Statistical analysis
AGC, AY
23 3 2021
Last updated: 2022-02-21
Checks: 7 0
Knit directory: CePTER_RNASeq/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20210315) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 56805fa. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/.Rhistory
Ignored: analysis/MDSplots/D62_mdsplots.RData
Ignored: data/.Rhistory
Ignored: data/Countmatrix.RData
Ignored: oldcode/
Ignored: output/D62_GOresWGCNA.xlsx
Ignored: output/D62_ResTabs_KO.RData
Ignored: output/D62_ResTabs_KO_WGCNA.RData
Ignored: output/D62_WGCNA_adj_TOM.RData
Ignored: output/D62_dds_matrix.RData
Ignored: output/GOres_Comparisons.xlsx
Ignored: output/GOres_D244DIFFRAPA.xlsx
Ignored: output/GOres_D244DIFFnoRAPA.xlsx
Ignored: output/GOres_D244noDIFFRAPA.xlsx
Ignored: output/GOres_D244noDIFFnoRAPA.xlsx
Ignored: output/GOres_D62DIFFRAPA.xlsx
Ignored: output/GOres_D62DIFFnoRAPA.xlsx
Ignored: output/GOres_D62noDIFFRAPA.xlsx
Ignored: output/GOres_D62noDIFFnoRAPA.xlsx
Ignored: output/GOres_DIFFRAPA.xlsx
Ignored: output/GOres_DIFFnoRAPA.xlsx
Ignored: output/GOres_ReNDIFFRAPA.xlsx
Ignored: output/GOres_ReNDIFFnoRAPA.xlsx
Ignored: output/GOres_ReNnoDIFFRAPA.xlsx
Ignored: output/GOres_ReNnoDIFFnoRAPA.xlsx
Ignored: output/GOres_noDIFFRAPA.xlsx
Ignored: output/GOres_noDIFFnoRAPA.xlsx
Ignored: output/ResTabs_KO.RData
Ignored: output/ResTabs_KO_WGCNA.RData
Ignored: output/Restab_D244_DIFF_RAPA.xlsx
Ignored: output/Restab_D244_DIFF_noRAPA.xlsx
Ignored: output/Restab_D244_noDIFF_RAPA.xlsx
Ignored: output/Restab_D244_noDIFF_noRAPA.xlsx
Ignored: output/Restab_D62_DIFF_RAPA.xlsx
Ignored: output/Restab_D62_DIFF_noRAPA.xlsx
Ignored: output/Restab_D62_noDIFF_RAPA.xlsx
Ignored: output/Restab_D62_noDIFF_noRAPA.xlsx
Ignored: output/Restab_ReN_DIFF_RAPA.xlsx
Ignored: output/Restab_ReN_DIFF_noRAPA.xlsx
Ignored: output/Restab_ReN_noDIFF_RAPA.xlsx
Ignored: output/Restab_ReN_noDIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_All_DIFF_RAPA.xlsx
Ignored: output/Restab_Repl_All_noDIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_D62D244_DIFF_RAPA.xlsx
Ignored: output/Restab_Repl_D62D244_DIFF_noRAPA.xlsx
Ignored: output/Restab_Repl_D62D244_noDIFF_RAPA.xlsx
Ignored: output/Restab_Repl_D62D244_noDIFF_noRAPA.xlsx
Ignored: output/WGCNA_adj_TOM.RData
Untracked files:
Untracked: Wrapper.sh
Untracked: data/HumanM1 singlecellseq/
Untracked: data/genelists_to_test/
Untracked: mattson/
Untracked: wflow_helper.R
Unstaged changes:
Modified: analysis/MDSplots/rsconnect/shinyapps.io/molgenlab/mdsplots.dcf
Modified: output/GOresWGCNA.xlsx
Modified: output/dds_matrix.RData
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with wflow_publish() to start tracking its development.
home = getwd()Statistical analysis
Which genes are differentially regualted upon KO of DEPDC5
Modelling the KO in all different combinations
if(reanalyze | !file.exists(paste0(home,"/output/ResTabs_KO.RData"))){
# calculate all combinations
CellLines=c("D62","D244","ReN")
Rapamycin=c("noRAPA", "RAPA")
Differentiation=c("noDIFF", "DIFF")
Type_sgRNA<-list(c("sgNTC","sg2.1"),c("sgNTC","sg2.2"))
target="KO"
r = Rapamycin[1]
d = Differentiation[1]
Tp = Type_sgRNA[1]
# no random effects included
for(r in Rapamycin){
Rapafilter = SampleInfo$RAPA %in% r
for(d in Differentiation){
Difffilter = SampleInfo$DIFF %in% d
for(Tp in Type_sgRNA){
sgRNAfilter = SampleInfo$gRNA %in% Tp
vs_label=paste0(Tp, sep="", collapse="_")
for(Cl in CellLines){
Celllinefilter = SampleInfo$CellLine %in% Cl
Set = rownames(SampleInfo)[Rapafilter&Difffilter&
sgRNAfilter&Celllinefilter]
lab = paste("restab", Cl, vs_label, d,r, sep="_")
print(lab)
assign(lab,
comparison(dds_object = ddsMat, samples = Set,
target =target,randomeffect = c()))
}
}
}
}
comparisons= apropos("restab")
save(list = comparisons, file = paste0(home,"/output/ResTabs_KO.RData"))
} else
load(file = paste0(home,"/output/ResTabs_KO.RData"))[1] "restab_D62_sgNTC_sg2.1_noDIFF_noRAPA"
[1] "restab_D244_sgNTC_sg2.1_noDIFF_noRAPA"
[1] "restab_ReN_sgNTC_sg2.1_noDIFF_noRAPA"
[1] "restab_D62_sgNTC_sg2.2_noDIFF_noRAPA"
[1] "restab_D244_sgNTC_sg2.2_noDIFF_noRAPA"
[1] "restab_ReN_sgNTC_sg2.2_noDIFF_noRAPA"
[1] "restab_D62_sgNTC_sg2.1_DIFF_noRAPA"
[1] "restab_D244_sgNTC_sg2.1_DIFF_noRAPA"
[1] "restab_ReN_sgNTC_sg2.1_DIFF_noRAPA"
[1] "restab_D62_sgNTC_sg2.2_DIFF_noRAPA"
[1] "restab_D244_sgNTC_sg2.2_DIFF_noRAPA"
[1] "restab_ReN_sgNTC_sg2.2_DIFF_noRAPA"
[1] "restab_D62_sgNTC_sg2.1_noDIFF_RAPA"
[1] "restab_D244_sgNTC_sg2.1_noDIFF_RAPA"
[1] "restab_ReN_sgNTC_sg2.1_noDIFF_RAPA"
[1] "restab_D62_sgNTC_sg2.2_noDIFF_RAPA"
[1] "restab_D244_sgNTC_sg2.2_noDIFF_RAPA"
[1] "restab_ReN_sgNTC_sg2.2_noDIFF_RAPA"
[1] "restab_D62_sgNTC_sg2.1_DIFF_RAPA"
[1] "restab_D244_sgNTC_sg2.1_DIFF_RAPA"
[1] "restab_ReN_sgNTC_sg2.1_DIFF_RAPA"
[1] "restab_D62_sgNTC_sg2.2_DIFF_RAPA"
[1] "restab_D244_sgNTC_sg2.2_DIFF_RAPA"
[1] "restab_ReN_sgNTC_sg2.2_DIFF_RAPA"samesign <- function(x) {abs(sum(sign(x)))==length(x)}
mypval=0.05
colors <- rev(colorRampPalette(brewer.pal(9, "Spectral"))(255))
genelists=lapply(list.files(paste0(home,"/data/genelists_to_test"), full.names = T),
function(x){read.table(x, header=T)})
names(genelists) = gsub(".txt", "", list.files(paste0(home,"/data/genelists_to_test")))
geneids=rownames(restab_D244_sgNTC_sg2.1_noDIFF_noRAPA)
getlinmodoutput = function(targetline,
targetdiff,
targetrapa){
samplesincl = SampleInfo$DIFF==targetdiff &
SampleInfo$RAPA==targetrapa &
SampleInfo$CellLine== targetline
pvalrep= get(paste0("restab_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj<mypval
betarep = apply(cbind(get(paste0("restab_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange), 1,
samesign)
idx=which(betarep & pvalrep)
hits=geneids[idx]
restab=data.frame(
entrezgene = geneids[idx],
genname=rowData(ddsMat)[hits,"hgnc"],
log2FC_2.1 = get(paste0("restab_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
fdr_2.1 = get(paste0("restab_",
targetline, "_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj[idx],
log2FC_2.2 = get(paste0("restab_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
fdr_2.2 = get(paste0("restab_",
targetline, "_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj[idx])
print(restab)
write.xlsx(restab, file=paste0(output, "/Restab_",
targetline,"_",
targetdiff, "_",
targetrapa, ".xlsx"))
SamplesSet=SampleInfo[samplesincl,]
plotmatrix = log_2cpm[hits,rownames(SamplesSet)]
rownames(SamplesSet)=SamplesSet$label_rep
colnames(plotmatrix)=SamplesSet$label_rep
rownames(plotmatrix)=rowData(ddsMat)[hits,"hgnc"]
cellcol = Dark8[1:nlevels(SampleInfo$CellLine)]
names(cellcol) = levels(SampleInfo$CellLine)
cellcol = cellcol[as.character(unique(SamplesSet$CellLine))]
gRNAcol = Dark8[c(1:nlevels(SampleInfo$gRNA))+nlevels(SampleInfo$CellLine)]
names(gRNAcol) = levels(SampleInfo$gRNA)
gRNAcol = gRNAcol[as.character(unique(SamplesSet$gRNA))]
diffcol = brewer.pal(3,"Set1")[1:nlevels(SampleInfo$DIFF)]
names(diffcol) = levels(SampleInfo$DIFF)
diffcol = diffcol[as.character(unique(SamplesSet$DIFF))]
rapacol = brewer.pal(3,"Set2")[1:nlevels(SampleInfo$RAPA)]
names(rapacol) = levels(SampleInfo$RAPA)
rapacol = rapacol[as.character(unique(SamplesSet$RAPA))]
ann_colors = list(
DIFF = diffcol,
RAPA = rapacol,
gRNA = gRNAcol,
CellLine=cellcol)
collabels = SamplesSet[,c("CellLine","DIFF","RAPA", "gRNA")] %>%
mutate_all(as.character) %>% as.data.frame()
rownames(collabels)=SamplesSet$label_rep
idx=order(SamplesSet$gRNA)
pheatmap(plotmatrix[,idx],
border_color = NA,
annotation_col = collabels[idx,],show_rownames = F, show_colnames = F,
annotation_colors = ann_colors,
clustering_method = "ward.D2",
cluster_cols = F,
col = colors,
scale = "row",
main = paste("Normalized log2 counts", targetline, targetdiff, targetrapa))
gene_univers = rownames(ddsMat)
gostres = getGOresults(hits, gene_univers)
toptab = gostres$result
write.xlsx2(toptab, file = paste0(output, "/GOres_",targetline,targetdiff,targetrapa,".xlsx"), sheetName = "GO_enrichment")
idx = grepl("GO|KEGG", toptab$source)
titlename = paste(targetline,targetdiff,targetrapa)
if(!any(idx)){
p = ggplot() + annotate("text", x = 4, y = 25, size=4,
label = "no significant GO term") +
ggtitle(titlename)+theme_void()+
theme(plot.title = element_text(hjust = 0.5))
} else {
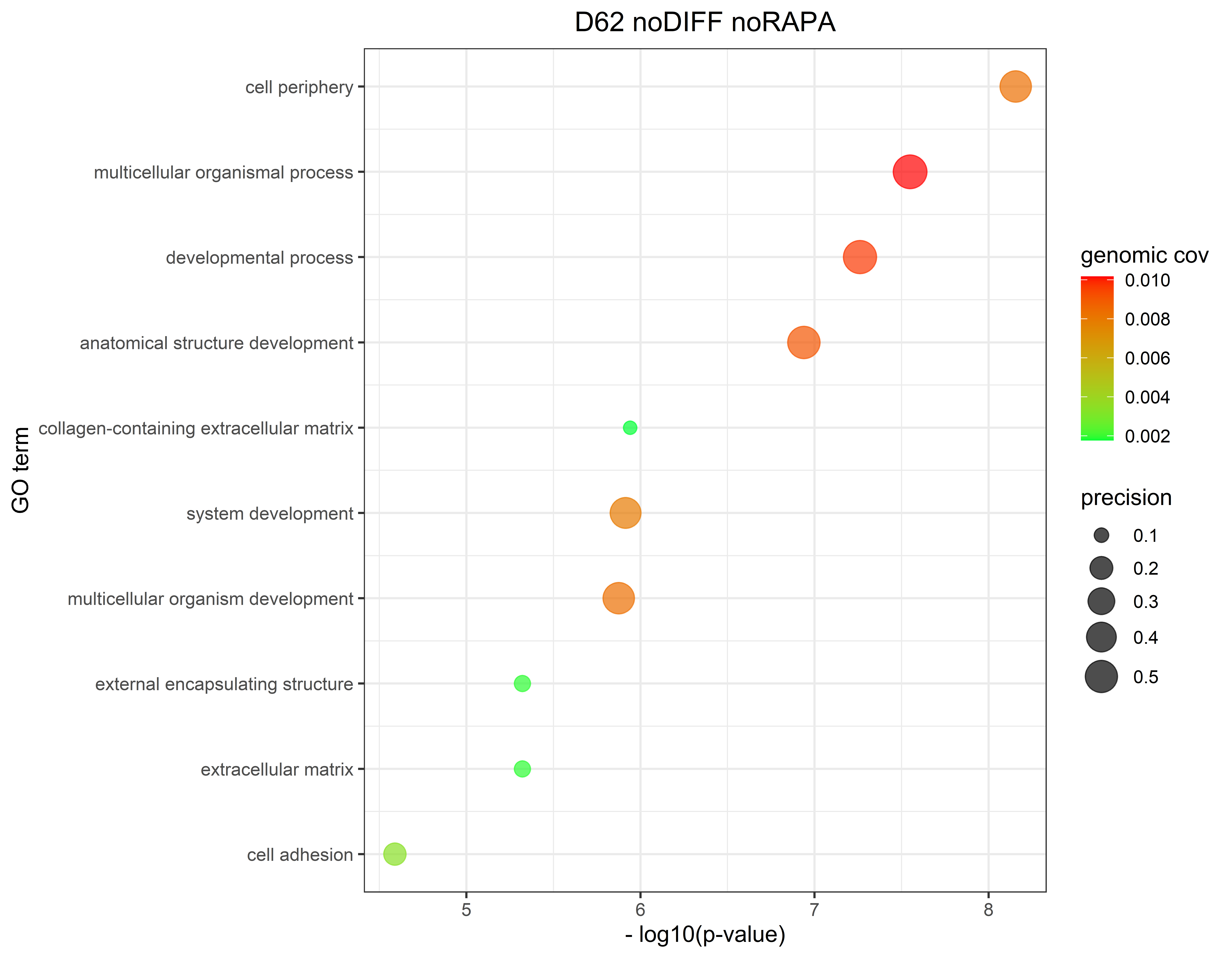
p=GOplot(toptab[idx, ], 10, Title = titlename)
}
print(p)
## enrichment test for genelist
Resall=data.frame()
for (gl in names(genelists)){
tmp=table(signif=geneids %in% hits, targetgene=geneids %in% genelists[[gl]]$ENTREZ_ID)
resfish=fisher.test(tmp)
res = c(resfish$estimate, unlist(resfish$conf.int), resfish$p.value)
Resall = rbind(Resall, res)
}
colnames(Resall)=c("OR", "CI95L", "CI95U", "P")
rownames(Resall)=names(genelists)
Resall$Beta = log(Resall$OR)
Resall$SE = (log(Resall$OR)-log(Resall$CI95L))/1.96
Resall$Padj=p.adjust(Resall$P, method = "bonferroni")
print(Resall)
multiORplot(Resall, Pval = "P", Padj = "Padj", beta="Beta",SE = "SE",
pheno=paste(targetline, targetdiff, targetrapa))
}
geneids=rownames(restab_D244_sgNTC_sg2.1_noDIFF_noRAPA)KO effect in noDIFF noRAPA
D62
getlinmodoutput(targetline = "D62",targetdiff = "noDIFF",targetrapa = "noRAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 64856 VWA1 1.0887369 3.357074e-12 0.7289010 3.093542e-06
2 26038 CHD5 2.6317640 1.274240e-07 1.4617556 1.601009e-02
3 3925 STMN1 -0.8137179 4.487672e-11 -0.4038794 1.435877e-04
4 90853 SPOCD1 1.5412353 1.172424e-05 0.9562654 4.592191e-02
5 23499 MACF1 0.3842645 4.982204e-02 0.3351333 2.463005e-02
6 59269 HIVEP3 0.8394967 5.265490e-03 0.7742331 8.323388e-04
7 11004 KIF2C -1.4726626 3.069666e-06 -1.2743887 6.113960e-08
8 5052 PRDX1 -0.5768855 1.130289e-09 -0.4276079 7.426535e-09
9 9077 DIRAS3 1.8022039 1.130961e-14 1.5359253 9.571880e-21
10 81849 ST6GALNAC5 -0.7860855 1.743212e-10 -0.4107442 1.195027e-05
11 64123 ADGRL4 1.3765063 1.449484e-03 1.8278999 5.108277e-10
12 2633 GBP1 4.3469675 1.478239e-04 4.0111487 9.206477e-05
13 84722 PSRC1 -0.4544078 8.415406e-04 -0.3012023 1.048519e-02
14 3321 IGSF3 -0.8447306 8.186979e-04 -0.5463340 7.250332e-03
15 10628 TXNIP -1.3807391 3.855992e-05 -1.1462088 4.458761e-07
16 140576 S100A16 0.8907779 7.208856e-10 0.3327141 4.177523e-02
17 10763 NES 0.7597404 9.985796e-14 0.4268953 7.990741e-07
18 57863 CADM3 -2.0291449 1.345453e-03 -1.0765544 3.564895e-02
19 400793 C1orf226 0.8128065 6.289715e-03 0.6009301 2.626275e-02
20 4921 DDR2 -0.8517003 3.949753e-02 -0.8841425 8.323388e-04
21 5999 RGS4 2.1839355 3.625474e-08 2.0253972 1.178328e-07
22 6004 RGS16 1.3221396 1.734986e-11 0.5614850 2.148512e-02
23 64710 NUCKS1 -0.4841646 2.456791e-06 -0.2907053 1.390388e-03
24 1063 CENPF -1.1466936 3.609332e-13 -0.4539131 2.168138e-04
25 149111 CNIH3 1.0685427 4.092541e-03 1.0633895 4.535816e-04
26 84886 C1orf198 0.5477246 3.424084e-02 0.5164650 2.383525e-02
27 4811 NID1 -1.5148942 1.904365e-03 -1.0105930 2.296366e-02
28 22944 KIN -2.0047172 5.502120e-04 -1.1833437 1.036235e-02
29 5660 PSAP 0.7103618 1.949853e-20 0.1806493 7.600955e-03
30 54541 DDIT4 1.0549545 3.214122e-03 0.8462576 2.066682e-02
31 5328 PLAU 1.2381776 1.514050e-02 1.0763653 3.554633e-02
32 84858 ZNF503 1.8248076 3.757224e-04 1.5518196 1.304512e-03
33 3988 LIPA 1.0229774 1.258333e-04 0.6516045 3.554633e-02
34 143282 FGFBP3 0.6236312 2.454262e-04 0.4636031 4.197354e-03
35 26509 MYOF 1.7150966 3.176843e-02 2.0884036 1.376061e-03
36 9124 PDLIM1 1.1007675 7.192111e-04 1.1011312 2.589242e-06
37 83742 MARVELD1 -0.8437692 1.245102e-02 -0.6506239 3.298481e-02
38 6319 SCD 0.5326328 1.598335e-05 0.3381076 1.082659e-03
39 6468 FBXW4 1.1765193 1.364798e-03 0.7725283 3.557110e-02
40 259217 HSPA12A 1.8426634 3.216693e-03 1.6189322 2.216133e-02
41 642938 INSYN2A 0.4021455 4.047089e-02 0.7784115 7.692471e-10
42 4288 MKI67 -1.0125441 1.226143e-06 -0.5883143 5.751303e-03
43 977 CD151 1.2624849 1.664143e-27 0.3302487 9.754474e-03
44 6240 RRM1 -0.8821072 1.121684e-02 -1.0392511 1.970332e-04
45 4004 LMO1 -2.3548878 2.048639e-07 -2.7124074 9.680117e-22
46 10553 HTATIP2 1.9347967 1.616995e-02 2.0024902 5.325513e-04
47 5080 PAX6 -0.9174372 1.022465e-02 -0.8138011 2.649379e-03
48 960 CD44 1.4577320 5.415566e-16 1.1687918 9.473981e-16
49 79026 AHNAK 1.5854085 1.414586e-09 0.9180973 4.510473e-04
50 1717 DHCR7 0.7292135 2.901922e-04 0.3966686 2.999299e-02
51 26011 TENM4 1.7228097 2.374904e-02 1.5164792 4.490345e-02
52 11098 PRSS23 0.8783873 1.873156e-02 0.9009730 3.505038e-04
53 1410 CRYAB 1.1162846 2.410543e-12 0.5939001 9.419441e-05
54 6876 TAGLN 0.6535509 1.315854e-13 0.3428368 4.037341e-08
55 1656 DDX6 -0.4627267 1.974148e-02 -0.3685693 5.784508e-03
56 2012 EMP1 0.6467583 1.466047e-02 1.1892730 8.967278e-13
57 196410 METTL7B 1.5592188 2.247789e-12 0.7194504 2.135379e-03
58 967 CD63 0.9012238 5.749064e-40 0.2548988 9.247364e-06
59 4035 LRP1 0.6450656 5.103070e-05 0.5926954 2.393841e-07
60 4193 MDM2 0.6720729 4.176205e-05 0.3624504 1.186010e-02
61 160335 TMTC2 -1.6150650 5.031119e-06 -1.0540218 1.594156e-04
62 490 ATP2B1 -0.7884871 1.345453e-03 -0.6051893 4.758216e-04
63 5829 PXN 0.7635220 3.848152e-04 0.5831564 7.783165e-03
64 3146 HMGB1 -0.7290124 6.419766e-16 -0.3207209 7.050456e-07
65 8848 TSC22D1 0.5605586 4.645096e-04 0.6643643 1.133959e-11
66 8660 IRS2 0.4095870 2.096683e-05 0.2994912 4.758216e-04
67 28526 <NA> 1.9269222 1.459106e-07 1.9184477 1.771068e-11
68 54916 TMEM260 -1.0305660 3.898016e-02 -0.9100269 3.893809e-02
69 57570 TRMT5 -0.6823055 1.092744e-04 -0.3822535 7.374547e-03
70 23224 SYNE2 -1.3797637 2.074033e-06 -0.7351991 6.014667e-05
71 64093 SMOC1 0.7195505 1.391368e-07 0.3608222 8.928172e-04
72 9369 NRXN3 -1.7172025 2.028184e-05 -1.0867269 7.910845e-04
73 55384 MEG3 1.6236408 2.580979e-22 1.1925473 2.944858e-15
74 113146 AHNAK2 2.2305093 1.032644e-04 2.6588628 1.577622e-14
75 11245 GPR176 2.1627304 3.572180e-03 2.0993058 1.663975e-03
76 90417 KNSTRN -1.3211373 3.587749e-03 -0.8164001 4.080416e-02
77 51203 NUSAP1 -1.3679769 1.839058e-06 -0.5187317 8.823202e-03
78 2628 GATM 0.6518976 3.394949e-02 1.0425253 7.502761e-12
79 302 ANXA2 0.3958604 1.837163e-04 0.3001945 9.344256e-03
80 7168 TPM1 0.5157933 2.102621e-02 0.4722003 1.661191e-03
81 9768 PCLAF -1.6219457 5.811956e-04 -1.0914799 3.160828e-03
82 3671 ISLR 1.4575124 6.307058e-08 0.8572040 9.297192e-04
83 23205 ACSBG1 0.6910454 1.236709e-02 0.5932626 2.935875e-02
84 9055 PRC1 -0.7447847 1.733945e-02 -0.5062243 4.744675e-02
85 56963 RGMA 1.6169697 1.508787e-22 0.7278730 4.458761e-07
86 23336 SYNM 0.7042096 6.696236e-05 0.7291826 2.725960e-08
87 51330 TNFRSF12A 1.4047167 1.598984e-20 0.9561323 4.156772e-16
88 2013 EMP2 -0.7553862 7.112681e-05 -0.4957736 7.600955e-03
89 26471 NUPR1 2.3604930 1.275840e-12 1.0670195 2.296366e-02
90 4313 MMP2 0.8695009 2.490348e-09 0.4227913 6.023638e-03
91 6376 CX3CL1 0.8933944 1.364798e-03 0.5525095 1.857524e-02
92 794 CALB2 -4.6134518 1.676816e-06 -2.1153464 2.076907e-03
93 23406 COTL1 0.7985791 1.235229e-09 0.4709035 1.578005e-06
94 79007 DBNDD1 1.9194455 9.495285e-25 0.6782882 1.055840e-03
95 9212 AURKB -1.2410536 2.093176e-03 -1.1951322 1.663975e-03
96 9423 NTN1 1.4853260 2.410543e-12 0.6565916 6.359039e-03
97 9953 HS3ST3B1 -0.7873216 1.713457e-04 -0.4180975 7.956870e-03
98 5376 PMP22 0.5562334 2.753659e-03 0.4810478 1.442876e-03
99 125144 SNHG29 -0.5333552 6.646735e-03 -0.2828075 2.579931e-02
100 7153 TOP2A -0.9719652 2.450201e-11 -0.8228819 1.091862e-15
101 284119 CAVIN1 1.0598237 1.045027e-04 0.6193031 4.515208e-02
102 2670 GFAP 1.0680802 1.138551e-30 0.2611641 4.975830e-03
103 6426 SRSF1 -0.9535097 1.971245e-06 -0.5774088 6.881405e-04
104 54549 SDK2 -0.9457414 4.464916e-02 -0.8399915 4.106571e-02
105 283991 UBALD2 0.5562493 2.846519e-05 0.3803418 3.807842e-03
106 332 BIRC5 -1.2794951 4.036746e-04 -0.7407098 8.323388e-04
107 284184 NDUFAF8 0.8491433 5.692768e-12 0.3481626 7.808378e-03
108 11031 RAB31 0.3240090 2.195959e-03 0.2517174 1.169403e-02
109 1000 CDH2 0.2476283 2.926897e-02 0.3202481 4.270465e-08
110 115106 HAUS1 -1.6978514 1.572448e-03 -0.9650103 8.536814e-03
111 4645 MYO5B -1.4849117 9.083405e-03 -0.8087432 3.513141e-02
112 5596 MAPK4 -1.1651266 3.734472e-03 -0.9121994 1.970914e-02
113 976 ADGRE5 1.5558481 1.840861e-10 0.9709443 2.429572e-04
114 9518 GDF15 1.7726956 9.495285e-25 0.9120218 2.725960e-08
115 348 APOE 2.0354059 3.708190e-20 1.2386377 5.932062e-11
116 83987 CCDC8 -2.7542907 4.471965e-04 -2.1432234 2.880361e-04
117 27113 BBC3 1.5019300 2.885492e-09 1.1144906 4.741474e-06
118 23645 PPP1R15A 0.7984083 3.369973e-04 0.5185689 3.082708e-02
119 79784 MYH14 -3.2167632 4.355945e-07 -3.3864313 8.830222e-10
120 151354 LRATD1 0.9969107 2.744760e-09 0.8078464 5.025839e-08
121 6432 SRSF7 -0.5313444 1.694898e-02 -0.4872875 3.135324e-02
122 91461 PKDCC -0.5131526 2.577269e-04 -0.4124175 7.177574e-05
123 25927 CNRIP1 -6.6985860 1.150131e-03 -5.2247905 2.200852e-03
124 3099 HK2 -1.7911145 3.646131e-03 -1.1136392 3.778682e-03
125 343990 CRACDL 1.5489677 8.186979e-04 1.5058582 5.099405e-06
