_1.6.0
[64] rpart_4.1.16 haven_2.4.3 cluster_2.1.2
[67] fs_1.5.2 magrittr_2.0.2 data.table_1.14.2
[70] flextable_0.6.10 reprex_2.0.1 truncnorm_1.0-8
[73] whisker_0.4 hms_1.1.1 xlsxjars_0.6.1
[76] evaluate_0.14 xtable_1.8-4 XML_3.99-0.8
[79] jpeg_0.1-9 readxl_1.3.1 gridExtra_2.3
[82] compiler_4.1.2 mice_3.14.0 writexl_1.4.0
[85] KernSmooth_2.23-20 crayon_1.4.2 htmltools_0.5.2
[88] later_1.3.0 tzdb_0.2.0 Formula_1.2-4
[91] geneplotter_1.72.0 lubridate_1.8.0 DBI_1.1.2
[94] dbplyr_2.1.1 Matrix_1.4-0 cli_3.1.1
[97] parallel_4.1.2 pkgconfig_2.0.3 getPass_0.2-2
[100] foreign_0.8-82 plotly_4.10.0 xml2_1.3.3
[103] foreach_1.5.2 svglite_2.0.0 annotate_1.72.0
[106] bslib_0.3.1 webshot_0.5.2 XVector_0.34.0
[109] rvest_1.0.2 callr_3.7.0 digest_0.6.29
[112] Biostrings_2.62.0 rmarkdown_2.11 cellranger_1.1.0
[115] htmlTable_2.4.0 gdtools_0.2.3 gtools_3.9.2
[118] lifecycle_1.0.1 jsonlite_1.7.3 viridisLite_0.4.0
[121] fansi_1.0.2 pillar_1.7.0 lattice_0.20-45
[124] KEGGREST_1.34.0 fastmap_1.1.0 httr_1.4.2
[127] survival_3.2-13 GO.db_3.14.0 glue_1.6.1
[130] zip_2.2.0 png_0.1-7 iterators_1.0.13
[133] bit_4.0.4 stringi_1.7.6 sass_0.4.0
[136] blob_1.2.2 latticeExtra_0.6-29 caTools_1.18.2
[139] memoise_2.0.1 