126 164832 LONRF2 0.8056629 9.967899e-04 0.6150822 1.054530e-02
127 4175 MCM6 -1.2006023 1.142748e-02 -1.0394463 6.465256e-03
128 130574 LYPD6 -1.2615303 4.390729e-02 -1.6173378 8.961483e-03
129 114793 FMNL2 0.8386577 2.025151e-08 0.5693890 1.320261e-05
130 55854 ZC3H15 -0.6073926 1.028732e-02 -0.4319460 3.689260e-02
131 26010 SPATS2L 0.4040408 4.069513e-02 0.5241896 6.406613e-04
132 389073 C2orf80 -0.6860560 3.021141e-05 -0.3989007 4.037353e-03
133 1373 CPS1 -2.4113344 3.135118e-02 -1.5363111 1.115901e-02
134 2637 GBX2 0.9261028 4.130967e-03 0.6671906 2.396204e-02
135 10267 RAMP1 1.3715391 8.566720e-12 0.8507912 2.765416e-05
136 2280 FKBP1A 0.6962166 2.508668e-05 0.3635890 4.886965e-02
137 5834 PYGB 1.0709729 1.027140e-07 0.6084430 2.308014e-02
138 22974 TPX2 -1.3233710 9.234264e-04 -0.7952465 5.345927e-03
139 1299 COL9A3 0.5755993 4.365583e-05 0.2846454 7.126419e-03
140 3785 KCNQ2 -0.6122840 2.104394e-03 -0.6171374 3.496026e-05
141 140578 CHODL -2.8003063 5.155278e-04 -1.5315831 2.071870e-02
142 116448 OLIG1 0.9333397 1.715356e-05 0.5884338 1.490735e-02
143 5121 PCP4 -1.5668394 7.672433e-03 -1.1879093 4.314608e-02
144 1826 DSCAM 1.5768898 6.716536e-04 2.1033772 2.500117e-11
145 80781 COL18A1 1.3174083 2.148859e-07 0.8091186 6.369037e-03
146 1312 COMT 0.5102825 1.021327e-05 0.3237802 7.783165e-03
147 23467 NPTXR 1.1213183 7.857622e-13 0.5433013 8.323388e-04
148 10752 CHL1 -1.0508060 2.599907e-12 -0.4931137 1.022821e-05
149 23171 GPD1L -1.6693632 7.709013e-04 -1.2523108 8.503905e-04
150 56992 KIF15 -1.1480147 1.244552e-02 -1.3058315 1.073764e-02
151 151887 CCDC80 0.7861593 3.884032e-08 0.5444540 5.158658e-06
152 2596 GAP43 1.6359187 4.388269e-60 1.0807341 3.797828e-33
153 11343 MGLL 0.9202739 4.176205e-05 0.6135539 8.415630e-03
154 5947 RBP1 -0.4686005 1.883104e-02 -0.7458186 2.387617e-07
155 10051 SMC4 -1.3308521 1.625283e-13 -0.5794336 1.187866e-03
156 8646 CHRD 1.6849809 3.167966e-02 1.7586855 2.058539e-03
157 6480 ST6GAL1 -4.2853399 1.578328e-02 -7.1401341 4.510762e-04
158 131583 FAM43A 1.7595220 8.504312e-05 1.2640110 5.157637e-03
159 10460 TACC3 -0.9068362 1.481627e-02 -0.9370790 5.973781e-05
160 80333 KCNIP4 -2.1218707 7.016198e-05 -1.2832211 3.364399e-02
161 5099 PCDH7 -2.6030063 5.882622e-03 -1.5811041 1.989404e-02
162 5156 PDGFRA -0.9547393 5.308860e-08 -0.4376841 5.750365e-04
163 3490 IGFBP7 1.4844042 6.295374e-19 1.8282513 1.823991e-53
164 79966 SCD5 0.4041952 1.713457e-04 0.7016327 2.148795e-28
165 8404 SPARCL1 2.1771329 5.091908e-10 2.1524214 3.073504e-16
166 3015 H2AZ1 -0.7067496 1.619655e-07 -0.4982645 2.218205e-05
167 1062 CENPE -2.1763776 6.800897e-20 -0.5713744 4.814100e-03
168 55013 MCUB -0.8277119 5.244963e-03 -0.6410976 1.125196e-03
169 2983 GUCY1B1 -1.9361826 9.943220e-03 -1.3455314 1.857524e-02
170 4889 NPY5R -1.2095153 3.665877e-02 -1.3273711 1.082659e-03
171 3148 HMGB2 -1.4512300 1.165034e-07 -0.5867647 1.795939e-04
172 55714 TENM3 -2.5366838 1.320516e-13 -0.5545306 2.331545e-02
173 1767 DNAH5 5.8173464 3.510642e-02 5.6528496 4.391011e-02
174 7204 TRIO 0.7074614 8.165427e-03 0.5640108 7.126419e-03
175 6507 SLC1A3 -0.5136744 3.320557e-07 -0.3891983 7.545064e-07
176 102467080 <NA> -2.2092766 4.296921e-02 -2.7772256 4.319364e-03
177 728723 <NA> 7.3304049 8.902443e-04 6.3859247 2.080350e-02
178 22836 RHOBTB3 -0.4283098 3.503150e-02 -0.3687419 6.599365e-04
179 4001 LMNB1 -1.2374118 4.005049e-11 -0.6979132 4.107474e-06
180 2201 FBN2 -1.3303062 5.846515e-03 -1.1768395 9.520106e-03
181 10112 KIF20A -1.3666360 7.533915e-05 -1.0740840 4.450953e-04
182 84418 CYSTM1 0.7223302 1.783180e-03 0.4655625 2.834089e-02
183 113829 SLC35A4 1.2354645 5.331330e-06 0.6330907 3.365639e-02
184 57717 PCDHB16 -1.3035828 4.956212e-04 -0.7859098 1.366067e-02
185 2246 FGF1 0.7326434 7.533915e-05 0.6738093 5.559947e-05
186 202374 STK32A 1.6464199 2.344297e-03 1.3008830 6.632451e-03
187 6678 SPARC 0.7424626 1.147023e-25 0.6113324 3.363226e-23
188 9232 PTTG1 -0.6683855 1.662312e-03 -0.5329348 4.107474e-06
189 221749 PXDC1 0.5202967 1.478111e-02 0.4275719 3.689260e-02
190 51299 NRN1 2.8330955 2.792238e-04 2.7357805 1.364090e-04
191 10590 SCGN -3.2952996 5.370178e-03 -1.0156297 2.414624e-06
192 222698 NKAPL 3.5957149 1.430573e-02 3.5843063 2.429572e-04
193 7726 TRIM26 6.5707922 2.067838e-03 6.5391753 2.715268e-03
194 221491 SMIM29 0.9411454 1.543927e-04 0.4827715 4.565178e-02
195 1026 CDKN1A 1.7451782 3.284453e-77 0.8937057 6.528518e-32
196 3910 LAMA4 0.4073207 4.502692e-02 0.4391435 1.254131e-02
197 57224 NHSL1 1.4575446 5.239096e-03 1.2023725 2.635414e-02
198 6196 RPS6KA2 0.8469458 2.974890e-07 0.4754517 4.349091e-03
199 221833 SP8 -0.9894228 5.967473e-06 -0.6121173 1.825002e-03
200 55536 CDCA7L -0.7781443 2.217032e-04 -0.5020097 1.625640e-02
201 10457 GPNMB 1.5149068 1.625164e-04 1.2501414 9.350606e-04
202 4852 NPY -1.6633964 1.919357e-08 -1.3042111 9.287714e-08
203 64409 GALNT17 2.4288004 4.072252e-06 1.4306650 3.090657e-02
204 23089 PEG10 0.3625560 1.736585e-02 0.5081485 6.768817e-09
205 5054 SERPINE1 1.2466133 5.238666e-15 1.2036148 1.243155e-18
206 5764 PTN 0.3452771 1.414878e-03 0.3264906 6.316862e-07
207 8795 TNFRSF10B 0.7811169 2.729630e-02 0.8057251 1.868080e-03
208 4741 NEFM 1.8233265 2.462344e-04 1.4783114 1.580777e-03
209 4747 NEFL 1.3034626 1.191912e-07 0.8377696 5.373116e-04
210 2185 PTK2B 1.8426546 1.929829e-13 1.0408686 8.209092e-05
211 1191 CLU 0.6341170 6.617167e-18 0.3197075 7.692471e-10
212 157570 ESCO2 -2.3357895 2.102632e-02 -1.5653693 9.297192e-04
213 5327 PLAT 0.5763984 4.705205e-04 0.4548138 1.342556e-03
214 10434 LYPLA1 -1.2319618 1.398297e-03 -0.6131828 4.010407e-02
215 11075 STMN2 -4.7702715 1.699120e-02 -4.3532595 1.785601e-02
216 5375 PMP2 -1.3014515 1.685462e-45 -0.2556988 2.445849e-03
217 1345 COX6C 0.4021484 2.102484e-04 0.2373740 2.160260e-03
218 29028 ATAD2 -1.3181418 6.171433e-04 -0.8511346 1.138536e-02
219 114907 FBXO32 0.7491730 1.467707e-02 0.9139246 7.397547e-06
220 169044 COL22A1 1.0211618 8.035480e-03 0.9891097 1.371098e-03
221 340371 NRBP2 0.8880338 4.727206e-06 0.5516024 4.241716e-02
222 286343 LURAP1L -6.4879486 1.743746e-02 -6.4904645 1.246119e-02
223 11168 PSIP1 -0.7578417 4.857954e-05 -0.3471516 1.758778e-02
224 6194 RPS6 -0.4735103 1.393820e-07 -0.1948928 3.245436e-02
225 9568 GABBR2 1.8836534 3.992926e-19 1.7700974 1.993618e-26
226 10592 SMC2 -1.5510976 6.965430e-07 -0.7484702 1.290462e-02
227 2934 GSN 0.9575078 3.658505e-03 0.8546227 7.231943e-05
228 64855 NIBAN2 1.0028233 3.331896e-07 0.4470281 1.500068e-02
229 56262 LRRC8A 1.0105367 2.351635e-15 0.3698767 6.306425e-03
230 5730 PTGDS 1.3848729 1.610059e-13 0.7796929 2.800884e-05
231 4571 <NA> 0.9819432 6.551433e-07 0.8385856 2.779733e-06
232 27286 SRPX2 6.4021162 4.092541e-03 6.4458707 7.808378e-03
233 51186 TCEAL9 0.9494553 5.497590e-13 0.3992640 4.511053e-05
234 27018 BEX3 0.2307380 1.488369e-02 0.2425153 1.364090e-04
235 1641 DCX -0.8995534 1.452689e-07 -0.7023485 1.328081e-05
236 5358 PLS3 0.6066581 1.358155e-02 0.7933757 5.554753e-08
237 10178 TENM1 -1.2340255 5.833969e-04 -1.1143038 2.019523e-05Lade nötiges Paket: gprofiler2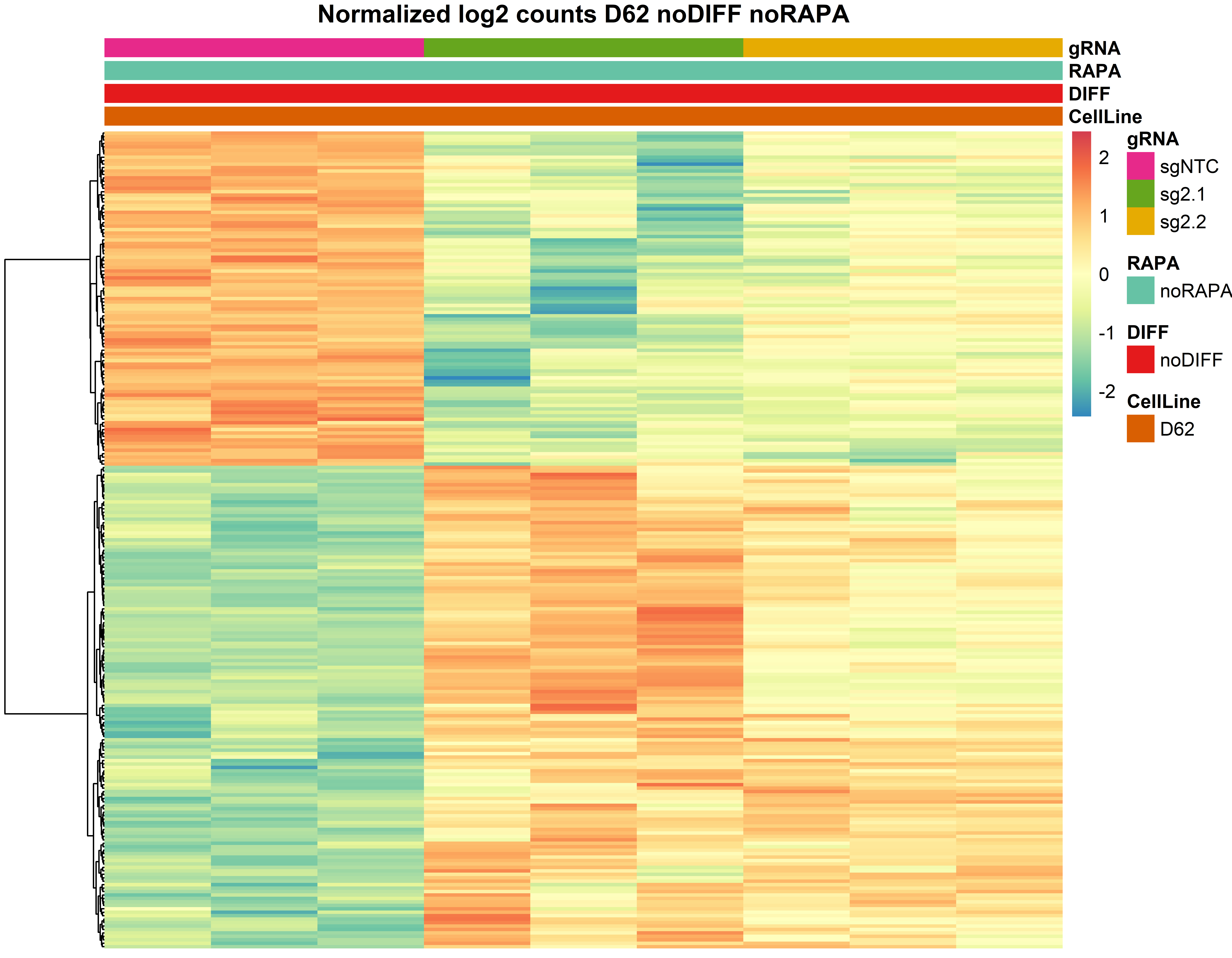
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.2877778 0.67089320 2.265422 3.358363e-01
02_TSC2_KO_Grabole_2016 1.7609750 1.34915108 2.296746 2.386415e-05
03_TSC2_Grabole-2016 1.9069744 1.44663760 2.529356 2.004220e-06
04_TSC2-tuber-Martin2017 3.8271742 1.48539266 8.292766 3.510059e-03
05_TSC2-SEN_SEGA-Martin2017 3.4990654 2.63839571 4.614603 2.952492e-17
06_all_EpilepsyGene 1.0615534 0.41907282 2.244720 8.409409e-01
07_high_conf_EpilepsyGene 0.9200612 0.10953205 3.432846 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 2.0229316 0.53726746 5.394379 1.458285e-01
09_Epi25_dataset 0.7008096 0.01745322 4.056387 1.000000e+00
10_SFARI_all 1.4771654 0.53098887 3.314241 3.117218e-01
11_SFARI_score4plus 1.3191449 0.35239855 3.485018 5.523534e-01
12_DeRubeis_RDNV 0.7379154 0.01836559 4.277341 1.000000e+00
13_Iossifov_RDNV 1.8942387 1.05548957 3.176803 2.242883e-02
14_FMRP_Darnell 1.4954934 0.87804477 2.409682 1.126561e-01
15_Voineagu_M12 0.4950929 0.10085139 1.477499 2.928806e-01
16_Voineagu_M16 2.8924093 1.60547796 4.875310 3.356070e-04
17_ID_genes_Neel 1.0612878 0.38257756 2.371358 8.287312e-01
18_ID_genes_pinto2014 0.4674959 0.05596242 1.726565 4.495240e-01
19_Fromer_SCZ_FunDN 1.4286915 0.72212593 2.569114 2.185662e-01
20_Cocchi_Pruning 3.3528738 1.05171443 8.246960 2.075668e-02
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.25291812 0.3326854 1.000000e+00
02_TSC2_KO_Grabole_2016 0.56586763 0.1359143 4.772831e-04
03_TSC2_Grabole-2016 0.64551788 0.1409571 4.008440e-05
04_TSC2-tuber-Martin2017 1.34212672 0.4828814 7.020117e-02
05_TSC2-SEN_SEGA-Martin2017 1.25249590 0.1440433 5.904983e-16
06_all_EpilepsyGene 0.05973332 0.4742061 1.000000e+00
07_high_conf_EpilepsyGene -0.08331512 1.0858280 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 0.70454776 0.6764321 1.000000e+00
09_Epi25_dataset -0.35551902 1.8840367 1.000000e+00
10_SFARI_all 0.39012496 0.5220098 1.000000e+00
11_SFARI_score4plus 0.27698373 0.6734573 1.000000e+00
12_DeRubeis_RDNV -0.30392608 1.8843625 1.000000e+00
13_Iossifov_RDNV 0.63881702 0.2983736 4.485766e-01
14_FMRP_Darnell 0.40245616 0.2716907 1.000000e+00
15_Voineagu_M12 -0.70300986 0.8117844 1.000000e+00
16_Voineagu_M16 1.06208981 0.3003410 6.712140e-03
17_ID_genes_Neel 0.05948303 0.5205647 1.000000e+00
18_ID_genes_pinto2014 -0.76036464 1.0830154 1.000000e+00
19_Fromer_SCZ_FunDN 0.35675900 0.3481198 1.000000e+00
20_Cocchi_Pruning 1.20981783 0.5915287 4.151336e-01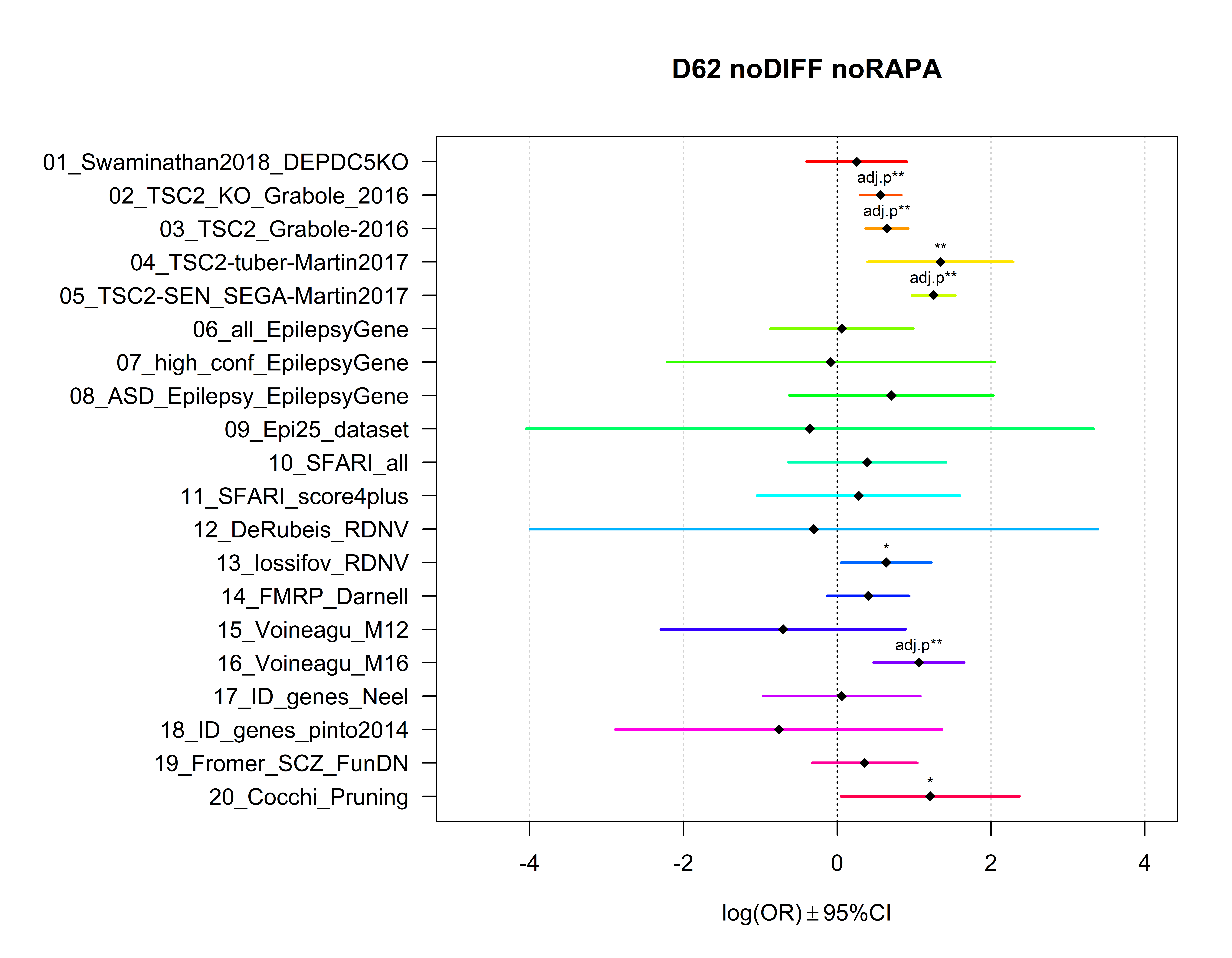
D244
getlinmodoutput(targetline = "D244",targetdiff = "noDIFF",targetrapa = "noRAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 54587 MXRA8 0.6447968 2.417795e-04 0.8182123 8.068229e-09
2 29101 SSU72 -0.9173927 2.763334e-03 -1.1082136 3.902258e-04
3 6497 SKI -0.8845508 3.016017e-07 -0.6570484 1.356475e-03
4 6146 RPL22 -0.3928134 4.692890e-03 -0.3572827 1.769757e-03
5 11315 PARK7 0.7491252 2.382166e-07 0.9060148 1.735854e-09
6 54206 ERRFI1 -1.1311270 4.743810e-04 -1.5296696 3.956574e-05
7 116362 RBP7 -2.2047409 4.864802e-02 -3.1121688 1.783481e-02
8 5226 PGD 0.5841230 2.843889e-03 0.5161229 7.892916e-03
9 23254 KAZN 1.2395553 8.332913e-08 0.9477767 1.053143e-03
10 23013 SPEN 0.9310611 4.471650e-04 0.8396755 3.787657e-03
11 84809 CROCCP2 -1.5358070 1.360397e-02 -1.9735378 3.140571e-02
12 5322 PLA2G5 0.8623372 3.016658e-04 0.7073064 1.489937e-03
13 55450 CAMK2N1 -0.4760729 1.275178e-03 -0.3857468 2.115330e-02
14 3399 ID3 1.7363641 2.589742e-06 1.8690969 1.464050e-08
15 6135 RPL11 -0.7132936 1.062736e-07 -0.4687796 2.108482e-03
16 25949 SYF2 -0.7167441 1.511816e-02 -0.8043068 1.057270e-03
17 57035 RSRP1 -0.6728268 5.240177e-03 -0.8006806 2.983479e-03
18 115572 TENT5B 5.7757449 1.505061e-02 5.7453215 4.484713e-02
19 2537 IFI6 1.5154830 8.533680e-13 0.7509509 2.270194e-06
20 93974 ATP5IF1 -0.6277827 6.324018e-05 -0.6097138 2.218699e-05
21 576 ADGRB2 1.5365554 5.780987e-05 1.0365313 3.807495e-02
22 3065 HDAC1 1.0545766 1.850515e-03 1.2706873 1.113050e-04
23 100128071 FAM229A 2.4783232 3.543597e-05 2.0452880 1.394958e-02
24 9202 ZMYM4 1.0336719 2.195512e-05 0.8845453 4.879582e-04
25 79647 AKIRIN1 -0.7141641 1.205169e-03 -0.8110174 1.386699e-04
26 643314 <NA> 2.0744004 3.544727e-06 2.7357402 2.704298e-11
27 65243 ZFP69B 2.8905770 4.750402e-02 3.2942760 2.060043e-02
28 59269 HIVEP3 -0.6210249 9.434418e-03 -0.4876863 2.914884e-02
29 4904 YBX1 -0.9467104 6.795124e-08 -0.5458329 8.100894e-03
30 6513 SLC2A1 -1.3114147 9.396373e-08 -0.5778601 8.917614e-03
31 9682 KDM4A 1.0221643 3.807963e-06 0.8188846 3.666041e-04
32 533 ATP6V0B -0.9324481 1.850625e-05 -0.6054301 5.726330e-03
33 7388 UQCRH -0.7643152 2.275698e-08 -0.5590674 1.200126e-03
34 9372 ZFYVE9 1.7869975 4.394748e-04 1.3874004 3.632698e-02
35 23318 TUT4 1.3570764 5.741589e-04 1.5929014 3.426310e-06
36 9528 TMEM59 -0.7735560 1.113853e-06 -1.0347099 2.281178e-11
37 51253 MRPL37 0.8619145 1.669471e-02 0.8493102 2.255944e-02
38 3725 JUN -0.8660046 1.153798e-10 -0.6198280 3.435749e-06
39 205 AK4 -2.6000671 2.034789e-17 -2.6459012 1.091889e-26
40 54741 LEPROT -0.5864790 1.389160e-02 -0.8981080 2.698948e-05
41 1647 GADD45A -1.3372732 1.155478e-14 -1.2436412 3.969845e-11
42 9295 SRSF11 -0.5021908 3.315065e-04 -0.6646528 2.036300e-06
43 257194 NEGR1 -0.5568808 2.555949e-02 -1.2006262 1.165253e-07
44 8880 FUBP1 0.9133557 2.452354e-02 1.0009577 1.840663e-02
45 64123 ADGRL4 -0.6100016 2.514503e-03 -0.3874754 3.217051e-02
46 2787 GNG5 -0.7878362 2.720673e-06 -0.5055382 4.065131e-03
47 3491 CCN1 -3.0068915 7.328821e-18 -1.6483774 9.005478e-07
48 9403 SELENOF -0.7382150 5.132402e-03 -0.6091142 3.288201e-02
49 2633 GBP1 2.0297059 2.515639e-02 2.0883616 2.907452e-02
50 7813 EVI5 0.9892861 3.443154e-03 1.1575739 3.326752e-04
51 6125 RPL5 -0.9418765 1.845797e-16 -0.7957400 1.667338e-09
52 9411 ARHGAP29 1.3663253 8.827226e-07 1.5754461 9.957316e-09
53 55119 PRPF38B -0.8389182 3.574626e-02 -1.2758728 2.312138e-08
54 6884 TAF13 2.7356784 5.595284e-13 2.1871193 1.826509e-05
55 84722 PSRC1 0.5366573 5.989141e-03 0.6938829 1.406718e-04
56 4893 NRAS -0.6826022 3.574626e-02 -0.7518060 2.671759e-02
57 4853 NOTCH2 -0.7896977 2.066373e-03 -0.8400555 5.425043e-04
58 9939 RBM8A 0.4701716 9.940688e-04 0.5115748 8.431092e-04
59 171423 <NA> 1.1888731 1.909574e-06 0.8145796 3.609831e-03
60 200030 NBPF11 -0.9055856 2.475034e-04 -0.6924015 9.653887e-03
61 51107 APH1A 1.2012888 3.693130e-24 1.0658835 2.574346e-18
62 29956 CERS2 -0.8834474 3.564355e-02 -0.9301237 4.595949e-03
63 10962 MLLT11 -1.5811599 3.095660e-09 -1.6835795 2.362632e-12
64 8991 SELENBP1 1.2547867 1.093970e-03 1.2207704 1.173326e-02
65 6281 S100A10 -1.9846326 3.813423e-18 -1.3322505 1.272432e-05
66 6282 S100A11 -0.6564485 1.243054e-04 -0.5045169 1.029874e-02
67 5872 RAB13 -0.5469641 2.903671e-03 -0.5719559 6.150246e-04
68 3782 KCNN3 1.1943319 2.086004e-04 1.2761001 1.077648e-03
69 54344 DPM3 1.0757316 2.723404e-05 1.0672900 1.201889e-04
70 23623 RUSC1 1.0832154 1.840788e-03 1.1752003 2.248440e-03
71 55870 ASH1L 0.9321681 9.934063e-06 0.9727356 1.165253e-07
72 112770 GLMP 0.7879062 3.886354e-02 1.1774971 3.818671e-05
73 10485 MIR9-1HG 1.1489320 8.257954e-11 1.6085905 9.168998e-20
74 63827 BCAN 1.8394101 8.450589e-03 1.8162346 6.072456e-03
75 10763 NES 0.3475318 2.516866e-03 0.6871561 1.662445e-11
76 8407 TAGLN2 0.8636301 5.565527e-03 1.0888752 9.437931e-05
77 477 ATP1A2 0.2746303 5.643269e-03 0.2505707 1.451028e-02
78 844 CASQ1 2.5185491 8.954515e-03 2.2659897 3.825698e-02
79 223 ALDH9A1 0.4402993 2.846024e-03 0.3789553 1.585687e-02
80 481 ATP1B1 0.7427634 3.407070e-09 1.5134022 1.486661e-46
81 9910 RABGAP1L 1.0777722 2.392776e-02 1.3284105 3.013888e-03
82 27101 CACYBP -0.8641518 4.606050e-03 -0.9597235 1.451028e-02
83 460 ASTN1 -0.5115954 1.305351e-02 -1.0060898 4.272994e-06
84 9857 CEP350 1.0052533 2.712106e-04 0.9324754 3.608563e-03
85 2752 GLUL -0.4312720 5.544555e-03 -0.5914626 2.878473e-04
86 10092 ARPC5 -0.6949867 1.364559e-05 -0.3897436 2.723603e-02
87 116496 NIBAN1 1.5849445 5.983495e-18 2.2824818 2.588458e-38
88 7175 TPR 0.8931670 3.382992e-06 0.8617858 2.095923e-05
89 9928 KIF14 2.2993429 4.224364e-02 2.2651655 4.949889e-02
90 6635 SNRPE -0.8584755 2.925472e-03 -0.6879693 4.496959e-02
91 64710 NUCKS1 0.9234332 8.264264e-09 0.6271025 3.258650e-11
92 947 CD34 6.8539720 2.178733e-03 6.8519457 7.351875e-03
93 5362 PLXNA2 1.6424306 2.145378e-07 1.5953150 5.867470e-07
94 3756 KCNH1 -7.3468321 3.699570e-04 -5.4322710 8.750087e-03
95 127018 LYPLAL1 -0.9879165 1.100170e-02 -0.8062422 4.027357e-02
96 6726 SRP9 -1.4157014 1.111334e-11 -1.0645445 7.213889e-08
97 2052 EPHX1 1.6046594 7.584594e-07 1.6860167 1.235717e-05
98 8476 CDC42BPA 0.6142316 1.179130e-03 0.5479464 1.451028e-02
99 2590 GALNT2 -0.7731319 6.670422e-05 -0.7149193 7.456542e-04
100 57568 SIPA1L2 0.9257859 1.890555e-04 0.6949482 2.554943e-02
101 9804 TOMM20 -0.4554094 2.820720e-02 -0.4458038 3.795679e-02
102 4811 NID1 -0.7110318 4.728859e-02 -1.2334604 3.579051e-03
103 116228 COX20 -1.8357792 2.152770e-07 -1.6485115 1.657246e-06
104 5209 PFKFB3 -1.3459402 2.238208e-09 -1.1345488 1.047416e-07
105 10659 CELF2 0.9977678 5.541618e-09 0.7432419 5.406799e-04
106 10006 ABI1 -0.7545308 2.849425e-02 -1.0733793 1.546463e-02
107 22931 RAB18 -0.5623941 1.961020e-02 -0.6927329 1.694852e-02
108 8829 NRP1 -0.8238021 2.178641e-05 -1.6445711 7.436750e-14
109 56288 PARD3 -1.0587320 2.799879e-03 -0.8265191 6.534399e-04
110 3185 HNRNPF -0.6858026 6.083684e-04 -0.6222294 1.412559e-02
111 220992 ZNF485 -6.3580771 1.320149e-02 -6.3600448 1.778639e-02
112 8031 NCOA4 -0.8830717 1.209904e-09 -0.8563953 6.575711e-08
113 288 ANK3 1.7275115 2.718835e-04 1.7514353 3.440427e-04
114 1959 EGR2 -3.0778980 1.216571e-06 -2.0601500 1.416154e-03
115 221035 REEP3 0.5389792 6.107508e-03 0.6018794 1.858649e-03
116 3189 HNRNPH3 -0.6417893 2.411780e-04 -1.0536549 2.235016e-07
117 80312 TET1 1.6967383 2.491949e-08 1.0827774 1.765652e-02
118 55749 CCAR1 -0.6191236 4.013120e-03 -0.8670215 3.794006e-06
119 9806 SPOCK2 1.4613572 3.499828e-09 0.9301164 1.207277e-03
120 5033 P4HA1 -1.7335366 3.370544e-12 -1.4498884 3.303972e-20
121 51021 MRPS16 0.7786055 9.652363e-05 0.5836621 5.868213e-03
122 7417 VDAC2 -0.6988929 9.794086e-03 -0.5341689 1.534269e-02
123 6229 RPS24 -0.6916832 3.106627e-10 -0.5704100 4.402772e-07
124 143244 EIF5AL1 1.5659324 5.049334e-03 1.6170947 5.558383e-03
125 2894 GRID1 -1.1036645 2.294590e-02 -1.3336066 4.951310e-04
126 27063 ANKRD1 -5.5137933 2.042192e-06 -8.6731657 7.834159e-08
127 80351 TNKS2 1.0299364 9.853310e-04 0.9336473 1.112778e-04
128 9849 ZNF518A 1.2378271 1.049081e-04 1.0746867 4.907204e-03
129 81894 SLC25A28 1.6968344 1.867188e-03 2.1183528 1.720933e-05
130 282991 BLOC1S2 -0.9321229 1.726778e-05 -0.9034756 2.252307e-04
131 55719 SLF2 1.2106297 6.250822e-20 1.0480260 2.875912e-10
132 6468 FBXW4 0.7390090 2.525435e-02 0.7973168 1.758357e-02
133 22984 PDCD11 0.8541384 2.583306e-02 0.9151997 2.759905e-02
134 374354 NHLRC2 1.3669600 4.767746e-02 1.8331192 6.306577e-03
135 340719 NANOS1 -1.3795954 1.639025e-02 -1.7742895 2.723133e-03
136 404636 DENND10 -0.9829849 1.678137e-04 -0.6144156 4.018666e-02
137 10935 PRDX3 -0.7389465 1.762494e-05 -0.5166271 2.564062e-03
138 2869 GRK5 -0.3854062 2.781705e-02 -0.4515866 8.548647e-03
139 9531 BAG3 -0.8898873 4.364275e-04 -0.7296086 2.123028e-03
140 10579 TACC2 0.9597943 3.319996e-03 0.8975968 4.173990e-02
141 5654 HTRA1 1.2390580 4.298624e-48 0.9493183 3.645379e-31
142 56647 BCCIP 0.9776739 1.284575e-03 0.7147603 4.706219e-02
143 664 BNIP3 -2.6809234 3.831802e-32 -2.5143484 5.095257e-33
144 5719 PSMD13 0.7484132 1.997567e-03 0.7823334 1.307554e-03
145 10581 IFITM2 1.8767842 2.456134e-14 1.0570806 1.018303e-02
146 100506211 MIR210HG -8.2586434 4.188804e-07 -8.2608220 7.402432e-07
147 6888 TALDO1 0.3771442 3.508497e-02 0.4072823 1.748263e-02
148 977 CD151 0.4651951 1.184530e-03 0.4485379 5.268446e-03
149 1509 CTSD 0.6420202 1.122989e-05 0.7755858 1.678842e-10
150 1200 TPP1 0.3684342 1.225323e-02 0.4018636 5.869324e-03
151 4004 LMO1 1.4755595 2.239385e-17 0.9912803 4.247282e-06
152 133 ADM -1.1528000 6.884112e-10 -0.8817726 8.411273e-07
153 100463486 MTRNR2L8 2.0760591 4.973333e-99 1.6383439 4.273056e-66
154 1982 EIF4G2 0.3431180 1.393095e-02 0.2990074 2.085604e-02
155 1315 COPB1 -0.7498241 9.088462e-03 -0.7499982 1.076310e-02
156 5682 PSMA1 -1.1672370 2.712690e-05 -0.7845991 5.267303e-04
157 55553 SOX6 0.6675824 1.383113e-03 0.5561424 8.674927e-03
158 6207 RPS13 -0.5854896 2.454053e-06 -0.3564525 8.678934e-03
159 3939 LDHA -3.4833879 5.917882e-07 -3.9180145 1.520274e-64
160 89797 NAV2 -1.9991799 7.869335e-07 -1.4428593 1.924487e-04
161 55366 LGR4 -0.6078966 1.919318e-03 -0.7625720 2.118318e-04
162 10480 EIF3M -1.2875213 3.990990e-02 -1.5846814 1.320877e-02
163 79832 QSER1 1.0949413 5.265493e-03 0.9822547 2.085115e-02
164 966 CD59 -0.4744186 3.538368e-03 -0.4490270 5.599200e-03
165 4005 LMO2 -0.7454562 1.728969e-06 -1.2056432 1.324486e-13
166 25841 ABTB2 -2.9561254 4.556237e-18 -2.2965434 8.098361e-17
167 24147 FJX1 -1.4503240 2.971714e-14 -1.3802161 1.799033e-14
168 2132 EXT2 0.9578585 4.364275e-04 1.3618098 6.821122e-08
169 3732 CD82 0.9955247 3.179847e-03 1.4382645 1.762720e-06
170 9793 CKAP5 1.3396702 6.779571e-05 0.8674058 1.371972e-02
171 4038 LRP4 -0.6929678 1.501417e-02 -0.5339822 4.390481e-02
172 53 ACP2 1.2286673 8.368271e-07 1.0066536 1.233090e-03
173 5795 PTPRJ -1.3350452 2.281039e-09 -1.0137659 6.996018e-05
174 710 SERPING1 1.9340720 4.385201e-20 1.1556381 8.334356e-05
175 219988 PATL1 -0.8415953 1.961020e-02 -1.8290073 3.685195e-05
176 2495 FTH1 -0.8232078 1.133042e-16 -0.7118784 3.813248e-12
177 51035 UBXN1 0.6918226 1.194236e-03 0.7230134 3.434057e-03
178 10963 STIP1 0.6548338 1.354956e-02 0.6768599 2.446924e-02
179 7423 VEGFB -0.8179653 1.047089e-04 -0.7177995 7.969386e-03
180 9379 NRXN2 1.4451059 7.683417e-14 0.9673922 9.412433e-04
181 823 CAPN1 1.0471023 2.863462e-04 1.0289059 2.279615e-04
182 11007 CCDC85B 0.5987630 1.440810e-02 0.7023948 1.763983e-03
183 10992 SF3B2 0.4417597 1.790684e-02 0.6146819 1.715440e-04
184 81876 RAB1B 0.8176175 3.056053e-02 0.8925297 1.230919e-02
185 9973 CCS 1.2196574 2.872499e-05 1.0860124 1.244910e-03
186 80194 TMEM134 0.6108532 4.654633e-03 0.5198541 1.739753e-02
187 2950 GSTP1 0.5971709 8.296952e-06 0.5144516 2.118318e-04
188 1374 CPT1A 0.9761002 3.474505e-21 0.8179089 1.851823e-16
189 116985 ARAP1 1.2395372 3.364267e-02 1.6189185 2.408494e-03
190 23201 FAM168A -0.6428070 2.296398e-02 -0.8234691 2.532603e-03
191 58473 PLEKHB1 0.5634513 1.372037e-02 0.6180136 2.117641e-02
192 408 ARRB1 1.7228956 8.174253e-03 1.7450086 9.636809e-03
193 6188 RPS3 0.6341066 1.870823e-07 0.7436506 5.681190e-10
194 5547 PRCP -1.3820226 5.954745e-23 -1.1224384 3.896844e-18
195 1740 DLG2 1.2647359 1.079473e-10 1.0743873 2.641466e-07
196 58487 CREBZF -1.1063279 1.804999e-03 -0.8379902 1.726749e-02
197 619383 SCARNA9 4.5607292 4.140064e-48 4.3806064 1.013526e-41
198 51503 CWC15 -0.8597363 9.142555e-04 -0.6153302 1.478114e-02
199 143686 SESN3 1.5739109 1.216622e-17 1.0316799 6.585544e-06
200 8898 MTMR2 -1.4221861 1.160066e-03 -1.3564063 6.961040e-04
201 80310 PDGFD -3.7537703 3.356242e-05 -1.6779752 3.840022e-02
202 5805 PTS -0.8402808 3.945673e-03 -0.9625137 5.917697e-03
203 4684 NCAM1 -1.0844064 6.229755e-04 -0.9184914 1.385297e-03
204 23705 CADM1 -1.2238272 4.691888e-23 -0.9632451 1.309892e-18
205 6330 SCN4B 1.6567324 1.733395e-06 1.6734741 1.432735e-07
206 10632 ATP5MG -0.4537306 5.381304e-04 -0.3536292 1.254254e-02
207 6230 RPS25 -0.8169508 3.441897e-08 -0.7294416 1.786325e-06
208 51399 TRAPPC4 -1.2063504 1.910954e-07 -1.0159225 6.123254e-05
209 6309 SC5D -1.7740606 1.143566e-13 -1.2179568 2.871699e-07
210 3312 HSPA8 -0.6503167 1.151615e-09 -0.3512107 5.901605e-04
211 9538 EI24 0.4420814 1.193928e-02 0.6816923 8.210376e-06
212 9743 ARHGAP32 -1.9969232 4.245443e-02 -1.4621260 3.491379e-02
213 50863 NTM 1.9202919 6.351849e-13 1.2243360 2.086549e-02
214 27087 B3GAT1 1.4140518 2.316604e-09 1.1640871 5.957845e-06
215 894 CCND2 0.4996225 2.743922e-04 0.6658611 6.640099e-10
216 4908 NTF3 -6.1515947 3.993723e-02 -6.1536348 4.936620e-02
217 928 CD9 -1.4717770 1.147028e-23 -1.7946156 2.531330e-31
218 7167 TPI1 -0.7463220 5.863670e-05 -0.7067538 4.273139e-04
219 2026 ENO2 -0.9007238 9.258185e-09 -0.6579910 2.930540e-04
220 1822 ATN1 -1.3915260 1.279368e-06 -1.2751632 3.949562e-06
221 715 C1R 0.9070775 1.895855e-11 1.0631194 4.060994e-11
222 6515 SLC2A3 -0.7474001 8.467466e-03 -0.7162774 9.403404e-03
223 8531 YBX3 -0.7778992 2.458787e-05 -0.5092774 1.087899e-02
224 51202 DDX47 -1.3307240 3.925541e-02 -1.3989661 2.478382e-02
225 2012 EMP1 -1.9226605 1.040745e-08 -1.5446940 1.635205e-09
226 55729 ATF7IP 0.9243519 1.184530e-03 1.2911304 2.936539e-07
227 5800 PTPRO -6.9596645 1.859337e-03 -6.9616527 2.648940e-03
228 11171 STRAP -0.5301000 4.645608e-02 -0.5421234 2.922212e-02
229 51026 GOLT1B -1.6521299 2.653729e-03 -1.0956044 2.030822e-02
230 3709 ITPR2 1.1914972 5.026342e-03 1.1596578 2.568866e-03
231 55297 CCDC91 -0.9491800 5.115343e-03 -0.8213005 7.732515e-03
232 441631 TSPAN11 1.1981092 2.743922e-04 1.5514002 2.602915e-07
233 636 BICD1 0.8302369 1.420847e-04 0.8885961 1.528456e-06
234 85437 ZCRB1 -0.7997047 1.833369e-04 -0.6686395 6.600812e-03
235 51564 HDAC7 -0.9381597 3.820831e-03 -0.8288939 2.974436e-02
236 1280 COL2A1 -2.0440771 3.180110e-05 -2.2969663 9.941474e-07
237 9416 DDX23 0.8370554 3.846728e-03 1.1092867 5.816490e-04
238 7009 TMBIM6 0.5951983 2.983066e-11 0.7133874 7.518873e-17
239 51474 LIMA1 -1.1029540 5.398832e-09 -0.6859229 2.347144e-03
240 57609 DIP2B 1.1206196 5.326370e-06 0.7783899 6.304220e-03
241 25840 METTL7A 1.1450889 2.028174e-13 0.8840038 1.737575e-06
242 81566 CSRNP2 -1.2118124 3.519716e-04 -1.0831552 1.874409e-03
243 9802 DAZAP2 -0.4733097 9.632950e-03 -0.4904441 1.294316e-02
244 160622 TAMALIN 1.1670602 5.849039e-03 1.1129331 4.059283e-02
245 2647 BLOC1S1 -0.9361823 1.063945e-02 -1.2069707 5.308265e-03
246 84324 SARNP 1.9850861 3.392792e-05 1.2924817 2.014432e-02
247 1606 DGKA -1.3416105 6.889470e-03 -1.2672936 1.706826e-02
248 4637 MYL6 -0.6208233 1.054238e-06 -0.4216945 4.501027e-03
249 10728 PTGES3 -0.7317474 3.951503e-05 -0.8735440 1.688367e-06
250 4666 NACA -0.9402961 1.195302e-09 -0.8107363 1.007170e-06
251 6472 SHMT2 1.1374543 7.009110e-03 1.4197326 1.260041e-04
252 22864 R3HDM2 1.0575085 1.864084e-03 0.9101495 3.574395e-02
253 10106 CTDSP2 -0.5228011 1.697879e-02 -0.6071262 9.827096e-03
254 4193 MDM2 0.5162521 1.420183e-03 0.5722706 9.204256e-06
255 4673 NAP1L1 -0.6268512 9.779490e-05 -0.5091138 2.586268e-03
256 1466 CSRP2 -0.8656306 2.940261e-06 -0.5731308 4.681778e-03
257 8411 EEA1 1.1609643 2.052218e-04 1.2419375 2.770534e-04
258 57458 TMCC3 2.4466383 3.053042e-02 2.7869660 1.660474e-02
259 6636 SNRPF -0.6886123 9.067862e-03 -0.8460276 1.302775e-03
260 23074 UHRF1BP1L 1.9891039 1.292456e-02 2.0868688 2.747768e-03
261 79158 GNPTAB 1.0095934 5.262357e-04 1.0638101 1.487775e-03
262 7184 HSP90B1 -0.8297707 2.472046e-10 -1.0026424 5.506803e-15
263 10970 CKAP4 -0.5231204 1.963472e-04 -0.6250150 5.957845e-06
264 121642 ALKBH2 1.1881260 1.508488e-03 1.0742132 9.059472e-03
265 10094 ARPC3 -0.7523080 1.326844e-06 -0.8765033 2.160986e-07
266 84329 HVCN1 4.0387920 2.008063e-04 4.4157216 8.959148e-05
267 10961 ERP29 -0.5296490 1.171193e-03 -0.4733213 1.011875e-02
268 643246 MAP1LC3B2 -1.1693202 1.726802e-04 -1.1514770 5.587250e-04
269 6175 RPLP0 -0.7427281 3.438281e-20 -0.2784224 3.519763e-03
270 4440 MSI1 1.3726026 3.957070e-04 1.0955685 7.351875e-03
271 8655 DYNLL1 -0.4910178 1.727812e-04 -0.6008993 1.239605e-05
272 9921 RNF10 0.7004251 7.923407e-04 0.6981440 9.754433e-04
273 9761 MLEC 0.4683059 1.188779e-03 0.5653529 4.661108e-05
274 55596 ZCCHC8 -1.7482099 1.961020e-02 -2.4142500 9.188144e-04
275 79720 VPS37B 0.9026088 1.485030e-03 1.1126039 1.197114e-06
276 51329 ARL6IP4 0.5401940 1.790684e-02 0.6683972 3.609831e-03
277 8099 CDK2AP1 -0.4985766 1.066706e-05 -0.4413100 2.049964e-04
278 9612 NCOR2 -0.7234303 5.381304e-04 -0.8248144 2.936539e-07
279 7316 UBC 0.3627002 2.787712e-03 0.3310509 1.899220e-03
280 2054 STX2 0.8726497 3.914194e-02 1.1685108 8.049816e-03
281 54737 MPHOSPH8 -0.6093420 7.666844e-03 -0.5502569 1.491265e-03
282 26278 SACS 1.0417896 1.649009e-03 1.0018200 3.940864e-03
283 221178 SPATA13 -1.2239684 8.945893e-09 -0.9779087 8.192199e-05
284 219287 AMER2 0.8334604 3.063447e-13 0.7072583 6.087483e-08
285 675 BRCA2 4.8437293 6.303308e-05 4.4783132 5.153121e-03
286 9201 DCLK1 1.0552536 2.276876e-14 0.9505832 6.874839e-10
287 341640 FREM2 0.9575750 3.226430e-03 0.8133285 1.858271e-02
288 10186 LHFPL6 -0.4650645 1.790684e-02 -0.8595817 7.458090e-05
289 11193 WBP4 -1.0962585 4.413549e-06 -0.9162024 3.088599e-04
290 11215 AKAP11 0.8422400 7.932182e-04 1.2606830 3.607048e-09
291 55068 ENOX1 -2.4177415 1.540034e-02 -3.8431654 2.452882e-04
292 7178 TPT1 -1.0430667 9.296532e-39 -0.7052410 1.074257e-16
293 9445 ITM2B -0.5852653 4.418694e-11 -0.6239563 6.611768e-12
294 51131 PHF11 1.5021135 7.180181e-06 1.8957506 6.093754e-08
295 8847 DLEU2 2.0522810 6.337342e-03 1.7427843 4.830400e-02
296 27253 PCDH17 -0.9566323 2.100556e-05 -1.1748425 5.231539e-08
297 122060 SLAIN1 -0.8106943 2.416954e-02 -1.1421408 2.583238e-03
298 5611 DNAJC3 1.1874172 1.434827e-09 0.7536076 4.266996e-03
299 5095 PCCA 1.5895316 3.052919e-04 2.4264321 1.685252e-13
300 55608 ANKRD10 -0.8445098 3.226430e-03 -0.9948470 1.119566e-03
301 55002 TMCO3 -0.7261552 2.866052e-02 -0.7889202 1.811970e-02
302 2621 GAS6 1.6850483 6.409206e-12 0.8078906 1.974866e-02
303 6038 RNASE4 -1.9302768 2.011118e-06 -1.2025272 8.033371e-04
304 57680 CHD8 0.9395286 4.839307e-05 0.8730603 2.538096e-03
305 4323 MMP14 1.0612673 4.393978e-11 1.3284949 1.962593e-17
306 10278 EFS 1.9268720 1.554770e-05 1.6053783 3.176664e-03
307 85446 ZFHX2 -1.5696793 1.028774e-06 -1.6434051 3.928918e-07
308 5720 PSME1 0.5950205 6.116031e-03 0.5471449 2.532466e-02
309 9985 REC8 0.8750887 2.357235e-03 0.8016210 8.410037e-03
310 26277 TINF2 0.7204108 3.873351e-03 0.6246591 2.328111e-02
311 29966 STRN3 1.3153498 1.658088e-07 1.5434901 5.395941e-11
312 25831 HECTD1 0.7356891 1.590851e-04 0.8104926 7.598210e-05
313 394 ARHGAP5 0.7443182 8.323117e-08 0.6647278 1.711393e-05
314 55012 PPP2R3C 0.5379489 3.657829e-02 0.5484802 4.693184e-02
315 5411 PNN -0.6463694 4.180690e-04 -0.7190232 4.968118e-04
316 23116 TOGARAM1 0.7574602 3.998225e-02 0.9200662 3.263452e-02
317 161357 MDGA2 1.1942387 1.678550e-04 0.9083236 1.928000e-02
318 6235 RPS29 0.4490748 2.362414e-06 0.5278008 1.321150e-07
319 6166 RPL36AL -0.9125391 8.257954e-11 -0.7907283 3.761478e-07
320 378706 RN7SL2 2.6170035 2.258347e-08 2.9161589 7.399495e-10
321 81542 TMX1 -0.9419960 1.845171e-05 -0.9141248 4.101185e-04
322 3895 KTN1 0.2886748 4.103657e-02 0.4045997 8.290912e-04
323 5684 PSMA3 -0.7017538 1.859337e-03 -0.8006992 1.806997e-04
324 51528 JKAMP -1.3726262 6.028115e-06 -1.2066665 1.475236e-06
325 6252 RTN1 -0.2886628 2.349734e-03 -0.4964596 4.923798e-08
326 51635 DHRS7 -0.8234558 1.673959e-04 -0.8150266 1.470560e-05
327 3091 HIF1A 1.8493151 2.503885e-17 1.8257181 3.576167e-15
328 23224 SYNE2 1.8616176 7.117427e-27 1.3688272 3.734839e-10
329 87 ACTN1 0.4284562 2.187020e-03 0.4684115 6.632582e-04
330 64093 SMOC1 0.9866785 6.718799e-19 0.6051735 2.779479e-05
331 8110 DPF3 -0.9789436 2.551990e-06 -0.7434020 6.008153e-03
332 8650 NUMB -0.9293603 6.133586e-03 -1.0166026 1.684982e-02
333 9240 PNMA1 -0.8285234 4.396808e-07 -0.5001593 4.612193e-02
334 4329 ALDH6A1 2.3638605 2.653510e-21 2.0569067 1.121320e-14
335 10972 TMED10 -0.9266083 7.998204e-17 -0.7999917 3.772776e-15
336 9517 SPTLC2 1.2379555 1.118082e-04 1.0955649 2.737931e-03
337 9369 NRXN3 -2.0815225 2.496293e-04 -2.9810474 1.170221e-06
338 5700 PSMC1 0.8246297 1.257916e-02 1.0463330 7.027737e-05
339 145567 TTC7B -0.9885643 2.732516e-02 -1.3407013 2.076992e-03
340 4707 NDUFB1 0.4369686 1.360353e-03 0.4696018 4.131010e-04
341 53981 CPSF2 0.7762509 1.909529e-02 0.9830912 1.315135e-03
342 5641 LGMN -0.6970986 2.561799e-03 -0.8213816 6.702576e-04
343 51466 EVL 0.6641817 1.624341e-02 0.6069795 4.979809e-02
344 8788 DLK1 -0.7610714 2.285698e-05 -1.7891293 4.347742e-23
345 3320 HSP90AA1 -0.9351361 7.328821e-18 -0.6908205 9.028993e-10
346 1983 EIF5 -0.6241797 5.686856e-03 -0.5320810 1.339059e-02
347 1152 CKB 1.0646996 7.958010e-24 0.7029645 1.555062e-09
348 1397 CRIP2 1.1480773 6.021030e-23 1.0006637 5.950226e-18
349 646214 <NA> 0.9896756 2.960410e-02 1.0235084 1.859654e-02
350 347746 <NA> 1.0173491 3.294376e-02 1.0088941 4.306992e-02
351 440248 HERC2P9 -2.0141815 3.196005e-04 -1.9771477 1.353172e-03
352 55505 NOP10 0.5100915 2.345297e-02 0.5693931 1.279535e-02
353 57082 KNL1 4.3518578 1.138427e-03 3.9533849 1.300403e-02
354 51332 SPTBN5 -1.5194173 9.394863e-03 -1.6763140 1.201892e-03
355 30844 EHD4 -0.5592482 3.523082e-02 -0.6276293 9.653887e-03
356 25963 TMEM87A -0.5712014 1.596924e-02 -0.4683078 3.551897e-02
357 64397 ZNF106 0.5888948 5.531800e-04 0.5229948 3.609831e-03
358 2628 GATM 1.0641065 8.803533e-06 0.8997106 3.609831e-03
359 2200 FBN1 0.8228623 4.586825e-02 0.9472631 6.130884e-05
360 9728 SECISBP2L 0.9054933 6.477263e-03 0.7180537 5.226021e-03
361 55930 MYO5C 1.6161840 4.872646e-03 1.9094068 8.189179e-04
362 4644 MYO5A 0.7821074 2.849359e-03 1.0527000 1.007170e-06
363 51187 RSL24D1 -0.7582322 4.833167e-02 -0.7211120 3.538759e-02
364 6938 TCF12 -0.7119689 1.361048e-02 -0.8529075 4.335729e-03
365 102 ADAM10 -1.0435933 2.176227e-07 -0.9847376 2.563821e-06
366 79811 SLTM -0.5261474 2.957613e-02 -0.7266043 3.961070e-04
367 2958 GTF2A2 -1.1390753 2.667342e-08 -1.1163038 2.828873e-04
368 302 ANXA2 -0.6174623 7.459776e-12 -0.6791950 4.316901e-13
369 54832 VPS13C 0.9913565 4.323899e-04 0.9832413 3.418495e-04
370 83660 TLN2 0.7075094 5.504391e-06 0.5277677 1.053720e-02
371 8925 HERC1 1.0538969 5.231586e-03 1.0321122 1.520674e-02
372 51324 SPG21 -0.7068337 5.191695e-04 -0.8068551 1.282304e-03
373 8766 RAB11A -0.6927286 1.137117e-04 -0.8830035 3.602168e-04
374 84465 MEGF11 -1.3726201 2.327044e-05 -1.5206484 1.603060e-03
375 6124 RPL4 -0.5214030 6.734381e-04 -0.4077637 1.088155e-02
376 10391 CORO2B 1.2558827 3.283149e-21 1.0136424 8.558249e-14
377 5315 PKM -0.7939105 3.670433e-11 -0.3837185 2.702039e-02
378 4756 NEO1 0.5201654 1.160066e-03 0.7084842 2.854245e-06
379 27020 NPTN -0.7695636 1.767261e-06 -0.7797756 1.423627e-07
380 3671 ISLR -0.2833417 2.028597e-02 -0.3222841 3.332679e-02
381 9377 COX5A -0.6967445 3.494099e-05 -0.5349109 1.390013e-02
382 5780 PTPN9 -0.8448890 1.322620e-03 -0.6038163 2.040272e-02
383 5955 RCN2 -0.8584015 1.643613e-05 -0.6520988 5.948056e-03
384 55466 DNAJA4 5.2930987 1.018488e-02 5.3991296 1.109825e-03
385 10933 MORF4L1 -0.5568219 6.649373e-05 -0.6284132 1.108807e-03
386 1512 CTSH 2.2502971 2.188462e-31 1.1993025 9.204256e-06
387 23423 TMED3 -2.5221986 2.083008e-04 -1.5214805 4.883821e-04
388 2184 FAH 1.3692678 4.251650e-02 1.4510803 3.151168e-02
389 100996492 CTXND1 -1.2594438 2.747066e-09 -0.6366117 1.639764e-02
390 58489 ABHD17C -0.7203873 6.935918e-03 -1.0637108 5.587250e-04
391 57214 CEMIP -3.9315748 1.063531e-02 -2.0436591 3.779308e-07
392 83640 RAMAC -2.5731740 4.573590e-02 -4.6556532 1.466011e-04
393 388165 UBE2Q2P1 2.1997492 1.586437e-03 2.0978713 3.694621e-02
394 4828 NMB 1.2409236 1.168253e-03 1.4003827 2.182028e-04
395 11057 ABHD2 1.0991177 1.173058e-11 1.0141111 1.555218e-09
396 3418 IDH2 1.0091847 7.629644e-03 0.9807063 1.950107e-02
397 8826 IQGAP1 0.6676281 2.298051e-04 0.5221466 4.544405e-03
398 5045 FURIN 0.7669524 6.354729e-03 0.8715731 3.469704e-03
399 100507217 <NA> -0.8697019 7.994831e-10 -0.9408703 1.569851e-09
400 56963 RGMA 0.6604419 6.084450e-04 0.4291084 2.922212e-02
401 80213 TM2D3 -1.0519095 3.625704e-04 -1.6898388 3.096144e-06
402 10573 MRPL28 1.1499369 3.029710e-02 1.2957882 1.485419e-02
403 58986 PGAP6 -0.8998415 1.804999e-03 -0.8592389 1.048363e-03
404 146330 FBXL16 -0.7899244 2.654058e-02 -1.2690415 4.662999e-04
405 79006 METRN 0.6003464 7.760361e-06 0.5046881 4.699858e-04
406 65990 ANTKMT 1.8619041 2.452679e-05 1.7352076 3.035851e-04
407 4832 NME3 0.9375875 2.140022e-04 0.6623158 3.996775e-02
408 735301 <NA> 1.0597505 8.099778e-04 0.9979261 8.293104e-03
409 5310 PKD1 -1.0304686 3.192947e-02 -1.1070155 2.521897e-02
410 527 ATP6V0C 0.4370069 3.907192e-04 0.4542608 8.966766e-04
411 54442 KCTD5 -1.4562156 2.964474e-03 -0.9402968 1.021544e-02
412 54985 HCFC1R1 -0.6079132 3.991626e-02 -0.6490186 2.634717e-02
413 54700 RRN3 -1.2351659 4.128261e-03 -0.8604663 3.409937e-02
414 6210 RPS15A -0.5960953 1.211274e-05 -0.4428597 1.003147e-03
415 23204 ARL6IP1 0.9383252 6.006087e-11 0.9058792 1.158777e-10
416 162073 ITPRIPL2 1.1726156 7.658727e-05 1.5954294 7.109072e-13
417 124152 IQCK -0.5657486 5.628584e-05 -0.5589493 1.117374e-04
418 100271836 <NA> 3.0737938 9.292262e-05 2.4881162 7.305543e-03
419 440345 NPIPB4 -1.5582785 2.743922e-04 -1.6215592 1.854642e-04
420 100132247 NPIPB5 -1.6525869 2.696048e-03 -1.5490723 8.293104e-03
421 4706 NDUFAB1 -0.5472785 6.751972e-04 -0.4580337 4.470866e-03
422 84516 DCTN5 0.8826744 2.374119e-02 1.2467702 1.967691e-03
423 5930 RBBP6 -1.2887842 3.735406e-08 -1.4071110 1.200944e-09
424 84901 NFATC2IP 1.0847510 1.773855e-02 1.1231429 2.177698e-02
425 83985 SPNS1 1.2340362 2.104477e-03 1.5747545 1.257186e-05
426 728888 NPIPB11 -2.8252509 3.901370e-08 -2.1850084 5.952707e-07
427 9961 MVP 0.8920615 7.547243e-05 1.0620417 2.463593e-06
428 283899 INO80E 2.1379343 1.465428e-03 2.0649537 4.561370e-03
429 226 ALDOA -1.5444400 1.280511e-36 -1.0558203 4.194413e-14
430 9274 BCL7C 1.0513226 5.261210e-04 1.4388208 1.866388e-09
431 6810 STX4 0.6977938 2.856031e-02 1.1719340 3.262791e-07
432 79001 VKORC1 -0.8027366 4.541522e-04 -0.6130450 9.319786e-03
433 2521 FUS 0.7632304 7.252086e-10 0.5075587 3.861127e-05
434 583 BBS2 -0.6261172 8.986188e-03 -0.7058018 3.188413e-03
435 4502 MT2A -1.0055816 3.944840e-16 -0.7554787 9.519839e-13
436 9289 ADGRG1 0.3976620 4.775778e-05 0.2441522 3.486594e-02
437 27183 VPS4A 0.7513295 8.744913e-03 0.6347523 4.998338e-02
438 91862 MARVELD3 2.9430332 3.746597e-03 2.4531545 2.922212e-02
439 5713 PSMD7 -0.6205772 1.408600e-02 -0.7739885 7.044908e-04
440 124491 TMEM170A 0.6723400 4.502382e-03 0.5489694 3.677470e-02
441 4166 CHST6 2.6962446 3.960340e-02 2.9333458 2.154713e-02
442 54386 TERF2IP -0.4463460 1.197770e-02 -0.5627876 6.426842e-04
443 100130958 SYCE1L 2.5517604 3.309420e-08 2.0546872 3.600374e-04
444 4094 MAF 1.0118798 1.150650e-10 0.8890634 1.552017e-07
445 10200 MPHOSPH6 0.8730754 4.180690e-04 0.7648047 6.037178e-03
446 3281 HSBP1 -0.5428740 1.600119e-02 -0.4276417 3.484602e-02
447 23406 COTL1 -0.6228931 1.202637e-09 -0.5049880 1.484791e-05
448 81631 MAP1LC3B -0.7188656 8.473933e-06 -0.6431186 6.301319e-05
449 29123 ANKRD11 -0.6278631 1.274030e-07 -0.8518794 3.709899e-09
450 10381 TUBB3 -1.6371122 8.191888e-09 -1.3555122 2.687430e-06
451 7531 YWHAE -0.4972274 3.319772e-04 -0.6790844 4.976348e-07
452 5306 PITPNA 0.8511018 5.672427e-06 0.8483277 1.945406e-05
453 124935 SLC43A2 0.5879152 6.469172e-04 0.4245842 3.069666e-02
454 6117 RPA1 -1.2230346 4.086653e-03 -1.1241705 4.712945e-02
455 23293 SMG6 1.3085509 1.983173e-06 0.8856096 8.608194e-03
456 83460 EMC6 7.3783605 1.556652e-04 6.3921739 2.837770e-02
457 101559451 <NA> -3.6325402 2.419142e-02 -3.4300353 3.630983e-02
458 23302 WSCD1 1.2867431 3.495667e-21 0.9934216 1.143742e-09
459 54478 PIMREG 1.8914490 8.193218e-03 2.0731304 7.415395e-03
460 1742 DLG4 0.6449161 2.187844e-03 0.6425151 1.379859e-02
461 37 ACADVL 0.9264694 3.956736e-07 0.7891664 3.943013e-06
462 23399 CTDNEP1 0.6252976 6.386210e-04 0.5288386 8.925890e-03
463 1984 EIF5A 1.0315814 4.543233e-06 1.3259060 1.274520e-12
464 6154 RPL26 -1.2272616 2.821646e-21 -1.1028759 1.559946e-20
465 9611 NCOR1 -0.8297948 5.964385e-04 -0.7314405 1.533090e-03
466 125144 SNHG29 -0.8703269 1.045102e-07 -0.5452850 5.894301e-03
467 3996 LLGL1 1.1900278 4.993151e-08 0.8556594 3.675840e-04
468 284047 CCDC144B 1.8884786 7.111301e-05 1.5906406 5.151420e-03
469 22905 EPN2 1.4370574 6.184815e-16 1.5810549 3.356685e-26
470 224 ALDH3A2 0.8853191 1.103245e-05 0.8671828 3.435749e-06
471 218 ALDH3A1 2.3556449 2.035509e-03 2.0678832 9.973890e-03
472 100462977 MTRNR2L1 0.7958907 1.369794e-11 0.5234227 1.051784e-04
473 26118 WSB1 -1.1530284 4.282675e-18 -1.2895732 6.796359e-18
474 6147 RPL23A -0.9519818 4.237693e-12 -0.8341970 1.106815e-09
475 147015 DHRS13 -1.2698348 2.472879e-02 -1.4027750 1.384453e-02
476 6347 CCL2 -0.8309797 1.080186e-11 -0.5559667 2.811113e-06
477 163 AP2B1 0.8419641 1.329770e-10 0.5182979 1.468526e-03
478 91608 RASL10B 1.1936587 5.115343e-03 0.9893958 2.742697e-02
479 9326 ZNHIT3 -0.8371984 3.465783e-02 -1.5988886 3.679887e-03
480 9349 RPL23 -0.8601711 1.368029e-11 -0.7530755 1.834738e-10
481 6143 RPL19 -0.5838960 2.960158e-07 -0.4368548 2.773448e-04
482 10948 STARD3 1.0312622 1.372149e-03 0.7799050 4.037384e-02
483 10209 EIF1 -0.5815818 6.679365e-05 -0.4465559 5.651855e-04
484 60681 FKBP10 0.9927652 4.868650e-09 0.7324452 5.532716e-04
485 2118 ETV4 1.1266639 7.095266e-03 1.0881048 2.325569e-02
486 7343 UBTF 0.6423764 4.736278e-03 0.5637409 2.799931e-02
487 51629 SLC25A39 0.7355032 2.081793e-04 0.5493490 1.216379e-02
488 2670 GFAP -0.9725394 8.074219e-04 -0.8263723 2.111793e-02
489 124790 HEXIM2 -2.1718223 1.020913e-03 -2.5862550 1.359943e-03
490 90507 SCRN2 1.2076429 5.834244e-04 0.9343968 3.683940e-02
491 4804 NGFR -1.8648504 3.212246e-04 -3.1054087 1.054488e-06
492 1277 COL1A1 -2.1977388 1.004864e-28 -1.8782416 1.355766e-15
493 51747 LUC7L3 -0.7556232 5.249112e-07 -0.8744854 1.787546e-07
494 404093 CUEDC1 1.0085939 3.429940e-06 0.9298476 2.985881e-04
495 6426 SRSF1 -0.7835381 1.563324e-03 -0.5706565 1.011060e-02
496 6827 SUPT4H1 0.6457652 2.576390e-02 0.5739720 1.820339e-02
497 653653 <NA> 1.3619291 1.746363e-16 1.3746073 1.757163e-16
498 8493 PPM1D -1.5052961 2.032412e-02 -1.1217445 2.987023e-02
499 342538 NACA2 -0.9918503 7.987224e-06 -0.9597353 1.813232e-06
500 9902 MRC2 0.8169026 4.846276e-12 0.8201005 3.549890e-12
501 1534 CYB561 3.0422758 3.438905e-08 2.1727194 4.956277e-03
502 5573 PRKAR1A -1.1025272 2.731797e-19 -1.1435375 1.047407e-19
503 388419 BTBD17 1.3016105 1.738123e-11 1.1857468 7.405348e-08
504 23510 KCTD2 0.8289403 3.560670e-03 0.6640987 4.150298e-02
505 10476 ATP5PD -0.9717998 5.852795e-06 -0.9420880 4.104210e-06
506 30833 NT5C 1.2280223 3.866066e-02 1.3086616 2.684832e-02
507 6613 SUMO2 -0.5930108 2.705062e-02 -0.7286626 1.625612e-02
508 2584 GALK1 1.7254135 1.310249e-02 2.0021461 3.579051e-03
509 3021 H3-3B -0.3994214 1.863027e-03 -0.4230520 1.282304e-03
510 201292 TRIM65 1.2293615 1.751758e-03 1.0188096 3.123313e-02
511 57690 TNRC6C -1.1507167 9.976401e-04 -0.9694068 5.387006e-03
512 332 BIRC5 2.0906223 6.319246e-06 1.4536843 4.356118e-02
513 9021 SOCS3 -2.0791411 8.463804e-13 -1.0958583 2.703121e-03
514 3959 LGALS3BP 0.6904852 1.327004e-05 0.5355953 8.451631e-04
515 114897 C1QTNF1 1.4696470 2.454798e-18 0.9856446 3.435749e-06
516 284184 NDUFAF8 0.5911756 2.083037e-03 0.6861791 2.647084e-05
517 71 ACTG1 -0.2961316 1.159036e-03 -0.2218373 2.331023e-02
518 2873 GPS1 0.6823021 3.766068e-02 0.7508669 2.494401e-02
519 9123 SLC16A3 -4.1330190 7.226284e-07 -3.3561812 4.548231e-05
520 7525 YES1 -1.2615927 3.266399e-04 -1.0557013 5.607378e-03
521 23347 SMCHD1 0.8597124 9.011347e-03 0.8732188 1.003663e-02
522 10627 MYL12A -0.7901486 3.179507e-06 -0.8546644 3.579980e-07
523 103910 MYL12B -1.0486207 4.662796e-15 -0.8754780 5.158171e-09
524 284217 LAMA1 -1.9117689 6.827497e-03 -2.7721247 3.188413e-03
525 201475 RAB12 1.2942048 3.444771e-05 1.1303184 1.006118e-03
526 23253 ANKRD12 -0.9347668 1.231226e-08 -1.0979806 2.631009e-12
527 11031 RAB31 0.5840879 9.078035e-06 0.7492132 4.453646e-09
528 57534 MIB1 1.0253310 1.369637e-05 0.8547497 1.325508e-03
529 114876 OSBPL1A 0.7727328 5.467397e-04 0.6463417 1.359824e-02
530 361 AQP4 1.3046907 4.415749e-12 1.4466123 3.593129e-19
531 498 ATP5F1A -0.6930482 1.508889e-06 -0.6093631 1.945666e-05
532 497661 C18orf32 2.3368170 8.243963e-03 2.0041399 2.645110e-02
533 10449 ACAA2 -1.1252042 9.418471e-06 -0.9019449 1.560490e-04
534 4677 NARS1 0.5722238 2.091157e-03 0.5704313 6.378848e-03
535 6209 RPS15 -1.0001926 1.043696e-08 -0.7964683 4.814394e-06
536 55643 BTBD2 0.8551346 3.004880e-04 0.7291902 1.318394e-03
537 8943 AP3D1 -0.4757093 9.636398e-03 -0.5547152 2.365682e-03
538 4946 OAZ1 -0.5678812 1.593618e-05 -0.5684510 9.197482e-05
539 51690 LSM7 0.8629005 4.461942e-05 0.7662276 1.043661e-04
540 10362 HMG20B 0.8689997 1.251490e-04 0.7084466 8.678934e-03
541 51341 ZBTB7A -0.7362040 3.299776e-05 -0.9163785 1.887914e-07
542 10226 PLIN3 0.6525927 6.166393e-03 0.6941874 3.886414e-04
543 5802 PTPRS 0.6513059 1.188465e-02 0.5756991 7.909267e-03
544 25873 RPL36 -0.4164091 2.349734e-03 -0.3654459 3.825698e-02
545 126328 NDUFA11 0.5011245 2.876334e-03 0.5154632 9.081134e-03
546 8570 KHSRP 0.5583323 1.269109e-04 0.4700514 8.504777e-03
547 59286 UBL5 -0.5032199 1.910347e-03 -0.3814452 3.069666e-02
548 93145 OLFM2 1.2958196 5.451070e-03 1.2964507 1.633293e-02
549 51073 MRPL4 1.1496204 3.708176e-04 0.8858085 1.234463e-02
550 11140 CDC37 0.5990434 2.052218e-04 0.5739898 1.201889e-04
551 6597 SMARCA4 -0.6024604 4.736583e-03 -0.5722336 8.274450e-03
552 3949 LDLR -1.0692312 9.146430e-05 -0.8628490 4.018303e-03
553 664709 HNRNPA1P10 -0.7790312 1.254641e-03 -0.6843172 2.785025e-02
554 84261 FBXW9 2.0842390 9.239159e-03 2.5722090 2.102751e-04
555 30000 TNPO2 0.9492195 7.131099e-10 0.8814543 2.400691e-09
556 811 CALR 0.7334217 4.393978e-11 0.5382697 1.786768e-06
557 4784 NFIX 1.2616492 8.226083e-09 1.1806997 5.782716e-08
558 22859 ADGRL1 -0.6358973 6.412381e-04 -0.8164053 7.858444e-07
559 4713 NDUFB7 1.3413248 1.327334e-08 1.1192722 6.515834e-06
560 10994 ILVBL 1.7209724 2.451358e-04 1.5464380 3.535904e-03
561 4854 NOTCH3 -0.9657688 5.154303e-09 -0.6352958 2.096314e-04
562 10270 AKAP8 -1.3286091 6.084450e-04 -0.8311861 4.546707e-02
563 58525 WIZ -0.7592523 3.166567e-03 -0.5806022 3.672069e-02
564 4218 RAB8A 1.0376511 2.830296e-03 0.9605041 2.023263e-02
565 79016 DDA1 0.7933349 9.067862e-03 0.9922694 8.085905e-05
566 93343 MVB12A 1.7769065 5.261210e-04 1.3674581 2.398354e-02
567 79709 COLGALT1 0.4124601 4.436015e-02 0.5796921 1.315433e-03
568 170463 SSBP4 1.1732690 3.074895e-02 1.2889058 2.671759e-02
569 55049 REX1BD 1.1068160 3.775347e-04 0.9570863 2.891174e-03
570 9454 HOMER3 1.0207052 2.694411e-07 1.2159963 4.374179e-09
571 1463 NCAN 1.8552424 2.188462e-31 1.6067155 1.535438e-23
572 7562 ZNF708 0.8905314 2.796814e-02 0.8959148 3.408261e-02
573 10775 POP4 -1.2569788 3.964031e-03 -1.2112346 8.921095e-03
574 9141 PDCD5 -0.8202988 2.650478e-03 -1.1732265 9.929970e-07
575 2821 GPI -0.7403914 3.824566e-05 -0.5062771 2.630139e-02
576 57655 GRAMD1A 0.9865818 1.070319e-04 1.1724726 9.852140e-07
577 7392 USF2 0.6781979 3.652474e-04 0.7192832 8.714748e-04
578 81 ACTN4 -1.0975926 7.328821e-18 -1.0113257 2.229865e-14
579 1891 ECH1 -0.8293285 2.034413e-06 -0.9450336 1.075059e-08
580 2091 FBL 0.7555994 3.926679e-04 0.9116386 5.870773e-05
581 208 AKT2 0.6588291 3.737784e-05 0.4835865 9.435243e-03
582 23646 PLD3 0.9610472 3.110679e-05 0.5979353 2.187981e-02
583 57716 PRX 4.6707467 9.114871e-04 3.5917022 4.911900e-02
584 11100 HNRNPUL1 0.3282188 3.652300e-02 0.3857864 7.751267e-03
585 2931 GSK3A 1.3518810 7.885800e-07 1.0514270 9.724454e-04
586 348 APOE 1.4498686 7.075051e-34 1.2934789 3.664330e-25
587 27113 BBC3 0.6373515 2.628145e-05 0.6024491 2.306262e-04
588 6415 SELENOW 0.9909652 2.244993e-13 0.9096428 1.029829e-10
589 9266 CYTH2 0.6943171 3.909534e-03 1.0140591 9.474857e-07
590 587 BCAT2 1.4421405 9.219084e-05 1.1208820 6.235893e-03
591 4924 NUCB1 0.9642368 4.599121e-08 0.6443797 3.636552e-03
592 581 BAX 0.6616982 1.733013e-03 0.6491224 6.961040e-04
593 2997 GYS1 -1.1704013 2.424543e-03 -1.0464762 3.949933e-02
594 6205 RPS11 -0.6835725 3.644532e-10 -0.5347524 2.214286e-06
595 57479 PRR12 -1.4598384 2.028532e-05 -1.1119260 1.034438e-02
596 284361 EMC10 0.6566259 2.508187e-05 0.5678333 1.687299e-03
597 81856 ZNF611 1.3370182 3.068669e-02 1.3784094 2.061382e-02
598 90338 ZNF160 -0.7951403 3.013453e-02 -0.8864334 1.891143e-02
599 22870 PPP6R1 0.7905539 1.541383e-02 1.0882563 2.127000e-05
600 57106 NAT14 1.0606447 2.298690e-06 0.9730329 8.991540e-05
601 11338 U2AF2 0.9576634 4.424857e-11 0.9220234 8.803951e-09
602 10782 ZNF274 -2.7420750 3.225211e-03 -1.5732952 4.183662e-02
603 1 A1BG 1.7764942 1.893902e-07 1.7191093 2.513362e-07
604 27243 CHMP2A -0.5828843 1.790711e-03 -0.5497006 1.227021e-02
605 100506054 RNASEH1-AS1 0.7400574 4.069356e-02 1.0344256 1.935911e-03
606 6664 SOX11 0.4954825 2.411350e-02 0.6476866 2.147638e-03
607 10971 YWHAQ -0.9029050 1.733962e-11 -0.6327886 8.085905e-05
608 23175 LPIN1 -1.0987236 1.704742e-02 -1.4486904 5.378831e-03
609 51594 NBAS 1.0570885 3.172130e-03 1.0400491 7.442351e-04
610 9741 LAPTM4A -0.4276444 9.273345e-03 -0.5209931 2.751332e-04
611 23369 PUM2 -0.8077769 1.395178e-03 -0.8828535 1.338400e-02
612 388 RHOB -0.4902705 7.362897e-03 -0.4944563 1.115636e-02
613 2281 FKBP1B 1.8587152 3.699570e-04 1.5071386 4.030367e-03
614 79172 CENPO 2.6346988 1.997567e-03 2.5050997 9.091497e-03
615 150946 GAREM2 2.2269391 1.410312e-02 2.5053629 2.933181e-03
616 64838 FNDC4 0.7282151 1.101089e-03 0.9049124 2.490206e-05
617 23160 WDR43 1.2714116 2.002280e-03 1.6945405 5.532833e-07
618 57448 BIRC6 1.3074670 2.605890e-03 1.7006418 4.685400e-05
619 4052 LTBP1 -0.9280523 4.631957e-02 -1.0822135 4.577571e-03
620 5610 EIF2AK2 1.3201542 2.865812e-05 1.3663565 6.162688e-06
621 130589 GALM 1.0114772 4.590935e-02 1.2592887 3.058976e-03
622 9581 PREPL 1.0783194 1.875154e-04 1.1955425 5.858133e-06
623 618 BCYRN1 0.9002005 9.506296e-09 0.8671163 1.688549e-06
624 6711 SPTBN1 0.4873272 1.535356e-03 0.4310155 1.241425e-02
625 6233 RPS27A -0.5685306 1.426542e-05 -0.3217539 2.004684e-02
626 2202 EFEMP1 -1.8673761 5.632408e-21 -1.3846348 8.727071e-15
627 53335 BCL11A -3.1478672 1.719544e-02 -3.9258961 3.136373e-03
628 100132215 <NA> 1.1237271 3.231566e-04 0.9652739 1.765376e-03
629 4190 MDH1 -0.8992681 9.385312e-05 -0.9554870 1.642206e-05
630 5861 RAB1A -0.6911741 4.231081e-05 -0.4161447 4.929642e-02
631 200734 SPRED2 -2.1771695 7.274667e-17 -2.3144806 9.369923e-11
632 5093 PCBP1 -0.8339433 1.669351e-07 -1.0072698 8.973847e-09
633 51449 PCYOX1 -0.6178456 9.297444e-03 -0.4927827 3.314734e-02
634 6637 SNRPG -0.7202623 3.978501e-04 -0.4936487 1.798174e-02
635 6697 SPR 6.3043835 2.616049e-02 7.3050653 5.168280e-04
636 94097 SFXN5 0.9450706 2.317729e-09 0.8784552 1.162711e-06
637 3099 HK2 -4.5134193 3.385477e-04 -2.6657547 4.626470e-07
638 9168 TMSB10 -0.5736203 7.823343e-09 -0.4355572 9.581289e-06
639 83439 TCF7L1 -2.2282879 2.068168e-07 -1.0079873 1.235070e-02
640 55818 KDM3A -2.4185131 1.269525e-23 -1.9326630 6.312661e-13
641 56910 STARD7 -1.1795632 1.458482e-07 -1.0762310 2.199631e-05
642 51239 ANKRD39 0.9589030 1.627096e-03 0.9555986 1.768028e-02
643 25972 UNC50 -1.5028819 2.830296e-03 -1.1770171 1.700158e-03
644 9669 EIF5B -1.0579362 2.960876e-09 -1.1704707 1.652664e-13
645 6160 RPL31 -1.0404040 2.146221e-19 -0.7049568 6.596252e-09
646 9448 MAP4K4 0.4832022 7.805133e-07 0.4274676 2.330796e-05
647 100506421 PANTR1 0.9229483 1.773272e-11 0.6850591 1.817861e-05
648 102682016 LINC01159 2.0497373 2.530975e-12 1.0275224 1.398310e-02
649 653489 RGPD3 2.0394914 2.545323e-03 1.9816520 5.670454e-03
650 100288695 LIMS4 -1.3865581 1.818117e-03 -0.9851257 2.064174e-02
651 1622 DBI 0.5536426 3.514880e-05 0.6702063 1.250429e-07
652 5899 RALB 1.0444103 4.665916e-06 1.0003916 3.543981e-05
653 56886 UGGT1 0.6895074 2.846024e-03 0.6133531 2.004684e-02
654 9394 HS6ST1 -1.3221679 1.377281e-12 -0.8731861 2.095159e-07
655 80097 MZT2B 0.9371221 2.801397e-08 0.8164168 8.323667e-08
656 100216479 <NA> 3.3341524 2.877854e-04 2.5693812 4.732672e-02
657 653784 MZT2A 1.0645548 6.488380e-05 0.7527903 2.523920e-02
658 23190 UBXN4 -0.8506191 4.585101e-08 -0.5523627 2.135634e-03
659 390 RND3 0.6240307 5.017165e-05 0.5629198 2.599160e-03
660 375287 RBM43 1.4135384 1.372155e-03 1.4670153 2.579034e-04
661 10254 STAM2 -0.7627544 4.571436e-02 -0.9253563 2.626900e-02
662 4929 NR4A2 -2.4191977 3.070374e-03 -3.0328624 2.034881e-02
663 728066 <NA> -1.4580202 5.873088e-06 -0.8553140 1.162056e-02
664 90 ACVR1 -1.4143322 1.029848e-02 -2.3034422 2.927626e-03
665 85461 TANC1 -1.2011594 2.309556e-06 -1.0267857 1.313677e-04
666 5937 RBMS1 -1.4408726 7.509361e-10 -2.0522384 1.580311e-13
667 5163 PDK1 -2.3289650 3.525734e-04 -1.9930010 1.604289e-02
668 1123 CHN1 0.7179388 6.412381e-04 0.8102291 7.179717e-04
669 518 ATP5MC3 -0.6512415 9.283356e-06 -0.4087624 1.168162e-02
670 10651 MTX2 -1.7298221 5.885057e-05 -1.4966179 2.707084e-03
671 10787 NCKAP1 -0.7908878 1.981423e-03 -1.0718763 3.330677e-08
672 1290 COL5A2 -1.0082829 1.426542e-05 -0.8442413 1.724989e-02
673 57181 SLC39A10 1.0657628 5.129329e-04 1.3850089 1.575471e-07
674 3336 HSPE1 -0.8989412 6.940125e-06 -0.7717259 8.062168e-04
675 8324 FZD7 -2.4628538 1.355483e-03 -1.7968246 4.096832e-03
676 7341 SUMO1 -1.0486226 8.494272e-07 -1.1280136 3.141158e-07
677 8828 NRP2 -0.3111674 2.653930e-02 -0.3753242 3.756314e-03
678 57683 ZDBF2 1.7742951 1.067700e-03 1.7175867 7.228121e-03
679 3417 IDH1 -0.9360679 4.259466e-05 -0.5796108 1.995971e-02
680 4133 MAP2 0.9217919 1.463112e-17 0.8554260 8.747995e-16
681 2335 FN1 -1.0414844 1.863797e-06 -0.5017425 7.137736e-03
682 3488 IGFBP5 -1.3041410 4.974055e-39 -1.1204189 6.556344e-19
683 27013 CNPPD1 1.0722193 3.285990e-06 1.0809768 2.572886e-05
684 10290 SPEG 1.1632199 3.113965e-03 1.1482104 1.893859e-02
685 79586 CHPF 0.5053566 1.080166e-02 0.5071932 1.645624e-02
686 6508 SLC4A3 1.0881830 5.205922e-03 0.9881034 2.703164e-02
687 7857 SCG2 -1.0545084 1.495772e-19 -0.8456456 1.752878e-16
688 81618 ITM2C 1.5341829 1.007894e-05 1.1343081 2.469452e-03
689 26058 GIGYF2 1.1186886 5.326370e-06 1.3335160 4.572632e-10
690 150678 COPS9 0.4398101 8.582452e-04 0.3808745 9.947854e-03
691 547 KIF1A 0.3862738 1.492197e-02 0.4182695 8.618599e-03
692 3069 HDLBP 0.5787054 1.318350e-03 0.4509784 5.380893e-03
693 2280 FKBP1A 0.6321454 2.398389e-04 0.7647272 9.299289e-07
694 55968 NSFL1C 0.7124404 5.639142e-03 0.6688523 1.289891e-02
695 65992 DDRGK1 0.7130558 4.259487e-02 0.9468888 1.581896e-02
696 8455 ATRN -0.8599651 1.733850e-03 -0.7983844 8.100894e-03
697 1059 CENPB 0.8905302 1.348676e-04 0.9085005 2.878473e-04
698 57506 MAVS 1.0247849 1.606832e-05 0.9371398 1.244129e-04
699 9770 RASSF2 0.9046947 9.744347e-03 0.8060801 2.817916e-02
700 92675 DTD1 0.5937663 7.442801e-03 0.5958019 4.935101e-03
701 3642 INSM1 -2.1627318 1.634353e-08 -1.7988030 7.958528e-05
702 1471 CST3 1.2574595 1.228132e-43 1.3989904 1.881513e-43
703 1473 CST5 3.4807533 2.130948e-02 4.4776118 1.137283e-04
704 57136 APMAP -0.4687479 3.452141e-03 -0.5532608 6.636269e-04
705 81502 HM13 0.6103657 4.926889e-02 0.7430691 7.799154e-03
706 6640 SNTA1 0.8609711 4.901495e-05 0.7187834 1.441268e-03
707 83737 ITCH 1.5383474 1.831554e-03 1.7961362 6.301319e-05
708 26133 TRPC4AP 0.6034704 2.640019e-02 0.6360617 2.894149e-02
709 8904 CPNE1 -0.7261921 3.963440e-03 -0.6508457 2.433891e-02
710 140679 SLC32A1 -6.9614078 1.665998e-03 -6.9633237 2.377326e-03
711 7150 TOP1 -0.4334168 2.734083e-02 -0.5474378 1.053712e-04
712 5335 PLCG1 0.9397782 1.294045e-03 0.8981406 1.967691e-03
713 6385 SDC4 1.6033788 1.919486e-03 1.6082534 2.882438e-03
714 51604 PIGT 0.6016027 3.172130e-03 0.6626024 1.048363e-03
715 441951 ZFAS1 -0.4844473 1.465428e-03 -0.5151909 2.065086e-03
716 56937 PMEPA1 -1.1234135 3.486397e-10 -1.1428644 5.947594e-15
717 2778 GNAS -1.1274375 7.936376e-35 -1.0341184 1.907468e-32
718 3911 LAMA5 -0.7958419 1.308276e-03 -0.5909412 3.441671e-02
719 3785 KCNQ2 -1.5208703 2.206510e-07 -2.3512731 1.761040e-10
720 79144 PPDPF -1.0589037 1.557186e-26 -1.1102100 6.796359e-18
721 102724594 U2AF1L5 -0.6142693 2.484599e-02 -1.0716140 5.729579e-06
722 8204 NRIP1 1.8390072 2.593748e-18 1.6222269 1.053896e-11
723 58494 JAM2 -0.8511409 2.458429e-03 -0.7355560 1.485775e-02
724 522 ATP5PF -0.8419146 4.664132e-11 -0.7679662 1.761040e-10
725 6647 SOD1 -0.6465754 9.934063e-06 -0.4027966 8.074615e-03
726 10215 OLIG2 1.4931531 8.338989e-04 1.6938804 3.444987e-04
727 6651 SON -0.7252499 3.708503e-07 -0.5982682 1.977253e-04
728 64968 MRPS6 -0.5600060 7.309418e-03 -0.7456760 8.192199e-05
729 51227 PIGP -1.0151924 6.434849e-05 -1.4169807 8.093641e-04
730 7485 GET1 -0.8782413 5.952885e-04 -0.9844376 3.070395e-03
731 4599 MX1 6.3558573 1.362144e-03 4.2851884 1.592019e-03
732 7307 U2AF1 -0.6778997 1.629854e-03 -0.7079508 1.866647e-03
733 56894 AGPAT3 0.4704983 4.182571e-02 0.4434549 3.450608e-02
734 754 PTTG1IP -0.8988419 1.572866e-09 -0.4989243 3.589519e-05
735 23275 POFUT2 -0.6750628 2.798413e-02 -0.6003035 3.970582e-02
736 3275 PRMT2 0.8618709 6.200504e-14 0.8946230 8.932975e-15
737 7122 CLDN5 8.3883777 1.673883e-07 8.0931000 3.435749e-06
738 84861 KLHL22 1.5055838 2.092838e-04 1.4678917 2.909649e-03
739 24144 TFIP11 0.9653725 2.788975e-02 1.2828603 9.860279e-04
740 84133 ZNRF3 0.9608432 2.313716e-02 0.9603653 3.300728e-02
741 25807 RHBDD3 1.1328601 3.180055e-02 1.1706749 2.703164e-02
742 8508 NIPSNAP1 1.7853732 3.888430e-15 1.7820268 1.803443e-11
743 253143 PRR14L 1.2457609 1.834090e-05 1.1781720 1.482524e-04
744 80830 APOL6 1.4413256 1.671602e-02 1.4310562 2.378136e-02
745 80832 APOL4 2.5210905 4.031680e-10 2.0696845 7.062728e-05
746 25828 TXN2 0.8354706 1.166943e-03 1.1201771 4.557875e-06
747 11078 TRIOBP -0.7775169 1.123170e-04 -0.7080126 3.620766e-03
748 3005 H1-0 1.0858638 9.104073e-05 1.3330747 7.405348e-08
749 10521 DDX17 -0.4657370 7.322971e-04 -0.3207491 2.887454e-02
750 23492 CBX7 -1.5490055 2.001006e-03 -1.9909370 6.265631e-03
751 23112 TNRC6B 0.6110730 1.797362e-04 0.5108770 1.789964e-03
752 158 ADSL -0.9459539 5.056202e-03 -0.8877531 1.911147e-02
753 9978 RBX1 -0.7025602 6.283363e-05 -0.6047769 1.734487e-03
754 27351 DESI1 0.9162164 3.548512e-02 1.0545973 1.730020e-02
755 706 TSPO 0.5150853 3.708176e-04 0.7888045 1.457786e-10
756 25814 ATXN10 -0.8446038 7.293622e-05 -0.7235980 3.696941e-04
757 25817 TAFA5 -1.3326552 2.726045e-03 -0.8881709 4.675505e-02
758 10752 CHL1 -0.4827870 8.236944e-03 -0.5881262 1.121250e-02
759 285362 SUMF1 -1.1779362 3.250033e-04 -1.0124457 2.983479e-03
760 8536 CAMK1 1.2315697 3.066155e-02 1.7299232 4.125938e-04
761 10093 ARPC4 1.0425886 1.572107e-07 0.8647963 1.343671e-04
762 7428 VHL 1.0147178 1.257900e-07 0.7398724 1.863685e-03
763 6396 SEC13 -0.5189447 3.986739e-02 -0.7715083 2.209743e-03
764 9686 VGLL4 0.7273346 3.351188e-02 0.7202849 2.612461e-02
765 6161 RPL32 -0.9977789 9.639138e-19 -0.7711331 4.635884e-12
766 2199 FBLN2 -0.7498440 4.371132e-04 -1.5063973 8.365366e-10
767 79750 ZNF385D 0.6772722 2.654633e-02 0.9190135 3.640681e-04
768 7325 UBE2E2 -1.2106877 1.038667e-02 -1.4987755 2.659635e-03
769 6138 RPL15 -0.7647532 3.373498e-18 -0.8472316 4.347742e-23
770 7048 TGFBR2 -2.6158518 1.354956e-02 -1.8877619 1.626682e-02
771 9045 RPL14 -1.0991655 3.060151e-12 -0.9295706 5.143351e-12
772 55129 ANO10 1.2277481 6.743602e-10 0.6732278 1.223579e-02
773 4134 MAP4 0.5676463 2.457925e-06 0.4497198 4.236870e-04
774 64925 CCDC71 1.2623676 1.081510e-02 1.4606370 4.125938e-04
775 2771 GNAI2 0.4000372 3.721473e-04 0.4367197 9.453246e-04
776 132160 PPM1M 1.3847062 1.284946e-02 1.3472098 2.064174e-02
777 378 ARF4 -0.6787331 4.854183e-04 -1.0599049 1.737575e-06
778 5162 PDHB -0.8998770 4.931384e-04 -0.7050772 1.005182e-02
779 11170 FAM107A 0.9019539 3.595037e-13 0.9156521 2.612074e-11
780 132204 SYNPR -4.5714795 1.342440e-04 -3.1543564 4.305321e-03
781 9861 PSMD6 -1.2462613 2.673589e-05 -1.3719776 1.298088e-03
782 166336 PRICKLE2 -1.2918039 4.676828e-04 -1.5444432 2.659635e-03
783 727936 GXYLT2 2.3185270 2.407880e-03 2.3473233 8.451631e-04
784 285237 C3orf38 -1.1868078 2.630981e-02 -1.6640196 1.964804e-03
785 131544 CRYBG3 1.6878201 2.613220e-02 1.7089412 4.547170e-02
786 55076 TMEM45A -6.3544942 2.237875e-03 -4.0427023 4.576852e-02
787 29114 TAGLN3 -2.0269659 3.682592e-03 -2.4059202 1.464662e-07
788 151887 CCDC80 0.8069456 2.122272e-15 0.7595701 3.271995e-14
789 25871 NEPRO 0.7898265 3.515523e-02 0.8704867 1.744985e-02
790 10063 COX17 -1.0122122 1.229871e-06 -0.9457776 2.720699e-06
791 4710 NDUFB4 0.4495966 7.309418e-03 0.4128587 1.443009e-02
792 2804 GOLGB1 0.5022046 7.909733e-03 0.4755504 9.953376e-03
793 26355 FAM162A -1.6172471 3.813423e-18 -1.3451385 1.041823e-12
794 54625 PARP14 1.2464407 3.350094e-04 1.0046250 2.974436e-02
795 54437 SEMA5B -0.5775444 4.530561e-02 -1.3440046 1.860633e-03
796 10954 PDIA5 -0.7754740 4.570312e-02 -0.7258743 2.742697e-02
797 57493 HEG1 -1.0557142 3.478100e-08 -0.6742118 1.076310e-02
798 84303 CHCHD6 1.6919593 7.496738e-05 1.3962226 4.434165e-03
799 11343 MGLL 1.1144299 9.215220e-17 1.0100009 1.558741e-07
800 29927 SEC61A1 0.4791172 1.790711e-03 0.3892516 2.795976e-02
801 8607 RUVBL1 1.0489119 2.763334e-03 0.9160656 2.363942e-02
802 8971 H1-10 1.4063725 2.340612e-18 1.3063005 1.126179e-14
803 6259 RYK -2.0211616 7.118626e-14 -1.4405850 1.545863e-06
804 22808 MRAS 0.7019507 1.266008e-02 0.8127516 1.315527e-02
805 92370 PXYLP1 -0.9637595 6.215956e-04 -1.1259169 2.790936e-05
806 9616 RNF7 -1.2038336 1.261723e-06 -1.2897244 6.862628e-09
807 483 ATP1B3 -0.6341125 6.438534e-03 -0.8346410 3.796594e-04
808 9435 CHST2 -1.4414572 4.268293e-16 -1.1660266 6.552169e-12
809 205428 DIPK2A -0.9895666 2.078672e-03 -1.2635912 3.914614e-05
810 5352 PLOD2 -1.1564791 8.469171e-06 -1.0093895 7.767172e-05
811 4071 TM4SF1 -1.0480331 6.563097e-10 -1.1388825 7.489678e-09
812 51714 SELENOT -0.8988647 1.330311e-03 -0.7237756 1.255979e-02
813 6747 SSR3 -0.5296982 1.839420e-05 -0.4132969 3.093032e-02
814 25976 TIPARP 1.2298068 1.874552e-05 0.9784203 3.242270e-03
815 647033 <NA> 1.4506093 9.219084e-05 1.8080614 3.645026e-06
816 5806 PTX3 -2.4429172 1.761816e-09 -1.8866608 2.550278e-05
817 101243545 LINC02067 -1.2758056 6.856263e-05 -1.2680884 9.724454e-04
818 7095 SEC62 -0.5057600 3.652376e-04 -0.6414447 3.540592e-06
819 64778 FNDC3B -0.7124311 8.757998e-04 -0.7662147 1.237268e-04
820 64393 ZMAT3 0.5898881 3.161822e-04 0.7586235 9.426139e-08
821 59345 GNB4 0.5375369 1.846323e-02 0.4834116 2.634587e-02
822 4711 NDUFB5 -0.6678206 7.309418e-03 -1.0444914 9.929970e-07
823 5708 PSMD2 0.8175908 6.570138e-06 0.9086990 1.218329e-06
824 1981 EIF4G1 0.6148957 1.149979e-03 0.6750565 6.235094e-04
825 23355 VPS8 0.9302267 1.671602e-02 1.4890532 4.486472e-05
826 10644 IGF2BP2 -1.8910339 2.716598e-13 -1.0584528 1.101850e-05
827 2119 ETV5 0.6231281 5.147271e-06 0.4834801 7.078013e-03
828 1427 CRYGS 1.0250111 6.286110e-04 1.0003041 7.732515e-03
829 1974 EIF4A2 -0.5422310 1.979869e-03 -0.7003313 1.027394e-03
830 3280 HES1 1.3067640 5.177006e-17 1.0026634 1.165196e-14
831 152002 XXYLT1 0.6107124 2.726715e-02 0.7424757 2.917558e-03
832 5504 PPP1R2 -0.8542790 1.198774e-02 -1.3598579 2.999403e-04
833 7037 TFRC -1.8902219 1.051211e-10 -1.3824688 7.618815e-06
834 22916 NCBP2 0.4690908 3.574626e-02 0.5596570 2.567591e-02
835 6165 RPL35A -0.9143664 1.463252e-16 -0.9048525 7.518873e-17
836 521 ATP5ME -0.3755903 4.960564e-02 -0.6288114 1.109825e-03
837 2580 GAK 0.7949277 3.912914e-03 0.8002064 3.260131e-03
838 10460 TACC3 2.0448673 3.311675e-08 1.3201499 1.334292e-02
839 677770 SCARNA22 3.5640783 4.500075e-24 3.5273401 9.813043e-20
840 401115 C4orf48 1.4948861 3.255711e-10 0.9191376 5.660188e-03
841 339983 NAT8L 1.1542053 2.673589e-05 0.9319789 4.935101e-03
842 10608 MXD4 1.1572425 6.551740e-18 0.7724441 3.540592e-06
843 118 ADD1 0.4879196 5.157894e-04 0.4971619 3.656670e-05
844 6002 RGS12 1.2826448 2.112503e-04 1.4270219 3.773619e-05
845 85013 TMEM128 -1.1757276 2.173370e-02 -1.1556747 3.313285e-02
846 114932 MRFAP1L1 1.0002344 3.512925e-08 0.6430647 8.824011e-03
847 259282 BOD1L1 -0.4510582 1.266008e-02 -0.5817596 1.280140e-03
848 5860 QDPR -0.6227368 1.189567e-03 -0.4796809 2.649954e-02
849 57620 STIM2 0.4937752 1.150648e-02 0.3581694 2.972976e-02
850 5099 PCDH7 -1.1263769 9.919625e-03 -1.2945977 1.285023e-03
851 55728 N4BP2 1.6450103 1.143343e-03 1.4412843 3.562897e-03
852 65997 RASL11B -6.6651138 1.125966e-02 -5.7044272 3.639798e-02
853 5156 PDGFRA -1.7558863 1.515392e-17 -1.8639525 2.069884e-30
854 3490 IGFBP7 0.6820844 4.251649e-03 1.0385101 4.750404e-07
855 258 AMBN -6.1985571 1.220498e-04 -4.1219349 2.647842e-03
856 25849 PARM1 -0.4986369 1.909574e-06 -1.0396695 1.014559e-17
857 950 SCARB2 -0.3059094 1.250141e-02 -0.3697860 2.749261e-02
858 10983 CCNI -0.6816002 6.122848e-08 -0.6780022 1.665411e-09
859 65008 MRPL1 0.8607050 8.027712e-03 0.8818691 3.535904e-03
860 9987 HNRNPDL -0.3992914 1.069576e-02 -0.6819008 1.255247e-05
861 79966 SCD5 -0.6425003 1.621696e-05 -0.3401131 1.057945e-02
862 6696 SPP1 0.7750330 9.473182e-05 0.6493998 1.390013e-02
863 10611 PDLIM5 -0.7791109 3.303880e-05 -0.8104113 2.829357e-05
864 8633 UNC5C -1.2619957 1.437038e-02 -1.1217370 3.346579e-02
865 3015 H2AZ1 -0.4375541 4.262233e-02 -0.6286706 8.081783e-03
866 7323 UBE2D3 -0.5800032 3.291082e-02 -0.4983115 1.412559e-02
867 6164 RPL34 -1.2316059 2.718076e-31 -1.1214041 1.559515e-35
868 58505 OSTC -0.9127847 2.491635e-05 -1.1003852 6.222865e-07
869 79071 ELOVL6 1.0002636 2.293579e-02 1.3548378 1.253234e-03
870 287 ANK2 0.8508230 1.318390e-05 0.9854018 9.838779e-08
871 401152 C4orf3 -0.7399140 3.738033e-06 -0.6491520 4.171132e-05
872 10252 SPRY1 1.3496250 1.608686e-29 1.0878301 2.718616e-22
873 10424 PGRMC2 -0.7313297 2.492626e-03 -0.7653054 2.841173e-03
874 57575 PCDH10 -0.5576917 7.778962e-04 -0.8800318 7.517228e-08
875 55294 FBXW7 0.8304382 3.218092e-02 0.9279142 9.319937e-04
876 79884 MAP9 -0.7048196 2.155945e-02 -0.7641540 4.737882e-03
877 6307 MSMO1 -1.2232146 7.766704e-11 -0.5305763 9.201490e-04
878 91351 DDX60L 1.7888210 3.214107e-02 1.7988656 3.483148e-02
879 9848 MFAP3L -1.1368037 1.119360e-03 -1.3853712 1.935911e-03
880 11341 SCRG1 -0.6441951 3.408611e-02 -1.2278129 2.102751e-04
881 2180 ACSL1 -3.0422754 2.757001e-02 -6.1202059 3.440427e-04
882 291 SLC25A4 -1.0196267 1.774482e-03 -1.2727577 1.421572e-05
883 65980 BRD9 0.7319604 7.404146e-03 0.7007806 1.168245e-02
884 4726 NDUFS6 -0.4293533 2.798413e-02 -0.7182884 3.700270e-04
885 170690 ADAMTS16 2.1418325 1.181011e-05 2.2530015 1.613799e-05
886 23379 ICE1 1.1141457 2.527353e-03 1.2641374 3.386557e-04
887 10299 MARCHF6 -0.5055872 4.142033e-02 -0.5933650 8.625888e-03
888 1008 CDH10 -1.8310709 3.009173e-07 -1.3356292 2.062147e-06
889 6897 TARS1 0.5449755 7.651393e-03 0.5618440 2.483078e-02
890 114899 C1QTNF3 -1.8710080 8.201995e-06 -1.5951577 1.451104e-02
891 92255 LMBRD2 1.2907673 3.843427e-02 1.2612212 2.782013e-02
892 202151 RANBP3L 5.9947575 4.769557e-02 7.3259727 2.960946e-04
893 6507 SLC1A3 1.2261073 2.073054e-27 1.0474464 9.826939e-22
894 3977 LIFR 0.5075304 2.438437e-02 0.5055484 4.009305e-02
895 9180 OSMR -1.6806252 2.166379e-17 -1.3079736 1.404824e-08
896 3157 HMGCS1 -1.2977528 5.202469e-28 -0.4698210 5.720666e-04
897 5144 PDE4D -1.8255781 2.438437e-02 -1.7623041 1.744985e-02
898 3796 KIF2A 0.7588020 1.641977e-02 0.6608915 2.634601e-02
899 285671 RNF180 1.1000820 2.106799e-05 1.1443162 7.176229e-06
900 285672 SREK1IP1 -1.2487584 7.253432e-08 -0.8534174 3.636552e-03
901 11039 <NA> 2.9008860 4.571436e-02 2.6864421 1.835619e-02
902 3842 TNPO1 1.0534232 1.098834e-08 1.1684308 3.641483e-11
903 689 BTF3 -0.6640619 1.440408e-04 -0.6391783 1.323864e-03
904 3156 HMGCR -1.0292318 1.317987e-08 -0.7945030 3.292241e-05
905 10087 CERT1 1.2562065 2.461916e-08 0.5547309 4.091248e-02
906 51426 POLK 0.9117751 3.534386e-02 0.9887011 2.568562e-02
907 6902 TBCA -0.5198963 8.391923e-05 -0.4901010 1.117374e-04
908 105379045 <NA> 5.5359017 7.264987e-05 4.8129384 2.490820e-03
909 256987 SERINC5 -0.8856079 2.007828e-09 -0.7677267 4.814394e-06
910 100462981 MTRNR2L12 2.3728233 4.412485e-121 2.0821714 9.216426e-90
911 5924 RASGRF2 0.6281567 1.306408e-02 0.8531180 7.074303e-04
912 6228 RPS23 -0.7586035 1.158385e-14 -0.4561300 2.072916e-05
913 1462 VCAN 2.1385062 2.385483e-34 2.0203203 4.347742e-23
914 9652 TTC37 0.8667440 2.934680e-04 0.7973857 3.535904e-03
915 285704 RGMB -2.2793041 2.506751e-02 -1.5081480 2.450929e-02
916 5066 PAM -0.5339104 4.312686e-03 -0.6884944 6.150246e-04
917 90355 MACIR -1.2077593 3.333911e-02 -1.6540499 1.093099e-03
918 2241 FER 1.8287641 1.670853e-04 1.9395679 4.256741e-06
919 4124 MAN2A1 -1.4805865 1.685329e-04 -1.2347478 1.275351e-02
920 324 APC 1.2938950 1.246320e-19 1.1667567 8.551660e-16
921 7905 REEP5 -0.8599144 7.253432e-08 -0.4905003 5.003620e-03
922 9140 ATG12 0.6628566 2.882654e-02 0.7136343 1.659067e-02
923 57556 SEMA6A -0.8584427 1.507903e-14 -0.8119492 3.459618e-14
924 153443 SRFBP1 1.7492238 7.651393e-03 1.5027332 8.100894e-03
925 4015 LOX -3.4492726 1.534331e-02 -5.5702533 2.751332e-04
926 4001 LMNB1 1.0735518 3.918699e-02 1.2470807 1.052093e-02
927 3094 HINT1 -0.9597675 1.443601e-13 -0.8270266 4.886536e-11
928 56951 C5orf15 -1.2436538 6.155303e-09 -1.0393249 2.422391e-07
929 6500 SKP1 -0.7388214 6.672718e-10 -0.6068449 2.981052e-04
930 23338 JADE2 1.6415751 3.538368e-03 1.5306703 2.562736e-02
931 9879 DDX46 -1.3232377 1.228772e-09 -1.3897567 5.601777e-10
932 6695 SPOCK1 -0.3992412 1.909529e-02 -0.9485331 3.482655e-09
933 51308 REEP2 0.5177892 1.398287e-02 0.8098869 2.218699e-05
934 1495 CTNNA1 -0.3223623 4.384920e-02 -0.3620190 5.612742e-03
935 5813 PURA -0.4739584 2.495881e-02 -0.8561705 9.168914e-06
936 56139 PCDHA10 1.6594768 1.243936e-02 2.0103726 4.414930e-04
937 1729 DIAPH1 1.0095936 1.129233e-02 0.9517830 7.517506e-03
938 9812 DELE1 0.9966876 3.618391e-03 0.9875318 2.873087e-03
939 10007 GNPDA1 0.7444086 4.772613e-06 0.8028246 3.387729e-07
940 81555 YIPF5 -0.7107069 3.262287e-02 -0.6276118 2.276521e-02
941 1836 SLC26A2 -0.7002361 3.667461e-02 -0.7535285 3.819847e-02
942 6949 TCOF1 -1.3559200 9.919625e-03 -1.1530379 9.727384e-03
943 972 CD74 2.6668660 2.020915e-19 1.8788221 6.903108e-07
944 309 ANXA6 1.1994955 8.360836e-16 0.6720061 1.064444e-04
945 2760 GM2A 0.9908944 1.062736e-07 0.6623566 3.097188e-03
946 57451 TENM2 -1.5403147 1.463252e-16 -1.2374595 2.430985e-13
947 79646 PANK3 1.3983193 6.510940e-03 1.3158478 1.827212e-02
948 4869 NPM1 -0.6629394 3.120819e-02 -0.8830174 1.857708e-04
949 91272 BOD1 -0.9615587 3.273182e-03 -0.9526690 4.429697e-02
950 1212 CLTB 0.7480007 8.434080e-03 0.7894512 6.304220e-03
951 64324 NSD1 1.7693606 1.346075e-12 1.5941898 4.279366e-09
952 80758 PRR7 0.5590812 2.596234e-02 0.5954623 2.232803e-02
953 91522 COL23A1 1.5331617 1.003958e-04 1.1047430 8.405361e-03
954 821 CANX -0.4401075 8.078890e-05 -0.2923362 8.504777e-03
955 8878 SQSTM1 0.3269261 6.827497e-03 0.2845370 2.733297e-02
956 55819 RNF130 -1.0273074 5.037306e-05 -1.1695436 4.841817e-05
957 10399 RACK1 -0.5853888 9.380366e-07 -0.3534830 5.979495e-03
958 7280 TUBB2A -0.6523625 6.820400e-10 -0.4223057 2.536242e-02
959 347733 TUBB2B -0.4068452 3.092731e-05 -0.3121213 2.065086e-03
960 221749 PXDC1 0.9963329 1.487422e-03 1.0337919 1.924487e-04
961 8899 PRPF4B -0.6382914 6.066757e-04 -0.6727835 2.568866e-03
962 54898 ELOVL2 0.8309062 3.086342e-12 0.6013948 1.990421e-05
963 10048 RANBP9 -0.8719225 1.763338e-02 -0.9313987 8.115302e-03
964 10486 CAP2 -1.2199630 2.316106e-03 -0.9460051 2.655741e-02
965 3400 ID4 -1.1729122 3.536546e-05 -1.4701369 4.163988e-08
966 55856 ACOT13 -1.0767539 3.581976e-03 -1.2781208 1.201889e-04
967 9750 RIPOR2 -4.3828824 7.814646e-03 -4.8433891 4.851746e-03
968 55604 CARMIL1 1.4839269 3.405275e-03 1.8063191 8.938578e-04
969 10590 SCGN -2.3919563 1.044409e-05 -3.9601061 4.182359e-23
970 8364 H4C3 0.9451051 1.366833e-05 0.7174858 2.036132e-03
971 493812 HCG11 4.7796185 2.029789e-03 4.3485079 3.707406e-02
972 3137 <NA> 1.3140214 8.496514e-09 1.3031711 1.009809e-07
973 3107 HLA-C 1.1193240 2.938337e-19 1.0818260 4.847682e-15
974 3106 HLA-B 1.1287090 1.325624e-06 1.2305854 8.705199e-08
975 534 ATP6V1G2 1.8892072 1.705161e-02 1.9257573 2.463332e-02
976 4758 NEU1 0.9330691 8.174253e-03 1.4729430 1.928330e-04
977 7922 SLC39A7 1.1109136 5.258606e-04 0.8742493 1.166589e-02
978 6222 RPS18 -0.8177342 4.819272e-03 -0.8081587 9.380303e-03
979 6892 TAPBP 0.6831594 3.319996e-03 0.5612667 1.213298e-02
980 221491 SMIM29 0.9451797 9.219084e-05 1.2516034 5.325344e-06
981 1026 CDKN1A 0.3671832 2.120348e-04 0.5293086 4.230156e-08
982 57699 CPNE5 1.7044231 2.456650e-43 1.4739600 2.887406e-20
983 9025 RNF8 1.1573508 1.384884e-03 1.3092432 2.102751e-04
984 2739 GLO1 0.6264107 1.973620e-03 0.7763592 6.301319e-05
985 6722 SRF -0.8589408 2.513043e-02 -0.7375109 1.937077e-02
986 7422 VEGFA -3.5771151 8.006230e-39 -1.8740998 7.189818e-14
987 55362 TMEM63B 0.9507098 9.349905e-03 1.1685571 1.792530e-05
988 9697 TRAM2 -1.3151451 3.507218e-10 -1.0203589 2.081117e-06
989 730101 <NA> -0.8255339 3.223422e-02 -1.5221937 1.105712e-05
990 2941 GSTA4 -0.6254280 2.752608e-03 -0.5224627 2.732302e-02
991 667 DST 1.0260073 8.346718e-11 1.1947347 7.062728e-05
992 577 ADGRB3 1.1970290 1.941937e-03 0.9884202 4.436115e-02
993 79627 OGFRL1 1.1446520 1.880664e-08 0.4622340 4.657382e-02
994 79940 LINC00472 3.0193372 1.401517e-08 2.9805289 1.577472e-08
995 1915 EEF1A1 -0.5659628 9.077925e-07 -0.4132856 5.424443e-04
996 55754 TMEM30A 1.4721523 3.562862e-02 2.1280078 3.197776e-05
997 55023 PHIP -0.8736034 2.098786e-03 -1.0001512 7.837069e-04
998 387066 <NA> -1.1806565 1.434827e-09 -0.7796893 6.761550e-05
999 26799 <NA> 4.2969273 5.619315e-15 3.1919402 8.062652e-05
1000 1268 CNR1 0.8407997 2.962184e-03 0.8385144 2.666680e-03
1001 10690 FUT9 1.6137422 2.910943e-10 1.0974774 3.088599e-04
1002 25957 PNISR -0.8351896 4.188804e-07 -0.7329497 1.859918e-06
1003 892 CCNC -1.3283579 6.107508e-03 -1.0828915 2.086549e-02
1004 10973 ASCC3 0.9740322 1.398450e-02 1.3374888 2.371890e-04
1005 100133941 CD24 -4.3815306 1.404673e-02 -5.6309490 5.191700e-03
1006 55084 SOBP -0.4245339 9.760040e-03 -0.6266803 2.779028e-04
1007 2309 FOXO3 0.7411300 1.225323e-02 0.7568960 9.478544e-03
1008 8763 CD164 -0.6601841 4.613712e-05 -0.5391174 2.143089e-03
1009 10758 TRAF3IP2 0.6459487 1.929006e-02 0.7564423 9.455946e-03
1010 2534 FYN -0.3430576 1.472462e-02 -0.4551027 2.967679e-03
1011 3910 LAMA4 1.2757771 6.167079e-05 1.4702063 2.159632e-06
1012 7259 TSPYL1 -0.6590721 1.554525e-02 -0.6814849 7.732515e-03
1013 57515 SERINC1 -0.5948738 2.187020e-03 -0.9072815 1.819267e-06
1014 2173 FABP7 -1.0895523 1.468732e-18 -0.8173959 3.235664e-02
1015 51020 HDDC2 -0.6087856 3.339516e-02 -0.7685111 1.599050e-03
1016 135114 HINT3 0.6125591 4.324743e-02 0.7963661 5.016529e-03
1017 1490 CCN2 -3.4233347 9.244045e-07 -3.4623379 9.118969e-27
1018 6206 RPS12 -1.0102756 3.852799e-13 -0.6113954 6.954987e-05
1019 81706 PPP1R14C 1.2884750 1.861711e-02 1.6802229 7.939616e-04
1020 56995 TULP4 1.4635305 1.210761e-06 0.7874619 4.409724e-02
1021 3482 IGF2R -0.4982640 7.559460e-03 -0.4628997 2.823296e-02
1022 51660 MPC1 -0.6362673 2.034413e-06 -0.4037930 1.466118e-02
1023 6196 RPS6KA2 0.4482844 1.645518e-02 0.7026317 3.622400e-07
1024 5689 PSMB1 -1.4306699 1.709341e-14 -1.3353015 1.844748e-10
1025 60 ACTB 0.4316069 1.156729e-04 0.3813709 1.549594e-03
1026 6624 FSCN1 -1.0650741 3.295316e-21 -0.8177755 5.345990e-12
1027 79034 C7orf26 1.3754879 9.808167e-03 1.3930069 4.397246e-03
1028 4697 NDUFA4 -0.5565546 3.034047e-08 -0.4021756 2.647842e-03
1029 9771 RAPGEF5 1.6813472 1.588777e-07 1.1190181 1.639858e-02
1030 55975 KLHL7 -1.3929368 4.169786e-06 -1.1058372 1.027960e-03
1031 10643 IGF2BP3 0.8121885 7.308784e-04 1.1770296 1.648070e-09
1032 11335 CBX3 0.6725664 2.937034e-05 0.6021006 1.255247e-05
1033 9865 TRIL -0.5413231 2.373205e-06 -0.4316549 1.174174e-03
1034 23333 DPY19L1 -1.9756419 1.870823e-07 -1.3396201 1.441338e-05
1035 83930 STARD3NL -1.4000936 4.106381e-07 -1.3635678 1.394166e-08
1036 27072 VPS41 0.9637214 3.367563e-04 0.7366929 9.455946e-03
1037 5683 PSMA2 -1.2694849 4.083299e-09 -1.2379262 4.584934e-05
1038 23386 NUDCD3 1.1975808 1.083489e-05 1.0873010 3.773619e-05
1039 83605 CCM2 0.9187804 2.916465e-03 0.8723152 1.442882e-02
1040 3486 IGFBP3 -2.1268987 3.475055e-02 -3.3522683 5.434873e-06
1041 23480 SEC61G -1.2225534 2.300721e-02 -1.0217223 2.102751e-04
1042 1956 EGFR 1.2505734 3.807672e-04 1.4236782 8.811623e-05
1043 908 CCT6A -0.8025697 1.260260e-05 -0.5358095 7.769446e-03
1044 51142 CHCHD2 0.4914996 2.099724e-04 0.5238814 3.391164e-04
1045 285908 LINC00174 -2.6778128 3.247166e-02 -3.0969462 1.190762e-03
1046 55069 TMEM248 0.5109028 4.264221e-02 0.5584811 1.641871e-02
1047 51119 SBDS -0.9086152 2.861771e-03 -1.0980977 4.180930e-03
1048 2969 GTF2I 1.8741678 2.115792e-08 1.4704149 3.949890e-04
1049 2970 GTF2IP1 0.4957421 9.760040e-03 0.3458507 4.197736e-02
1050 5447 POR 1.2552945 5.719273e-06 0.8218520 1.600230e-02
1051 3315 HSPB1 -0.5108580 2.458787e-05 -0.4151774 1.012390e-03
1052 222194 RSBN1L -1.1127770 6.827497e-03 -1.2398893 1.855899e-03
1053 3082 HGF -6.1562404 3.626441e-02 -6.1579361 4.498105e-02
1054 5244 ABCB4 -6.1965700 3.675018e-02 -6.1991912 4.528461e-02
1055 6717 SRI -0.7268927 4.375343e-09 -0.6253537 3.047018e-07
1056 5218 CDK14 0.9575605 6.388307e-04 0.7238509 3.593873e-02
1057 54467 ANKIB1 0.9857906 5.437996e-05 0.6429197 2.703164e-02
1058 1021 CDK6 0.8255013 1.909068e-04 0.6914380 1.934690e-03
1059 1278 COL1A2 -1.7032506 3.730046e-35 -3.0662598 4.949363e-58
1060 8910 SGCE -0.9947101 4.297143e-03 -1.1717772 2.244372e-06
1061 7979 SEM1 -0.8872191 4.444586e-10 -0.6656241 3.769006e-05
1062 1749 DLX5 -2.4091798 3.538169e-05 -2.4808466 3.053985e-05
1063 10552 ARPC1A -1.5505765 1.132553e-06 -1.4014472 3.374089e-05
1064 9551 ATP5MF -0.9122261 1.046184e-12 -1.0013636 2.320850e-12
1065 441272 SPDYE3 3.2323583 2.483159e-06 2.3129106 1.878336e-02
1066 81628 TSC22D4 0.7218744 2.534931e-02 0.6623944 4.866324e-02
1067 100316904 SAP25 -6.5211249 7.075835e-03 -6.5232547 9.746457e-03
1068 7205 TRIP6 0.5184782 6.242458e-03 0.7052049 4.814394e-06
1069 5054 SERPINE1 -0.8147019 3.306630e-04 -0.7419385 6.179144e-06
1070 51024 FIS1 0.6621925 9.567496e-04 0.7320772 2.610772e-03
1071 5439 POLR2J -0.8340540 5.136801e-03 -0.6312595 4.167769e-02
1072 5701 PSMC2 -0.8227119 6.566488e-04 -0.8105875 2.164361e-04
1073 3912 LAMB1 -1.7129157 7.869335e-07 -1.6398376 3.792587e-07
1074 5803 PTPRZ1 0.8499531 8.857353e-16 0.7828030 2.986159e-13
1075 2861 GPR37 -1.0058424 1.062736e-07 -0.9841472 6.116840e-07
1076 29923 HILPDA -0.8644915 2.690730e-03 -1.3336372 3.192238e-06
1077 813 CALU -0.5642215 1.098449e-03 -0.6757290 1.711393e-05
1078 9296 ATP6V1F 0.5605802 1.204329e-03 0.4319463 2.040272e-02
1079 23534 TNPO3 0.5267186 3.753410e-03 0.4674936 8.772606e-03
1080 800 CALD1 -1.0604498 3.095809e-25 -0.9969709 1.525995e-19
1081 5764 PTN -0.5561775 1.263635e-05 -0.5236909 2.155123e-05
1082 9747 TCAF1 -0.6596205 1.331632e-02 -0.7793796 3.302720e-04
1083 26047 CNTNAP2 1.0529656 8.191008e-03 1.3018181 1.239605e-05
1084 9601 PDIA4 0.5400641 3.166494e-03 0.4304019 2.485455e-02
1085 155066 ATP6V0E2 0.8836044 1.188315e-09 0.7137228 1.087412e-03
1086 10922 FASTK 0.6739543 2.956378e-02 1.0148587 8.069758e-04
1087 6009 RHEB -0.6596723 7.255424e-04 -0.8501332 2.907693e-06
1088 58508 KMT2C 1.2837004 1.511755e-07 0.6270360 3.737328e-02
1089 3638 INSIG1 -1.5311896 1.530070e-09 -1.2334013 5.062690e-05
1090 9690 UBE3C 0.7247088 1.094544e-02 1.1348315 9.544714e-06
1091 55326 AGPAT5 0.6809654 1.034671e-02 0.6855259 1.045075e-02
1092 9258 MFHAS1 1.8839473 1.540980e-04 1.8457996 2.305208e-03
1093 2222 FDFT1 -0.9574911 6.468122e-07 -0.4416615 2.561598e-02
1094 7991 TUSC3 -0.6248532 1.410312e-02 -0.8541539 1.322641e-02
1095 51201 ZDHHC2 -1.1095686 1.298308e-02 -1.3326518 3.590577e-02
1096 4023 LPL -0.8077300 2.695365e-07 -1.7464661 1.149501e-25
1097 4747 NEFL -1.5177214 2.056881e-19 -1.5656654 1.525995e-19
1098 665 BNIP3L -1.1564470 3.336063e-11 -0.9224700 4.196975e-13
1099 1808 DPYSL2 0.6517512 6.182593e-05 0.6439877 8.411273e-07
1100 148 ADRA1A 1.1247519 1.180486e-09 1.0022627 3.841822e-07
1101 1191 CLU 0.4500957 2.038659e-05 0.3534327 2.737931e-03
1102 51435 SCARA3 1.0436263 7.111232e-03 0.9383369 2.483078e-02
1103 5368 PNOC -2.4193744 5.896725e-06 -3.1203224 7.834159e-08
1104 55893 ZNF395 -1.1092006 1.141548e-04 -1.2240649 2.687430e-06
1105 7976 FZD3 0.4565468 3.899663e-06 0.3478755 6.458641e-03
1106 2137 EXTL3 -0.8286334 1.063351e-11 -0.9122561 7.306328e-13
1107 51669 SARAF -1.1054450 2.685201e-23 -0.8435553 2.861997e-12
1108 10671 DCTN6 -0.7865972 1.060526e-02 -0.9294487 1.026461e-02
1109 692084 SNORD13 3.5814883 1.084694e-14 3.1406624 2.981701e-09
1110 83877 TM2D2 -0.6954581 3.989551e-02 -0.6575888 4.432710e-02
1111 51125 GOLGA7 -0.8253894 1.184741e-03 -0.7949723 6.108184e-04
1112 1052 CEBPD -1.3986854 2.748017e-06 -1.6086805 1.498142e-05
1113 5591 PRKDC 1.0564994 6.098003e-07 0.7960897 1.725305e-03
1114 6224 RPS20 -0.4287188 7.568176e-04 -0.3019994 2.604493e-02
1115 54928 BPNT2 -0.8045056 6.665623e-06 -0.6206243 1.523335e-02
1116 6386 SDCBP -0.7543932 4.594914e-05 -0.5516403 2.873087e-03
1117 9760 TOX 1.0267727 2.116667e-05 0.7767222 1.072121e-02
1118 641638 <NA> -1.1172364 6.670521e-13 -0.9594642 8.276891e-09
1119 80243 PREX2 2.0993992 1.109976e-03 1.9029162 3.136373e-03
1120 23213 SULF1 -2.2849884 1.666922e-11 -2.8757807 4.827200e-17
1121 27067 STAU2 1.5194868 3.060151e-12 1.5477622 5.051800e-15
1122 6921 ELOC -0.5749804 4.189718e-03 -0.6347296 1.215240e-03
1123 83690 CRISPLD1 -0.8573380 1.880054e-03 -0.6742713 4.791183e-02
1124 793 CALB1 -3.3433224 3.608602e-08 -1.6612826 3.262349e-03
1125 169200 TMEM64 -1.3885216 6.328290e-06 -1.0534055 5.380893e-03
1126 157657 CFAP418 2.1406114 2.826441e-03 2.3907474 1.858846e-04
1127 7381 UQCRB -0.8441729 2.247430e-08 -0.7093605 9.355824e-06
1128 6383 SDC2 -1.3612925 9.876046e-18 -0.7310451 2.871879e-05
1129 10404 CPQ 1.4685766 2.247100e-06 1.5448830 8.279331e-08
1130 6156 RPL30 -0.7385175 4.846276e-12 -0.6084917 5.564470e-09
1131 157680 VPS13B 1.7788432 1.983173e-06 1.2807797 1.300299e-02
1132 1345 COX6C -0.4642074 1.578547e-04 -0.3599264 4.485192e-03
1133 26986 PABPC1 1.0219107 4.500075e-24 1.1563470 2.443693e-32
1134 7534 YWHAZ -0.5247511 5.664507e-03 -0.7528931 2.603378e-05
1135 51123 ZNF706 -0.5884937 3.214430e-02 -0.8084932 3.126144e-03
1136 83988 NCALD 1.6421306 1.483546e-02 1.8446474 6.913283e-03
1137 51582 AZIN1 -0.8535790 2.801529e-04 -1.0227270 1.346778e-06
1138 3646 EIF3E -1.4448954 2.003885e-17 -1.2243319 4.831108e-17
1139 7227 TRPS1 0.7751844 1.354956e-02 0.7603250 2.567591e-02
1140 8667 EIF3H -0.7123231 9.434711e-07 -0.5797577 1.247850e-03
1141 5168 ENPP2 -1.4693316 1.122237e-07 -1.1867265 3.192238e-06
1142 157378 TMEM65 1.2862627 1.568367e-04 1.2341949 2.760661e-04
1143 6713 SQLE -1.2353531 5.249112e-07 -1.1082942 5.359539e-04
1144 4062 LY6H 1.8013510 9.983239e-05 2.2937684 1.653706e-09
1145 1936 EEF1D -0.8904676 2.286553e-02 -0.9067862 4.098628e-02
1146 2907 GRINA 1.5349975 5.482041e-53 1.3910913 5.752684e-41
1147 8733 GPAA1 0.5220514 1.721679e-02 0.5497770 2.954568e-02
1148 414919 C8orf82 1.1133290 2.775911e-03 1.1195292 1.022057e-02
1149 65265 C8orf33 1.5395053 3.492828e-06 1.7829171 4.361312e-10
1150 23189 KANK1 1.3522040 9.534509e-10 1.0487020 4.857602e-08
1151 7436 VLDLR -1.4084607 5.876598e-03 -1.3625483 8.221977e-03
1152 4781 NFIB 1.0576175 4.980551e-11 1.2948008 3.376685e-14
1153 158326 FREM1 -1.8314618 6.374590e-03 -2.0223534 2.784290e-03
1154 10670 RRAGA -0.6663678 2.029789e-03 -0.5921200 7.947954e-03
1155 1029 CDKN2A 1.2077224 3.173660e-08 0.9230954 1.122198e-03
1156 100129250 SMIM27 0.5825178 3.645523e-02 0.8522364 6.301319e-05
1157 10488 CREB3 1.2460254 2.015862e-04 1.2922152 1.513145e-03
1158 347252 IGFBPL1 2.2282164 3.374804e-11 1.1530911 4.282239e-02
1159 23670 CEMIP2 -1.1221064 6.404097e-07 -0.9477786 2.324785e-04
1160 7763 ZFAND5 -1.0843667 1.588232e-04 -0.7649882 1.691653e-02
1161 301 ANXA1 -1.3431326 2.241250e-20 -1.2152902 7.856257e-11
1162 5125 PCSK5 0.8934213 8.467466e-03 1.0360546 3.047012e-02
1163 7091 TLE4 0.6382555 7.758070e-09 0.4169314 7.094943e-03
1164 7088 TLE1 -1.8155925 1.716142e-04 -1.0141598 3.662671e-02
1165 2619 GAS1 -1.4648877 3.038473e-04 -2.2040747 8.323667e-08
1166 53358 SHC3 0.9262683 9.956158e-04 0.9269721 1.283522e-03
1167 84270 CARD19 -1.0415437 7.583025e-04 -0.6751315 1.294730e-02
1168 23196 FAM120A -0.4236324 1.512396e-02 -0.6795777 4.824212e-04
1169 10541 ANP32B -0.6589726 8.645788e-05 -0.8931879 3.350677e-05
1170 9568 GABBR2 1.0262131 4.391028e-02 1.8380988 4.032745e-07
1171 10952 SEC61B -0.6166122 1.576844e-05 -0.5613319 9.947854e-03
1172 23732 FRRS1L 2.8744991 3.542662e-04 2.7652934 1.314673e-03
1173 158399 ZNF483 2.8536433 2.203106e-08 2.5895244 2.209743e-03
1174 9550 ATP6V1G1 -0.4754534 2.910711e-03 -0.4181778 1.424583e-02
1175 7099 TLR4 -1.0528558 1.790711e-03 -0.8600661 4.274876e-02
1176 1620 BRINP1 -0.7489547 2.710170e-02 -1.4410807 2.107563e-03
1177 55755 CDK5RAP2 0.5912534 1.092101e-02 0.6132454 1.205363e-02
1178 169611 OLFML2A -2.3305029 4.853510e-02 -6.4601904 4.314329e-05
1179 11224 RPL35 -0.7050636 3.874540e-06 -0.7387057 4.147536e-05
1180 3309 HSPA5 -0.3808976 4.279457e-03 -0.7013340 9.275859e-10
1181 2801 GOLGA2 0.7559263 3.485763e-05 0.5452990 2.941887e-02
1182 81605 URM1 0.6158857 3.723992e-02 0.7681568 6.155077e-03
1183 89891 DYNC2I2 1.4614778 3.925263e-03 1.8047762 1.491239e-04
1184 883 KYAT1 2.4885843 1.199615e-04 1.8104104 3.632698e-02
1185 23048 FNBP1 0.6397174 6.488380e-05 0.6757149 1.367674e-07
1186 23064 SETX 0.7723261 4.244227e-03 0.9832935 1.506867e-03
1187 266655 BRD3OS 0.8046130 1.111845e-02 0.8628447 6.304220e-03
1188 8019 BRD3 0.8545019 9.394896e-05 0.7572487 1.306473e-04
1189 10422 UBAC1 1.1458949 3.309420e-08 0.9976798 6.705950e-06
1190 138151 NACC2 -0.9237412 2.595748e-08 -0.5638195 2.186773e-02
1191 4851 NOTCH1 -0.4646589 3.078629e-02 -0.5488352 3.300728e-02
1192 29085 PHPT1 0.5660772 1.850625e-05 0.5193602 1.091906e-04
1193 8721 EDF1 -1.0902562 1.172236e-08 -1.2199517 5.335663e-12
1194 26012 NSMF 0.9098442 5.205922e-03 0.8318681 1.451284e-02
1195 64975 MRPL41 0.7681481 4.830750e-05 0.4915816 2.435590e-02
1196 4549 <NA> 1.2454552 1.138345e-07 0.6615865 3.076121e-07
1197 4577 <NA> 1.4925311 1.377281e-12 1.0180558 4.740407e-06
1198 4550 <NA> 0.9482599 6.394072e-19 0.7538563 3.384186e-13
1199 4535 MT-ND1 -0.8328308 4.935918e-17 -0.9112035 1.030044e-20
1200 4565 <NA> 1.6392351 1.082964e-02 2.0354395 3.830498e-03
1201 4572 <NA> 1.2552975 1.252343e-15 1.3358005 6.181298e-06
1202 4569 <NA> 2.6387640 1.781356e-10 2.2340554 1.635875e-06
1203 4536 MT-ND2 0.5553390 5.106459e-11 0.5067176 1.082195e-08
1204 4578 <NA> 1.9672975 3.056260e-05 1.3723878 3.326214e-02
1205 4570 <NA> 0.9449237 6.122967e-13 0.9874331 6.552169e-12
1206 4579 <NA> 0.9604742 1.704628e-07 0.8534016 3.024507e-05
1207 4513 MT-CO2 -0.5329465 2.494799e-06 -0.6268429 1.895303e-09
1208 4509 MT-ATP8 1.1981271 1.295503e-21 1.0473691 6.355908e-06
1209 4508 MT-ATP6 -0.6794116 9.314834e-13 -0.9285691 2.426533e-34
1210 4514 MT-CO3 -0.2902918 3.011073e-03 -0.4383352 4.361843e-07
1211 4538 MT-ND4 -0.7403678 1.312136e-14 -0.8838916 1.368263e-25
1212 4564 <NA> 2.5719804 1.628081e-05 2.8232814 8.323667e-08
1213 4575 <NA> 3.0493965 3.521144e-07 2.1281057 1.157273e-02
1214 4540 MT-ND5 1.1911849 2.126822e-31 0.8676954 7.463550e-19
1215 4541 MT-ND6 0.8500680 1.826693e-13 0.4351731 4.429108e-04
1216 4519 MT-CYB -0.5003529 3.080074e-10 -0.6889478 4.827200e-17
1217 4571 <NA> 3.4044633 2.362771e-119 2.8563893 1.746726e-69
1218 8905 AP1S2 -0.9664051 6.101350e-23 -0.7749899 4.260762e-21
1219 94056 SYAP1 0.6064412 5.684941e-03 0.7330270 1.867686e-04
1220 10549 PRDX4 -1.6675877 3.694873e-10 -0.9225504 6.156778e-04
1221 10159 ATP6AP2 -0.8275559 4.426843e-07 -0.6952396 3.103277e-05
1222 54539 NDUFB11 -0.7516247 1.103419e-02 -1.1640156 1.678749e-04
1223 56850 GRIPAP1 0.6015785 4.095064e-02 0.6798442 1.516274e-02
1224 57477 SHROOM4 1.0039269 5.299002e-03 1.0496827 5.380893e-03
1225 728239 MAGED4 1.4320558 1.426777e-02 1.2798849 4.379649e-02
1226 3028 HSD17B10 0.8381355 1.739762e-03 1.0876016 1.345276e-06
1227 54552 GNL3L 1.2637831 6.868196e-07 1.0683116 3.149558e-04
1228 550643 NBDY -0.7225578 3.803228e-03 -0.6603237 3.214588e-03
1229 442454 <NA> -0.6373378 3.749090e-03 -0.4452347 4.347245e-02
1230 158586 ZXDB -3.1663152 1.104407e-03 -1.4865472 3.632698e-02
1231 9843 HEPH -1.7205327 1.415837e-04 -3.0352356 9.922827e-06
1232 1947 EFNB1 0.9924894 2.290356e-04 0.8597194 1.249707e-02
1233 340527 NHSL2 2.3301845 4.622585e-05 2.1087034 2.424107e-04
1234 5303 PIN4 -0.8835492 3.820335e-03 -0.8437007 3.188040e-03
1235 6191 RPS4X -0.5281537 1.614387e-05 -0.4433029 2.976157e-03
1236 441531 PGAM4 -0.7227182 1.497859e-02 -0.6859448 3.628419e-02
1237 5230 PGK1 -2.2418518 1.719764e-80 -1.2305879 2.661201e-26
1238 2846 LPAR4 1.4091085 8.177171e-08 1.2141436 5.276508e-05
1239 79366 HMGN5 -1.6773764 2.458787e-05 -2.1269753 1.565924e-07
1240 7105 TSPAN6 -0.4770235 2.708350e-02 -0.6324670 1.385789e-03
1241 27286 SRPX2 0.9507795 5.249112e-07 0.7486537 6.254085e-04
1242 94121 SYTL4 -2.1392611 2.369016e-12 -1.2710173 2.146449e-05
1243 51566 ARMCX3 0.5644596 3.988277e-02 0.9271075 2.139683e-04
1244 9823 ARMCX2 -0.9830875 1.782986e-04 -0.9581115 3.435068e-03
1245 84460 ZMAT1 -0.8542285 3.925541e-02 -1.3734403 2.677183e-03
1246 340543 TCEAL5 1.1268906 3.934522e-07 1.0011318 2.095923e-05
1247 9643 MORF4L2 -0.6747360 5.247299e-03 -0.4717278 3.263452e-02
1248 1641 DCX -0.8457536 7.306344e-03 -0.9610581 5.303766e-03
1249 5358 PLS3 -1.5233899 4.816454e-06 -1.0025877 4.267737e-03
1250 10857 PGRMC1 -0.7314699 2.612498e-04 -0.6340352 7.388822e-03
1251 4694 NDUFA1 -0.5530361 2.258081e-05 -0.5041327 5.871543e-04
1252 55026 TMEM255A -1.1946326 4.352120e-03 -1.1077157 1.121250e-02
1253 2239 GPC4 -2.1473273 5.147271e-06 -1.2110122 1.201094e-02
1254 2719 GPC3 -0.5884722 2.270755e-03 -2.9646777 6.067464e-67
1255 27316 RBMX 1.1630289 9.116097e-16 0.6873829 4.661108e-05
1256 1038 CDR1 3.7692197 4.289889e-59 3.9196558 7.957925e-73
1257 83692 CD99L2 0.8195141 5.631666e-04 0.7089770 1.177798e-03
1258 29944 PNMA3 5.4634513 2.323953e-04 4.9629362 1.922443e-02
1259 84968 PNMA6A 2.4873948 4.238186e-08 2.3903937 1.589204e-06
1260 4204 MECP2 0.8163152 1.867538e-02 0.7692576 3.750467e-02
1261 2316 FLNA 0.4895471 2.194210e-04 0.3709100 1.352675e-02
1262 2010 EMD 0.8932399 9.310564e-07 0.8302233 8.241662e-06
1263 7411 VBP1 -0.7364146 6.114222e-03 -0.4752162 4.356118e-02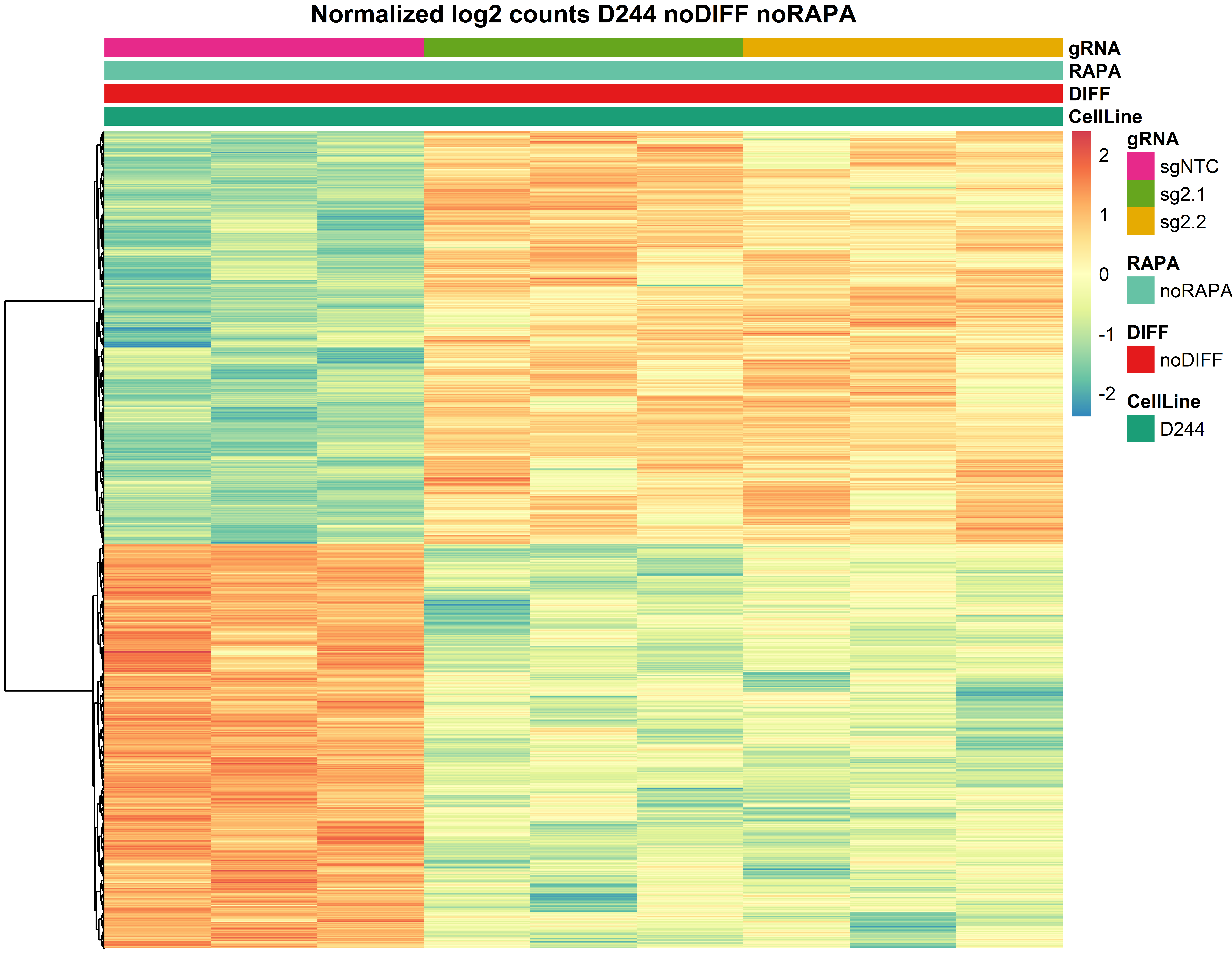
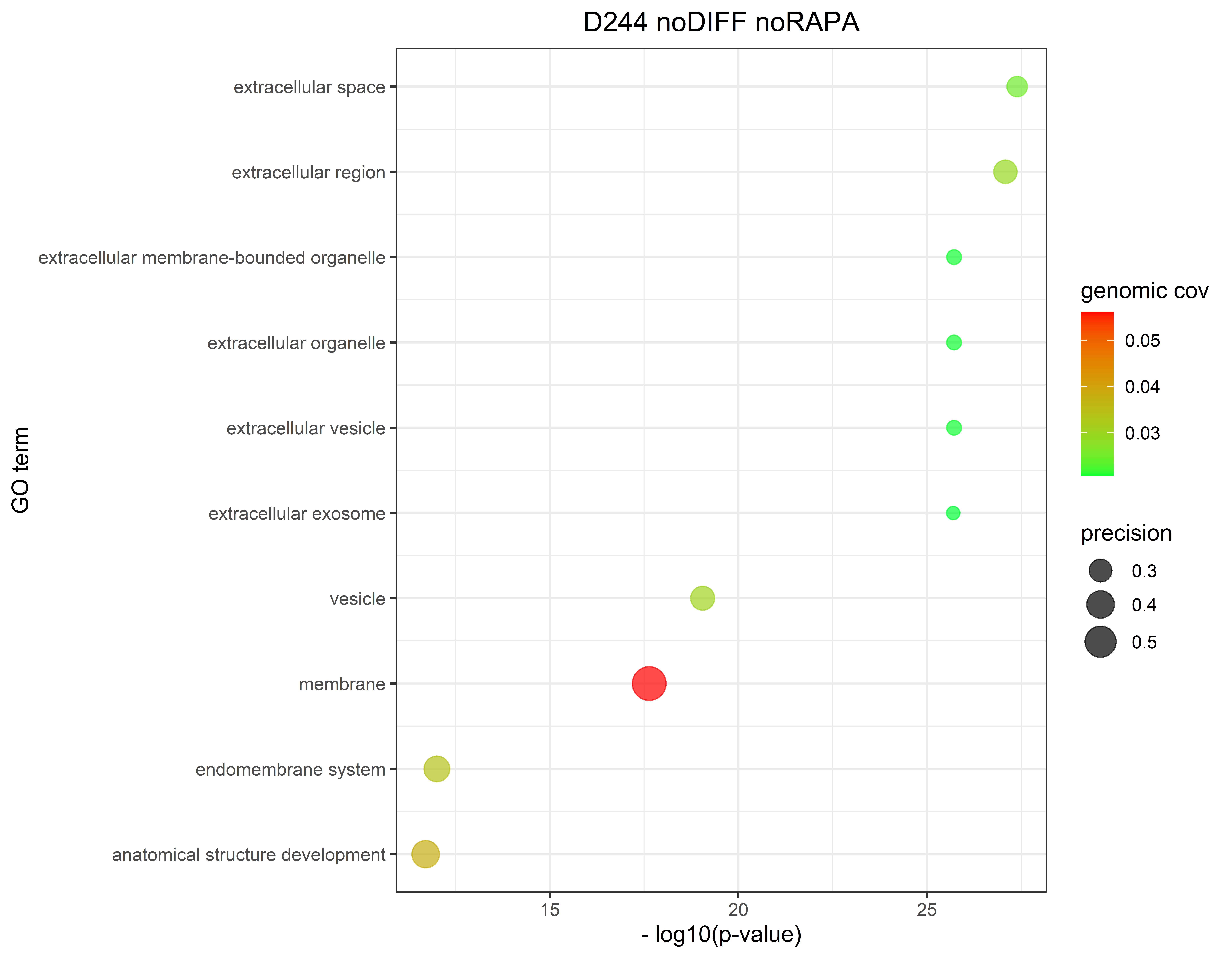
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.2004215 0.9049203 1.570216 1.908726e-01
02_TSC2_KO_Grabole_2016 1.4255145 1.2645679 1.606005 5.064721e-09
03_TSC2_Grabole-2016 1.4160309 1.2575323 1.595280 5.493296e-09
04_TSC2-tuber-Martin2017 2.1323041 1.2404072 3.500877 4.664193e-03
05_TSC2-SEN_SEGA-Martin2017 1.2105824 1.0290531 1.419102 1.969641e-02
06_all_EpilepsyGene 1.2232596 0.8608799 1.699740 2.425478e-01
07_high_conf_EpilepsyGene 1.1406076 0.5870818 2.039154 6.405796e-01
08_ASD_Epilepsy_EpilepsyGene 1.4498686 0.7801469 2.516390 1.964988e-01
09_Epi25_dataset 1.0663946 0.4427013 2.220974 8.472133e-01
10_SFARI_all 1.3711794 0.8919403 2.039237 1.140568e-01
11_SFARI_score4plus 1.5564891 0.9633182 2.416097 4.943506e-02
12_DeRubeis_RDNV 2.3692151 1.2478920 4.230333 5.024359e-03
13_Iossifov_RDNV 1.3502214 1.0090736 1.780511 3.517499e-02
14_FMRP_Darnell 1.9356981 1.5642182 2.380164 2.958261e-09
15_Voineagu_M12 0.6363092 0.3869344 0.994238 4.643397e-02
16_Voineagu_M16 2.7503442 2.0789844 3.601668 9.031342e-12
17_ID_genes_Neel 1.0676875 0.7134921 1.549303 6.990195e-01
18_ID_genes_pinto2014 1.2911033 0.8341784 1.930107 2.167127e-01
19_Fromer_SCZ_FunDN 1.2638669 0.9332924 1.684041 1.133299e-01
20_Cocchi_Pruning 1.9566540 1.0403460 3.449224 2.649349e-02
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.18267273 0.14417404 1.000000e+00
02_TSC2_KO_Grabole_2016 0.35453283 0.06112364 1.012944e-07
03_TSC2_Grabole-2016 0.34785780 0.06056455 1.098659e-07
04_TSC2-tuber-Martin2017 0.75720312 0.27640988 9.328387e-02
05_TSC2-SEN_SEGA-Martin2017 0.19110154 0.08288902 3.939282e-01
06_all_EpilepsyGene 0.20151907 0.17924455 1.000000e+00
07_high_conf_EpilepsyGene 0.13156107 0.33885319 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 0.37147292 0.31619692 1.000000e+00
09_Epi25_dataset 0.06428339 0.44854259 1.000000e+00
10_SFARI_all 0.31567124 0.21940169 1.000000e+00
11_SFARI_score4plus 0.44243273 0.24479809 9.887012e-01
12_DeRubeis_RDNV 0.86255871 0.32709335 1.004872e-01
13_Iossifov_RDNV 0.30026858 0.14858972 7.034999e-01
14_FMRP_Darnell 0.66046802 0.10871525 5.916522e-08
15_Voineagu_M12 -0.45207062 0.25379061 9.286794e-01
16_Voineagu_M16 1.01172608 0.14277887 1.806268e-10
17_ID_genes_Neel 0.06549505 0.20565255 1.000000e+00
18_ID_genes_pinto2014 0.25549714 0.22285977 1.000000e+00
19_Fromer_SCZ_FunDN 0.23417599 0.15470039 1.000000e+00
20_Cocchi_Pruning 0.67123588 0.32228701 5.298698e-01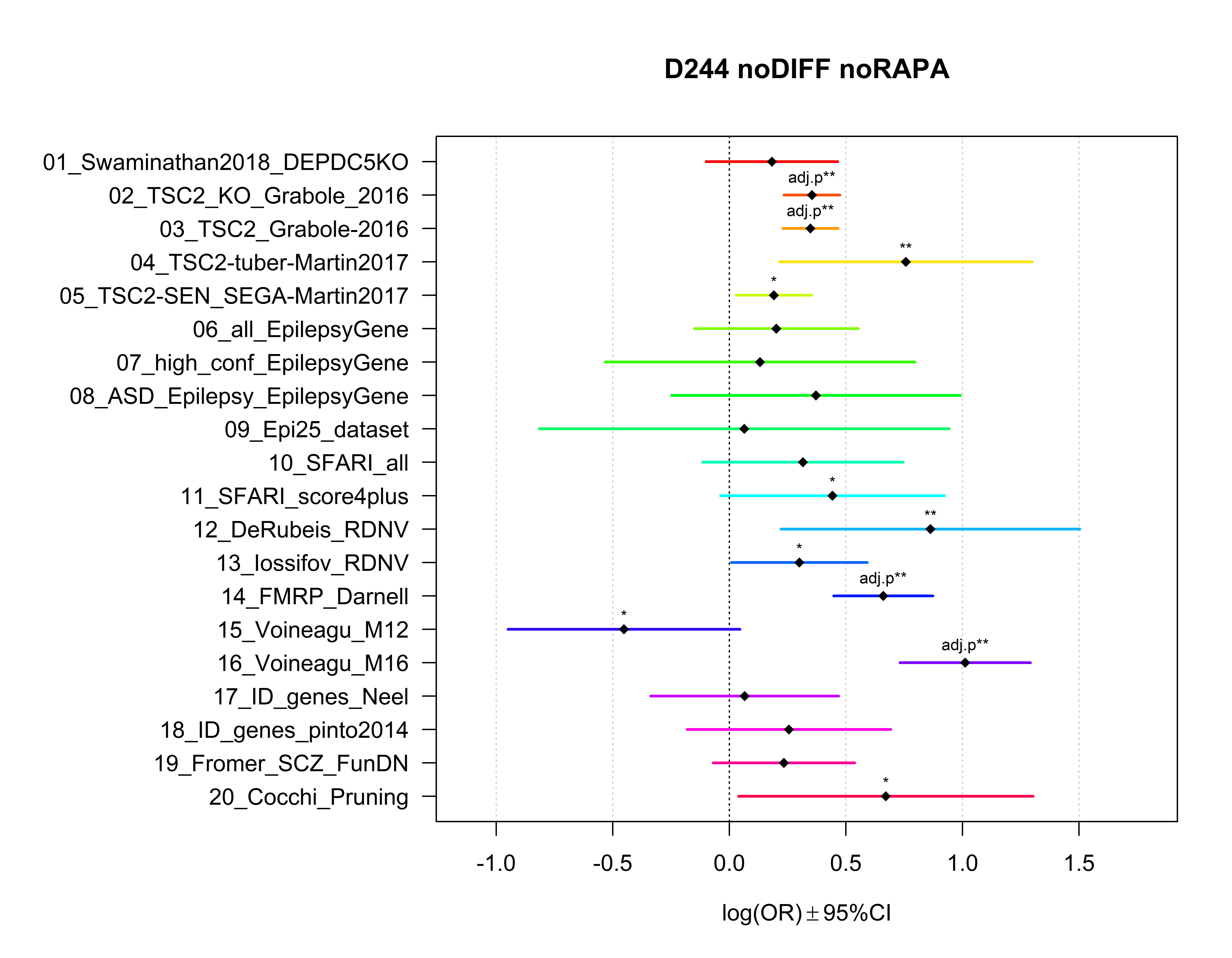
ReN
getlinmodoutput(targetline = "ReN",targetdiff = "noDIFF",targetrapa = "noRAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 55920 RCC2 -0.7709626 4.129780e-05 -0.2645249 3.812275e-02
2 2152 F3 -1.6052562 4.812020e-05 -1.1057348 6.860234e-03
3 116496 NIBAN1 -0.8567869 2.608034e-02 -0.9061142 1.725769e-07
4 10625 IVNS1ABP 1.0111945 1.290996e-02 0.8932301 1.349441e-03
5 1316 KLF6 0.6226359 6.285458e-03 0.4374648 2.393860e-03
6 27063 ANKRD1 0.7284664 8.767738e-03 0.6484243 1.845423e-03
7 10410 IFITM3 0.8639180 1.900099e-04 0.8986399 1.358229e-15
8 9415 FADS2 -0.5463766 1.048817e-02 -0.4041426 7.790489e-04
9 6876 TAGLN 0.5665027 3.412600e-03 0.3089471 3.634781e-05
10 8788 DLK1 1.3271225 2.339702e-06 1.0858472 4.185648e-07
11 55384 MEG3 0.6186050 6.986237e-03 0.3983264 1.395902e-02
12 7168 TPM1 0.6153430 9.929395e-04 0.7254517 9.832148e-29
13 8482 SEMA7A -1.1493099 2.996955e-02 -1.3520616 1.078898e-04
14 9289 ADGRG1 0.5742315 1.781405e-02 0.4019053 1.383476e-02
15 6560 SLC12A4 1.3887778 5.524029e-03 1.0250149 9.911553e-03
16 2670 GFAP 0.7380134 2.981670e-02 0.3964766 1.934040e-04
17 1277 COL1A1 1.1694356 3.505403e-08 0.7581812 3.017104e-08
18 114897 C1QTNF1 -1.6933139 4.743532e-04 -1.0442519 2.674684e-06
19 6597 SMARCA4 0.9510221 8.633707e-08 0.3447233 4.739736e-02
20 3589 IL11 1.0942977 7.595885e-10 0.7528246 5.200636e-17
21 7837 PXDN 1.2479747 7.686976e-10 0.8975230 3.194388e-13
22 6664 SOX11 0.5491863 2.902291e-02 0.4725256 3.590150e-03
23 64838 FNDC4 1.0766732 3.651082e-02 0.7477309 3.004000e-02
24 57142 RTN4 0.6548997 3.022107e-02 0.5092739 6.213421e-04
25 72 ACTG2 1.5107705 4.003181e-04 1.6657161 1.910836e-09
26 1622 DBI 0.6069698 9.572129e-03 0.2668891 5.430482e-03
27 8324 FZD7 -0.9560900 2.981670e-02 -0.9506187 3.864097e-04
28 3485 IGFBP2 0.4572250 8.559464e-03 0.2640600 1.483333e-02
29 547 KIF1A 1.0191769 3.837016e-03 0.8084061 1.349441e-03
30 1299 COL9A3 0.6652539 7.257546e-03 0.5082300 8.421927e-09
31 1291 COL6A1 -0.7239440 2.337870e-05 -1.3280713 2.904372e-67
32 1292 COL6A2 -0.9028436 6.275974e-03 -1.8399877 1.015325e-15
33 6285 S100B 1.0618307 2.499636e-04 1.1014556 4.978164e-11
34 151887 CCDC80 1.3269840 1.910378e-09 0.5349808 2.674684e-06
35 4071 TM4SF1 0.7983308 7.460130e-03 0.5525958 2.134321e-02
36 55752 SEPTIN11 0.5614741 2.309992e-02 0.6041739 5.856423e-06
37 10769 PLK2 -1.0628661 2.897441e-02 -0.9051113 1.805202e-02
38 2297 FOXD1 -1.1949549 3.907876e-02 -1.1498402 3.107899e-02
39 1839 HBEGF 0.7418055 3.092105e-03 0.7139983 4.059826e-08
40 8728 ADAM19 1.3682698 4.303558e-05 1.4944132 5.024004e-12
41 64065 PERP -0.9466032 5.083185e-05 -0.6466159 3.712832e-03
42 6624 FSCN1 0.6434701 1.733283e-06 0.3170077 5.802062e-05
43 800 CALD1 0.7665582 1.166677e-04 0.8241889 3.424143e-30
44 5764 PTN 0.5227649 2.820302e-02 0.7267524 3.128694e-23
45 1052 CEBPD 0.6269253 4.612829e-02 0.5429092 1.433602e-02
46 641638 <NA> 0.6140200 1.963879e-02 0.3201642 1.862983e-02
47 793 CALB1 -1.1418089 6.542001e-09 -1.3893524 3.320383e-23
48 28998 MRPL13 -0.9837532 2.905522e-04 -0.4863646 2.251966e-02
49 1515 CTSV -1.3687163 3.534266e-06 -2.5548099 1.628854e-55
50 9568 GABBR2 0.8591791 1.299630e-04 0.7804877 1.030234e-16
51 3371 TNC -1.0395490 1.382104e-06 -0.3561205 4.590609e-02
52 1289 COL5A1 -1.5101448 7.312207e-03 -1.1785048 4.211205e-07
53 4539 MT-ND4L 0.4547687 1.940962e-02 0.2676648 4.270763e-03
54 92745 SLC38A5 1.3495871 6.273324e-07 0.8102770 4.087766e-06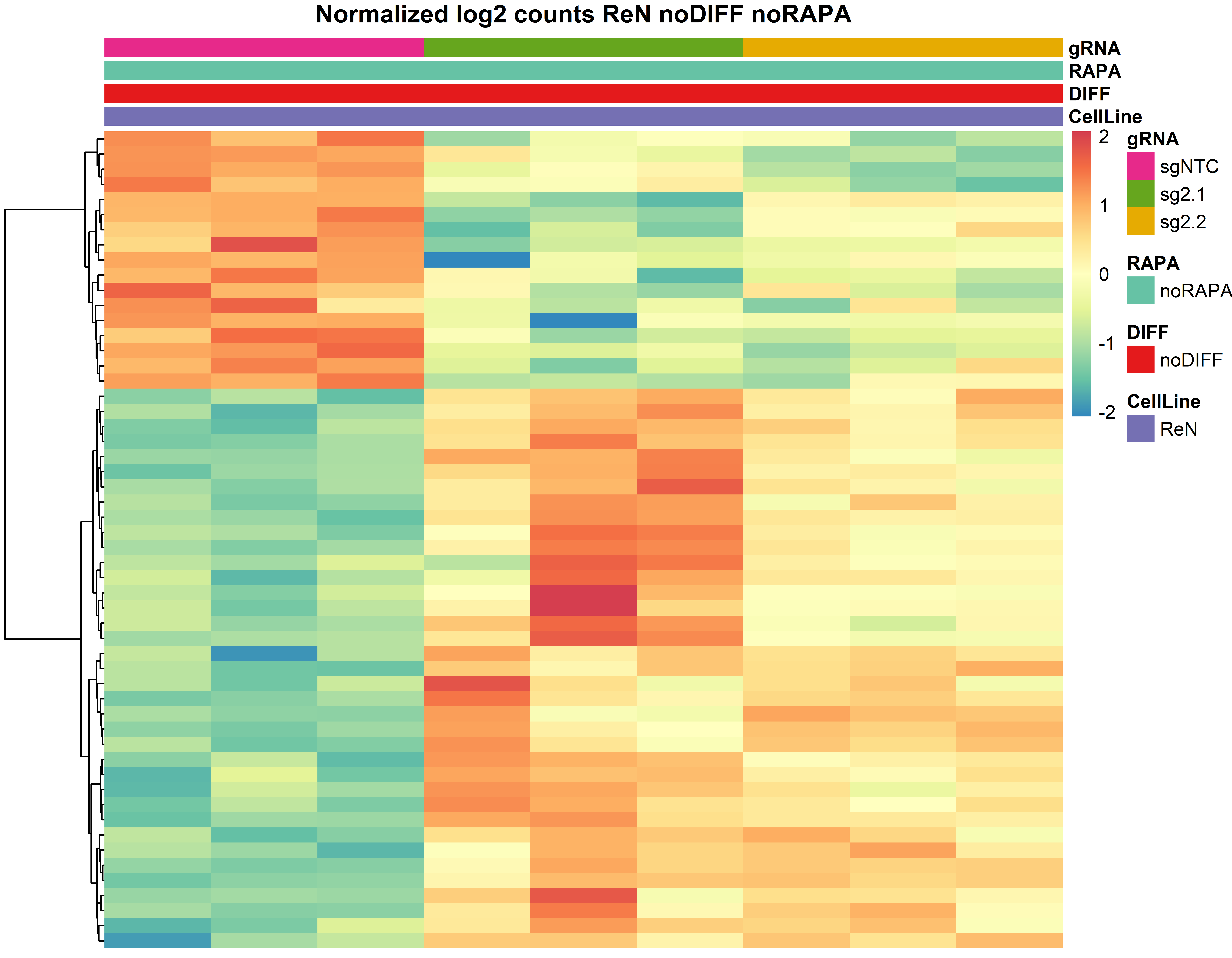
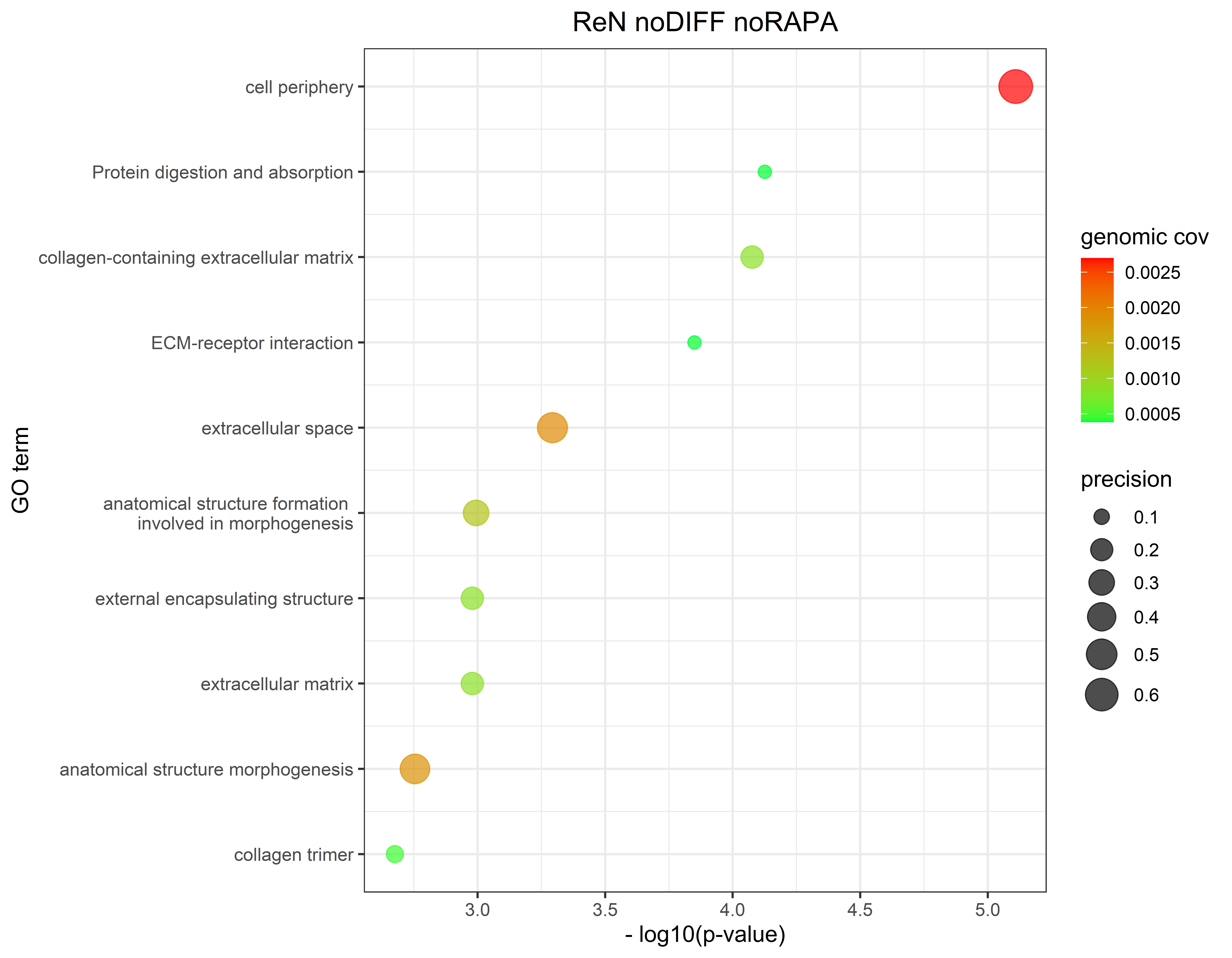
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.8487259 0.09989353 3.237908 1.0000000000
02_TSC2_KO_Grabole_2016 1.3967597 0.77680995 2.477698 0.2518780221
03_TSC2_Grabole-2016 1.4536366 0.81785072 2.628758 0.2194194589
04_TSC2-tuber-Martin2017 7.2212983 1.41995499 22.889012 0.0100471083
05_TSC2-SEN_SEGA-Martin2017 2.1290741 1.06776822 4.006525 0.0185447289
06_all_EpilepsyGene 1.3418222 0.15775609 5.130774 0.6634089148
07_high_conf_EpilepsyGene 2.0509246 0.05057317 12.147996 0.3919490878
08_ASD_Epilepsy_EpilepsyGene 2.1950289 0.05409370 13.015784 0.3720290812
09_Epi25_dataset 0.0000000 0.00000000 11.970736 1.0000000000
10_SFARI_all 0.0000000 0.00000000 4.002205 1.0000000000
11_SFARI_score4plus 0.0000000 0.00000000 5.436925 1.0000000000
12_DeRubeis_RDNV 0.0000000 0.00000000 12.602077 1.0000000000
13_Iossifov_RDNV 0.4853168 0.01204121 2.836155 0.7236633027
14_FMRP_Darnell 1.7411550 0.53979560 4.364462 0.2245150282
15_Voineagu_M12 0.0000000 0.00000000 2.758299 0.6468121238
16_Voineagu_M16 5.8663885 2.22056745 13.187271 0.0003854323
17_ID_genes_Neel 0.7693916 0.01906445 4.506239 1.0000000000
18_ID_genes_pinto2014 3.2898328 0.65218159 10.289161 0.0704745817
19_Fromer_SCZ_FunDN 1.5676565 0.31188937 4.874852 0.4470978815
20_Cocchi_Pruning 0.0000000 0.00000000 10.741481 1.0000000000
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -0.1640190 1.0916487 1.000000000
02_TSC2_KO_Grabole_2016 0.3341551 0.2993442 1.000000000
03_TSC2_Grabole-2016 0.3740684 0.2934408 1.000000000
04_TSC2-tuber-Martin2017 1.9770348 0.8298008 0.200942165
05_TSC2-SEN_SEGA-Martin2017 0.7556872 0.3521003 0.370894578
06_all_EpilepsyGene 0.2940285 1.0922111 1.000000000
07_high_conf_EpilepsyGene 0.7182907 1.8890943 1.000000000
08_ASD_Epilepsy_EpilepsyGene 0.7861952 1.8894045 1.000000000
09_Epi25_dataset -Inf NaN 1.000000000
10_SFARI_all -Inf NaN 1.000000000
11_SFARI_score4plus -Inf NaN 1.000000000
12_DeRubeis_RDNV -Inf NaN 1.000000000
13_Iossifov_RDNV -0.7229534 1.8859525 1.000000000
14_FMRP_Darnell 0.5545487 0.5975068 1.000000000
15_Voineagu_M12 -Inf NaN 1.000000000
16_Voineagu_M16 1.7692392 0.4956512 0.007708647
17_ID_genes_Neel -0.2621552 1.8866199 1.000000000
18_ID_genes_pinto2014 1.1908367 0.8256474 1.000000000
19_Fromer_SCZ_FunDN 0.4495818 0.8238207 1.000000000
20_Cocchi_Pruning -Inf NaN 1.000000000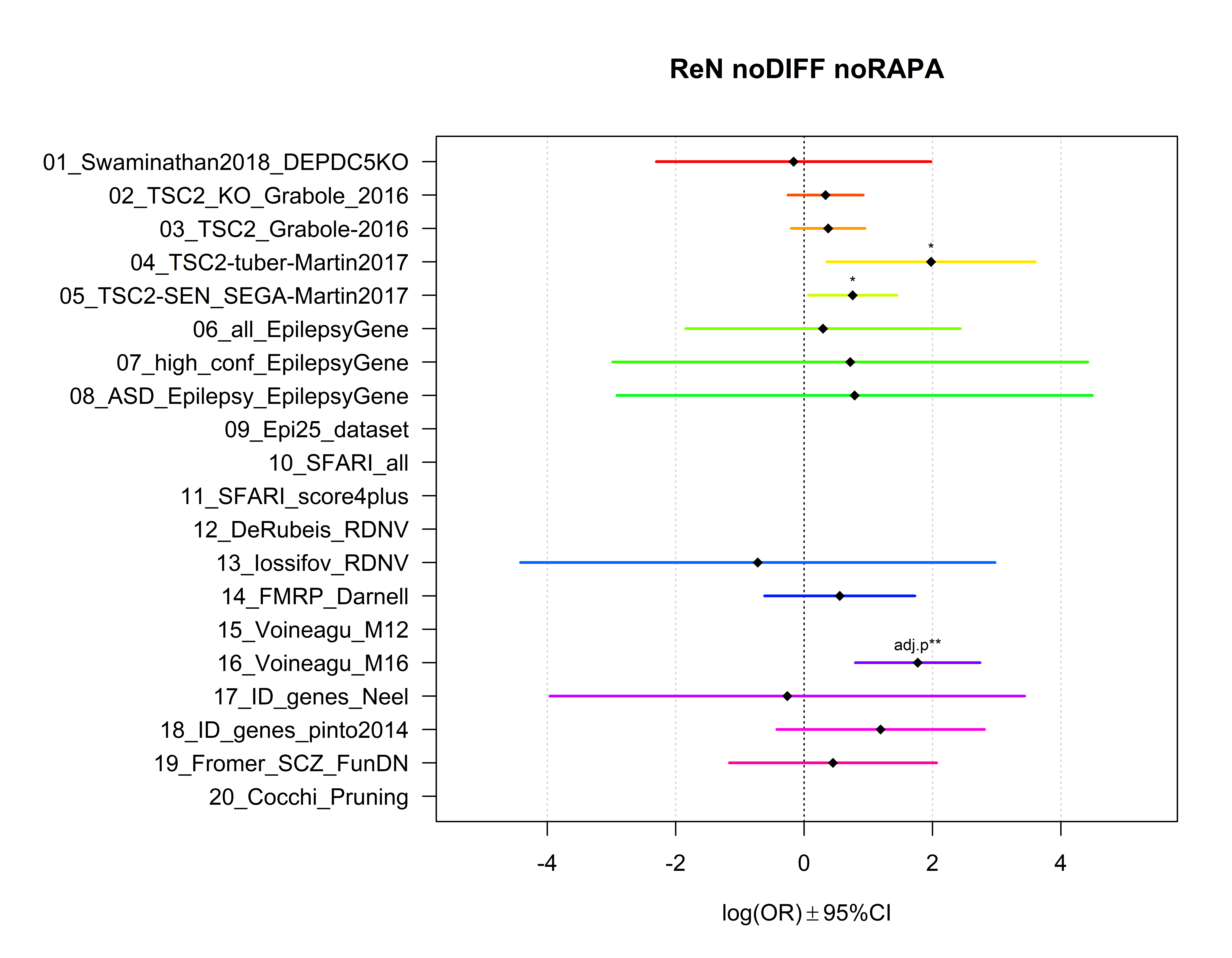
KO effect in DIFF noRAPA
D62
getlinmodoutput(targetline = "D62",targetdiff = "DIFF",targetrapa = "noRAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 51150 SDF4 -0.4955025 2.579359e-03 -0.5953468 1.781040e-04
2 64856 VWA1 -0.5621746 3.771578e-03 -1.4107532 1.828404e-14
3 1953 MEGF6 -1.0425165 1.794545e-02 -2.6466407 8.328495e-05
4 26038 CHD5 -1.2094340 9.476205e-03 -1.2741363 2.815505e-02
5 9249 DHRS3 -1.3097280 2.111936e-15 -1.2296776 2.617139e-18
6 2048 EPHB2 -0.8641387 3.887602e-09 -1.0985671 1.035345e-12
7 3925 STMN1 0.7944170 3.016039e-07 1.4274906 1.254741e-25
8 2537 IFI6 -0.8433904 4.332154e-06 -0.8189382 2.284061e-04
9 10657 KHDRBS1 0.4700116 6.464077e-03 0.4598147 2.286086e-02
10 65108 MARCKSL1 0.5000844 3.804763e-02 0.9989000 1.168573e-06
11 6421 SFPQ 0.4515981 1.693582e-02 0.5128704 1.572492e-02
12 63967 CLSPN 1.5137939 2.851279e-02 1.6319799 4.375498e-02
13 84879 MFSD2A 0.9763791 9.131033e-06 1.8248040 1.138677e-20
14 6513 SLC2A1 -0.5300470 2.822089e-02 -0.7679623 2.811444e-03
15 1996 ELAVL4 3.2608311 4.962903e-06 4.0652987 4.144930e-09
16 5865 RAB3B 3.5894843 1.843733e-05 4.3813828 3.762860e-08
17 3725 JUN 0.7298672 1.942181e-08 0.5537306 4.001398e-03
18 4774 NFIA 0.5175837 7.823789e-04 0.7298703 1.415834e-05
19 91624 NEXN 1.0028329 9.119499e-04 1.2187668 3.222356e-05
20 3491 CCN1 1.1287146 4.877936e-14 1.2278310 2.873031e-13
21 9411 ARHGAP29 1.7634825 1.769450e-02 1.8714486 3.230913e-02
22 84722 PSRC1 -0.3789217 2.535810e-02 -0.5226016 9.529778e-03
23 3752 KCND3 4.4704034 1.059294e-02 4.7438914 1.505603e-02
24 7482 WNT2B -1.4543823 3.547978e-03 -3.1408787 3.528764e-02
25 54879 ST7L -1.1880679 3.140635e-03 -2.0706668 1.253519e-07
26 389 RHOC -0.7041464 5.115743e-04 -0.7909251 2.023271e-04
27 476 ATP1A1 -1.0997358 6.225788e-09 -0.6174432 9.665606e-03
28 10628 TXNIP -1.0151132 8.325040e-03 -1.5573992 9.558770e-04
29 25832 NBPF14 -0.5872187 1.314383e-02 -0.6992549 9.981604e-03
30 4170 MCL1 0.5884118 8.485303e-03 0.5492956 3.046150e-02
31 10962 MLLT11 0.6808427 3.689879e-03 0.9870859 3.654889e-04
32 6282 S100A11 -0.6620088 2.674714e-04 -1.4333294 1.723954e-08
33 112770 GLMP -1.1545239 1.783433e-04 -0.8114086 1.604436e-02
34 10485 MIR9-1HG 0.3922581 1.894780e-03 0.4052353 2.092622e-02
35 93183 PIGM -0.8527513 2.001797e-03 -0.9327189 2.348428e-03
36 477 ATP1A2 1.4390816 4.482580e-05 2.0217080 1.246246e-09
37 5999 RGS4 1.4843432 1.246320e-02 1.8144199 4.411380e-03
38 387597 ILDR2 -0.7472582 3.654285e-04 -0.8916310 1.466669e-04
39 481 ATP1B1 1.0621640 5.828589e-03 1.2966800 1.205500e-03
40 23057 NMNAT2 3.4158547 8.593309e-04 3.5208565 3.846423e-03
41 23127 COLGALT2 -0.6453155 4.170684e-03 -1.0025729 2.990741e-06
42 5997 RGS2 2.3549801 2.908959e-03 2.4140555 1.135038e-02
43 23046 KIF21B 1.4025501 4.017314e-02 2.9011030 2.747197e-10
44 23612 PHLDA3 -0.9978168 8.918304e-08 -0.5631703 2.399817e-02
45 89796 NAV1 0.9189098 7.922638e-05 1.2236795 1.733921e-08
46 55220 KLHDC8A 0.4315198 3.954693e-02 1.1871507 1.409696e-12
47 254428 SLC41A1 -0.6641172 1.320366e-02 -1.0159950 5.463962e-04
48 50486 G0S2 -1.2769869 3.776809e-02 -3.7478581 2.744552e-36
49 1063 CENPF 0.9674909 1.276505e-04 0.6984687 1.927045e-02
50 3930 LBR 1.1691924 2.189956e-04 1.1824021 1.402408e-03
51 7044 LEFTY2 -1.8967514 2.076460e-02 -5.4919034 1.299317e-06
52 56776 FMN2 1.8223429 1.818342e-05 1.4585189 1.179215e-02
53 5214 PFKP -0.7085246 4.103217e-03 -0.9698443 1.182711e-03
54 1316 KLF6 0.4591444 4.092246e-02 0.9649889 2.229439e-06
55 5209 PFKFB3 0.9025962 1.244501e-06 0.6596269 5.396264e-03
56 56243 KIAA1217 -1.0092171 1.487638e-02 -1.6358087 4.374010e-04
57 57512 GPR158 6.2531163 1.041083e-02 6.1967773 4.088838e-02
58 3185 HNRNPF 0.4924999 1.043753e-02 0.5434778 3.751870e-02
59 118738 ZNF488 -1.3591800 1.781176e-05 -1.5052581 3.235236e-05
60 100507008 <NA> -1.0683994 1.666217e-02 -1.1833009 2.512927e-02
61 84159 ARID5B 1.1067873 7.036831e-10 0.7182760 8.550844e-04
62 55680 RUFY2 1.4665537 2.349213e-02 1.7921536 1.081271e-02
63 140766 ADAMTS14 -1.0590339 2.629807e-02 -1.9314582 1.261456e-04
64 5660 PSAP -0.5893935 1.614984e-09 -0.6307739 1.718370e-08
65 9806 SPOCK2 -0.7006081 1.000618e-06 -0.9393863 2.122881e-09
66 5328 PLAU -0.7107724 2.041878e-02 -1.7229682 1.575495e-11
67 132 ADK 0.8457418 3.776809e-02 1.2331920 3.288831e-03
68 23522 KAT6B 0.7195214 1.092992e-02 1.1942312 9.256950e-05
69 311 ANXA11 -0.8715186 9.406076e-05 -1.0391239 7.087740e-04
70 27063 ANKRD1 -1.4181691 1.258581e-03 -2.0199847 4.082433e-04
71 9124 PDLIM1 -0.4570125 4.949466e-03 -1.8256331 1.310934e-20
72 10580 SORBS1 0.7949338 2.625106e-02 1.2342037 5.463962e-04
73 83742 MARVELD1 -1.0196936 7.773859e-03 -3.0128114 1.626479e-05
74 84171 LOXL4 -1.0820720 5.359873e-03 -2.0032920 3.739933e-05
75 25911 DPCD 1.1158041 1.179168e-03 0.9881699 3.230913e-02
76 9118 INA 2.3042902 1.740790e-09 3.0690799 1.090343e-18
77 63877 FAM204A 1.2483183 4.111141e-03 1.1495418 3.713935e-02
78 9531 BAG3 -0.6141248 2.872818e-02 -0.8006766 2.551578e-03
79 64787 EPS8L2 -1.8841773 2.413899e-02 -2.5508625 2.033169e-03
80 6181 RPLP2 -0.7084403 7.711357e-12 -0.4562317 7.819783e-05
81 977 CD151 -0.8589280 1.329522e-07 -0.7195245 1.046971e-03
82 54472 TOLLIP -0.4889611 4.074004e-02 -0.6314934 4.081618e-03
83 1509 CTSD -1.3844459 6.042722e-11 -1.1546733 1.485188e-11
84 975 CD81 -0.6898415 3.572604e-10 -0.5108469 3.696366e-05
85 1200 TPP1 -0.8395789 5.673328e-09 -0.6679043 4.991740e-06
86 283298 OLFML1 -2.5901081 3.728262e-04 -4.3858271 1.744180e-11
87 8495 PPFIBP2 -1.9365285 4.201254e-03 -2.6640963 2.508021e-02
88 6157 RPL27A -0.4484639 2.172998e-05 -0.5557320 1.299543e-07
89 56672 AKIP1 -0.8593692 2.797337e-04 -1.0236992 1.320957e-02
90 100463486 MTRNR2L8 -0.5578237 2.828289e-05 -0.7469176 7.922861e-11
91 10335 IRAG1 2.7773162 1.257846e-03 3.4613748 2.067171e-05
92 10418 SPON1 2.3371384 1.821001e-25 2.8657805 2.055482e-37
93 100126784 <NA> -1.0947445 3.748100e-02 -2.0601972 5.716200e-04
94 90139 TSPAN18 -1.2350599 9.075399e-03 -3.1872211 1.471856e-07
95 4192 MDK -0.7403873 6.530164e-10 -1.2928391 2.363787e-22
96 54972 TMEM132A -1.5686110 2.263621e-06 -1.2014332 1.988352e-02
97 1937 EEF1G -0.6993356 3.122941e-08 -0.3454940 3.230913e-02
98 6094 ROM1 -0.7216586 7.087820e-03 -0.7315965 2.598470e-02
99 51035 UBXN1 -0.6519263 5.898459e-03 -0.6014790 1.409058e-02
100 6520 SLC3A2 -0.9796559 3.111335e-12 -0.9942900 3.719212e-09
101 5920 PLAAT4 -0.8592724 7.548369e-07 -0.6612061 1.541733e-05
102 402 ARL2 -1.0917134 1.144262e-06 -0.7172434 5.602278e-03
103 4054 LTBP3 -0.8975351 1.180630e-04 -1.4336403 4.685666e-09
104 29984 RHOD -2.1359526 1.651096e-02 -3.5853260 6.808726e-03
105 54961 SSH3 -0.9106481 1.023444e-02 -1.2660190 2.632959e-03
106 80194 TMEM134 -0.8017841 5.114206e-04 -0.7902311 3.070781e-02
107 1374 CPT1A -0.5505969 4.013752e-02 -0.7928566 1.142001e-03
108 55191 NADSYN1 -1.2535007 4.782090e-04 -1.0527382 3.797532e-02
109 6188 RPS3 -0.5129980 1.255454e-06 -0.4465642 7.555908e-05
110 2893 GRIA4 1.1796260 2.790154e-02 1.3712466 1.521117e-02
111 23086 EXPH5 5.5069765 1.744228e-02 7.1085098 1.182043e-04
112 1410 CRYAB -0.9064181 2.412081e-06 -1.0487832 5.878247e-09
113 23705 CADM1 0.2874183 4.487948e-02 0.4231700 2.510206e-02
114 50937 CDON -1.2766445 1.525007e-02 -2.0779999 1.173198e-04
115 334 APLP2 -0.6772377 1.015275e-06 -0.5802617 7.651342e-08
116 84318 CCDC77 1.9658368 1.749660e-02 2.1758950 3.412593e-02
117 2597 GAPDH -0.6653002 1.981316e-10 -0.3687779 1.983706e-03
118 2 A2M 1.3571253 3.286528e-04 1.3799321 3.349517e-03
119 53919 SLCO1C1 2.1983350 3.924279e-03 3.5345281 7.474492e-10
120 51474 LIMA1 1.3709008 4.390829e-26 0.9187943 3.001160e-08
121 3164 NR4A1 1.9392213 7.720419e-03 2.9236343 1.977458e-06
122 84926 SPRYD3 -0.8912767 1.242525e-03 -1.0968278 3.581571e-04
123 3489 IGFBP6 -1.6066792 3.470231e-09 -2.8097426 3.037210e-19
124 4327 MMP19 -1.0206521 3.718852e-02 -2.3634379 1.791381e-03
125 1606 DGKA -0.8334538 8.538833e-03 -1.6919856 1.766773e-07
126 10956 OS9 -0.5352587 1.315690e-03 -0.3777409 3.741461e-02
127 4193 MDM2 -0.8774030 1.265300e-06 -0.7506638 1.422245e-05
128 10576 CCT2 0.6548356 1.818342e-05 0.5180339 2.065548e-02
129 6857 SYT1 1.6645278 3.586655e-16 1.7366885 7.554068e-12
130 1848 DUSP6 0.9472608 1.507218e-05 0.8270646 4.384776e-03
131 694 BTG1 0.8678603 2.142367e-04 1.5305075 9.496891e-11
132 7112 TMPO 1.1775440 8.630785e-03 1.2966858 9.083012e-03
133 429 ASCL1 -2.4634672 7.050646e-09 -1.5122563 1.529090e-03
134 51559 NT5DC3 -1.6497901 1.121657e-04 -2.5550160 6.192312e-07
135 7296 TXNRD1 0.5299716 2.765136e-02 0.6784375 4.983759e-02
136 5992 RFX4 0.8119488 3.438064e-02 1.1235226 4.497281e-03
137 9921 RNF10 -0.8848528 4.794095e-10 -0.5315531 3.288831e-03
138 100293704 <NA> 4.1652044 2.637579e-05 3.4557775 2.613152e-02
139 9612 NCOR2 -0.7511655 1.501871e-05 -0.5527530 1.783153e-03
140 100507206 LINC00943 6.2760701 1.017513e-02 7.3674812 9.033309e-04
141 219287 AMER2 2.3065591 5.995112e-03 2.6089411 3.298906e-03
142 2963 GTF2F2 -0.5957940 1.811377e-02 -0.8237930 4.213769e-03
143 1948 EFNB2 0.7194535 8.451578e-05 0.9336074 2.764288e-05
144 1284 COL4A2 -0.8395538 1.828348e-08 -0.8404414 1.655976e-08
145 4323 MMP14 -1.5188045 3.672983e-05 -0.9287367 3.491573e-02
146 26020 LRP10 -0.7033103 1.818342e-05 -0.5537884 1.546205e-03
147 5720 PSME1 -0.5382084 1.867755e-03 -0.5772061 4.373057e-03
148 9472 AKAP6 1.7389756 6.851832e-03 1.8568791 4.070817e-02
149 64067 NPAS3 -0.4163694 2.398464e-02 -0.5080488 1.384654e-02
150 6235 RPS29 -0.7327891 3.199018e-11 -0.3803609 6.761011e-03
151 54331 GNG2 1.7462712 1.438339e-06 2.3850711 1.281297e-12
152 10979 FERMT2 0.4541759 1.001712e-02 0.6317041 3.403977e-04
153 652 BMP4 3.4322241 2.358141e-02 3.9413514 1.737453e-02
154 1033 CDKN3 1.4212585 1.657082e-02 1.5963138 1.861682e-02
155 5015 OTX2 -3.8471754 1.568726e-04 -8.7646450 5.096832e-08
156 23002 DAAM1 0.5756500 1.144014e-02 0.6480771 1.098956e-02
157 6252 RTN1 1.2138304 8.122873e-11 2.0253347 5.279550e-25
158 57570 TRMT5 0.5475171 1.728621e-02 0.7457291 1.358172e-03
159 3306 HSPA2 1.0124703 9.266707e-03 1.4676988 3.998716e-04
160 87 ACTN1 -0.4948545 3.613831e-05 -0.6160727 8.692213e-06
161 10577 NPC2 -0.5255423 3.558149e-03 -0.6101615 3.882748e-04
162 83694 RPS6KL1 3.8278595 2.284983e-02 4.3859902 7.594668e-03
163 7043 TGFB3 -1.7661103 7.289027e-05 -1.5817604 8.359027e-04
164 9369 NRXN3 1.2169767 2.413395e-02 2.4311314 2.659111e-08
165 145508 CEP128 7.1384851 3.603826e-04 6.8877576 3.834139e-03
166 10516 FBLN5 1.3856734 8.048815e-09 2.7589733 5.797998e-30
167 90050 FAM181A 0.6710977 1.314873e-02 0.8880721 1.142563e-03
168 83982 IFI27L2 -0.7111727 1.971316e-04 -0.8337053 6.760721e-04
169 283596 SNHG10 -0.9834750 4.264469e-02 -1.3534864 3.597421e-02
170 1778 DYNC1H1 -0.5395047 3.662869e-04 -0.6729980 1.119043e-03
171 7127 TNFAIP2 -1.2376403 1.806957e-07 -1.8796597 9.493629e-13
172 26153 KIF26A 1.5457261 1.467521e-03 2.5721679 2.241496e-12
173 7057 THBS1 1.1410705 4.491190e-02 2.3285348 1.071863e-07
174 388115 CCDC9B -0.9614209 1.019494e-02 -0.9618937 3.587554e-02
175 825 CAPN3 -2.4966123 7.173925e-03 -2.6874120 1.027918e-02
176 10169 SERF2 -0.5003563 5.689073e-04 -0.4259759 1.565503e-02
177 2628 GATM 1.0133473 1.046865e-04 2.1557044 4.223549e-22
178 9768 PCLAF 1.0640307 3.781109e-02 1.4935236 6.760721e-04
179 6176 RPLP1 -0.6093418 3.166160e-07 -0.4406568 7.279779e-05
180 5315 PKM -0.9369895 3.555936e-13 -1.1770603 2.540533e-20
181 3073 HEXA -0.6652942 5.121717e-04 -0.9594898 7.852808e-06
182 80381 CD276 -1.1989176 1.371933e-11 -0.7033776 6.626380e-04
183 4016 LOXL1 -1.1530879 1.557437e-11 -2.2150069 1.393321e-19
184 57611 ISLR2 -1.5523167 1.047715e-04 -1.4906241 1.194952e-03
185 1512 CTSH -0.9493288 6.993697e-03 -2.4987702 7.651342e-08
186 9055 PRC1 0.8509816 1.334434e-02 1.1448669 4.181127e-04
187 6627 SNRPA1 0.9370428 1.515239e-02 0.9710838 1.530019e-02
188 146330 FBXL16 1.2609745 6.780349e-03 1.1989549 3.072066e-02
189 1186 CLCN7 -0.9821757 1.938327e-04 -0.9597895 2.136651e-03
190 9235 IL32 -0.6723417 2.063015e-02 -1.5711091 1.688161e-07
191 84662 GLIS2 -0.8156735 2.638713e-03 -0.7750988 2.533532e-02
192 2903 GRIN2A 2.2116964 1.102525e-03 2.3194333 2.811444e-03
193 8303 SNN 0.4548868 4.175567e-02 0.5396832 3.066528e-02
194 51704 GPRC5B -0.4588105 1.525190e-02 -0.5810413 4.486396e-03
195 388228 SBK1 0.6975649 3.087182e-02 1.3383912 1.252054e-05
196 4502 MT2A -0.8592943 5.139522e-13 -0.7215908 6.547722e-07
197 6236 RRAD -1.2863347 6.482764e-08 -1.6024059 3.329564e-10
198 8824 CES2 -0.6507556 2.915376e-04 -0.6322111 3.907452e-03
199 5699 PSMB10 -0.9345479 1.674125e-03 -0.8847187 1.605926e-02
200 6560 SLC12A4 -1.2011669 1.168969e-04 -1.1745485 2.658996e-05
201 794 CALB2 1.4654634 1.245084e-14 2.7505470 3.771214e-49
202 497190 CLEC18B -1.7376001 1.636610e-05 -2.3390890 1.813716e-09
203 57687 VAT1L 1.0555940 3.343727e-10 1.3598243 2.662264e-15
204 55839 CENPN 1.9185524 4.318828e-04 1.9128202 2.755764e-03
205 80790 CMIP 0.6352677 1.065092e-02 1.0207875 4.669550e-05
206 23199 GSE1 0.6591221 2.943672e-03 0.9168692 7.672086e-04
207 8140 SLC7A5 -1.0733358 3.036524e-21 -2.2277424 1.646229e-54
208 10381 TUBB3 0.5102387 1.912244e-03 1.0074982 5.187304e-11
209 79007 DBNDD1 -1.2036565 5.626879e-04 -1.3212792 2.580847e-04
210 5187 PER1 0.7742327 2.911848e-02 1.7299740 3.482076e-10
211 4628 MYH10 0.4704751 1.176551e-02 0.4854399 3.784166e-02
212 9423 NTN1 -0.8038572 1.879074e-05 -0.5145178 1.565503e-02
213 5376 PMP22 0.5913604 3.709520e-03 0.5934196 1.042772e-02
214 201161 CENPV 1.2216823 1.590302e-07 2.0786215 1.601383e-23
215 51655 RASD1 0.9291624 3.037219e-03 1.6067946 1.018530e-08
216 146691 TOM1L2 -0.5988929 8.648250e-04 -0.5822012 4.891828e-03
217 8851 CDK5R1 0.7281006 2.902766e-02 1.4368297 1.688161e-07
218 124842 TMEM132E -1.1128654 5.631934e-05 -0.9711416 4.506951e-03
219 7153 TOP2A 0.6603221 1.020516e-02 1.2133368 1.522068e-08
220 60681 FKBP10 -0.4693712 1.111818e-02 -0.8194118 1.663741e-05
221 4669 NAGLU -1.1758444 2.360348e-05 -0.7471729 2.925943e-02
222 10493 VAT1 -0.4738040 2.095758e-02 -0.6318269 1.729325e-03
223 8153 RND2 -0.9296475 8.389526e-04 -0.6963927 4.627494e-02
224 10900 RUNDC3A 3.4327764 9.617462e-04 4.2369216 2.005898e-06
225 51629 SLC25A39 -0.7622986 6.773908e-04 -0.6448171 2.065242e-02
226 2896 GRN -0.8639623 1.193912e-09 -1.0190800 4.723064e-10
227 23131 GPATCH8 0.9948362 8.268025e-03 1.0406734 2.169798e-02
228 2535 FZD2 -0.8518293 6.488938e-03 -0.7678688 2.815505e-02
229 2670 GFAP -0.8788076 8.688770e-14 -0.6135068 7.450698e-12
230 4836 NMT1 -1.1632887 2.217884e-14 -0.6361625 5.493712e-04
231 113026 PLCD3 -0.6464323 1.203694e-03 -1.0225654 4.338592e-07
232 10951 CBX1 0.3944676 4.649747e-02 0.5992823 1.055582e-02
233 4804 NGFR -1.8961919 1.638191e-04 -2.0647461 7.699124e-05
234 9902 MRC2 -0.4674889 3.502325e-04 -0.7894924 7.125825e-08
235 10351 ABCA8 7.1845356 2.807586e-04 6.5117911 2.349140e-02
236 2232 FDXR -1.0998184 2.512139e-04 -0.8961472 9.375928e-03
237 3021 H3-3B 0.4481503 1.388105e-02 0.5709138 1.727819e-03
238 91107 TRIM47 -0.5521811 4.337940e-02 -1.0892316 3.109138e-04
239 6427 SRSF2 0.6570711 8.949572e-04 0.7749439 8.856202e-04
240 57690 TNRC6C 0.9318899 1.811934e-03 1.2590142 3.536824e-05
241 7077 TIMP2 -0.5451779 1.909010e-05 -0.5143953 4.559581e-03
242 3959 LGALS3BP -1.0807887 1.016548e-10 -0.9944939 1.156218e-09
243 2548 GAA -1.1008862 3.334932e-04 -1.1046775 2.438171e-03
244 57674 RNF213 -0.8289028 1.166942e-03 -0.8022541 1.096290e-02
245 284184 NDUFAF8 -0.7803367 4.361201e-07 -0.3851560 2.844284e-02
246 124565 SLC38A10 -0.9196737 1.967783e-06 -0.7500777 9.981604e-03
247 5034 P4HB -0.4811759 6.537472e-04 -0.4358903 1.725798e-02
248 23136 EPB41L3 0.8043025 1.809110e-02 1.2410497 4.155530e-05
249 361 AQP4 1.9832329 1.458331e-21 2.4813295 4.518687e-20
250 8715 NOL4 2.5687040 4.710820e-05 2.3750310 3.748654e-03
251 56853 CELF4 2.8308665 1.337532e-02 3.6788376 1.542505e-04
252 5596 MAPK4 2.1620912 1.378009e-05 3.1944381 4.168558e-15
253 51320 MEX3C 0.7216292 1.368868e-03 0.7288804 1.327124e-02
254 6925 TCF4 0.5224426 8.286071e-03 0.6992144 2.433102e-03
255 51046 ST8SIA3 1.7236224 5.223639e-03 2.0225337 1.231036e-03
256 115701 ALPK2 -0.3642425 1.318485e-02 -0.9000492 4.287961e-06
257 90701 SEC11C 0.6393863 9.080238e-03 0.7131786 1.199206e-02
258 22850 ADNP2 0.9382335 2.980606e-02 1.0730372 3.364371e-02
259 10272 FSTL3 -1.9141863 1.507138e-08 -2.4098435 1.145709e-08
260 2879 GPX4 -0.7943588 1.117022e-04 -0.5399049 1.042772e-02
261 4616 GADD45B 0.6442761 4.617392e-02 0.8482295 1.711765e-02
262 60680 CELF5 1.6397514 9.718171e-05 2.8299477 9.724418e-19
263 10362 HMG20B -0.5705072 2.482387e-02 -0.6321276 4.569560e-02
264 1938 EEF2 -0.7389372 1.785278e-10 -0.6395235 1.845226e-06
265 80700 UBXN6 -1.3315384 3.883207e-07 -0.5436964 4.416257e-02
266 25873 RPL36 -1.1881207 3.462815e-23 -0.4791971 1.583682e-03
267 3383 ICAM1 -0.5382711 3.466183e-02 -0.9588536 7.172713e-03
268 7297 TYK2 -0.8352119 3.919247e-03 -0.8395900 1.042772e-02
269 1995 ELAVL3 1.6008247 2.454648e-14 2.9572941 4.249882e-64
270 51398 WDR83OS -0.6621618 7.344926e-06 -0.4834366 1.179665e-02
271 4784 NFIX -0.5776782 1.388105e-02 -0.6800133 1.051906e-02
272 9592 IER2 0.7907178 1.128910e-03 0.8540200 7.832869e-03
273 23025 UNC13A 1.9799400 3.834358e-02 2.9789190 4.894679e-04
274 54858 PGPEP1 -1.1812771 4.796700e-11 -1.1865815 1.170981e-08
275 1463 NCAN 0.5462462 4.655457e-04 1.3779361 2.286572e-20
276 57616 TSHZ3 2.2062795 8.996888e-04 2.7318589 2.240349e-05
277 1054 CEBPG 2.4944429 5.594227e-11 1.9096803 2.498586e-04
278 9710 GARRE1 -0.6462264 1.658445e-02 -0.8139986 7.634639e-03
279 826 CAPNS1 -0.9150501 1.243570e-09 -0.6479115 2.443150e-04
280 7538 ZFP36 2.1059243 6.969675e-05 2.1236973 1.194859e-03
281 645 BLVRB -1.0333420 1.228879e-03 -1.0952873 5.846189e-04
282 602 BCL3 -0.6322777 1.674787e-02 -1.4886064 9.936794e-09
283 27113 BBC3 -1.3305512 2.285361e-14 -1.1325714 3.830730e-09
284 8605 PLA2G4C -1.3554795 3.958831e-02 -1.6706418 4.784355e-02
285 10945 KDELR1 -0.3167301 4.213416e-02 -0.5231799 5.564150e-03
286 6141 RPL18 -0.3924189 1.383462e-03 -0.3167985 2.360646e-02
287 51171 HSD17B14 -2.4382627 8.095397e-06 -3.0486255 9.880131e-07
288 4924 NUCB1 -0.8808132 6.334853e-08 -0.5112236 9.375928e-03
289 581 BAX -1.4924872 2.618953e-13 -0.7377792 1.239803e-05
290 2512 FTL -0.4942241 2.712866e-03 -0.6672808 2.482689e-08
291 2217 FCGRT -0.8782572 7.301084e-04 -1.0374652 2.021297e-03
292 6237 RRAS -1.0491362 2.177220e-05 -1.7450213 2.652380e-06
293 2109 ETFB -0.9079624 1.045505e-05 -0.5234932 3.723936e-02
294 101060391 <NA> 2.4792585 1.390130e-02 3.7223004 2.194727e-06
295 6664 SOX11 0.6401704 2.796227e-05 1.8647460 3.791295e-32
296 57498 KIDINS220 0.6170120 3.915208e-03 0.6849782 5.593621e-03
297 6382 SDC1 -1.3289872 4.026500e-03 -1.7592725 3.278351e-03
298 5500 PPP1CB -0.3473838 2.187066e-02 -0.9982070 1.894902e-10
299 6432 SRSF7 0.9228610 3.402040e-04 0.9014630 1.710025e-03
300 9378 NRXN1 3.2548950 1.117022e-04 4.3738759 2.987959e-10
301 55704 CCDC88A 1.3880390 1.399888e-20 1.1390501 1.632985e-08
302 51255 RNF181 -0.6008622 1.419132e-03 -0.7086328 1.174351e-03
303 51652 CHMP3 -1.0607526 2.116479e-05 -0.8821282 1.791381e-03
304 400986 ANKRD36C 1.0527632 9.243063e-04 0.9333155 2.028577e-02
305 57730 ANKRD36B 1.0015210 2.195924e-04 0.8999155 1.711765e-02
306 9486 CHST10 -0.8993879 2.266197e-07 -0.9206192 7.125825e-08
307 84417 ECRG4 1.0248005 2.217257e-03 1.1840566 8.394612e-04
308 3625 INHBB -1.1471836 1.615238e-05 -1.6018996 1.446473e-04
309 2863 GPR39 -1.6489253 2.178554e-03 -2.2435522 2.325506e-04
310 4249 MGAT5 0.7442919 9.599486e-03 0.9842356 9.405533e-04
311 3800 KIF5C 0.4649209 1.525190e-02 0.7388030 2.977731e-04
312 114793 FMNL2 -0.7898825 1.705881e-05 -0.8536632 4.788315e-05
313 56475 RPRM 2.6483233 2.564757e-02 3.7565992 3.335295e-04
314 90 ACVR1 1.2507968 1.781456e-02 1.6051601 2.597708e-03
315 220988 HNRNPA3 0.6482484 4.706088e-04 0.6659135 1.978731e-03
316 91404 SESTD1 1.6691029 9.604536e-05 1.5514432 7.832869e-03
317 51602 NOP58 1.2115301 2.267611e-06 1.0686516 8.546641e-04
318 8828 NRP2 -0.5226133 3.204906e-03 -0.9573256 1.599583e-08
319 8609 KLF7 1.0025878 1.033785e-06 1.0033664 3.078591e-04
320 2066 ERBB4 2.2485676 3.804598e-04 2.9170132 8.735657e-08
321 79582 SPAG16 0.6284534 3.959769e-03 0.7885585 1.342963e-03
322 6168 RPL37A -0.9303554 3.042872e-18 -0.3655451 8.019089e-03
323 3488 IGFBP5 -1.4806287 1.305233e-12 -2.6276423 1.373751e-26
324 5798 PTPRN 6.8817517 8.593309e-04 7.8030319 3.307515e-05
325 7857 SCG2 2.6113161 6.033672e-10 3.7349230 1.200548e-25
326 5270 SERPINE2 -0.9926334 1.123782e-07 -1.4256796 1.036254e-07
327 151473 SLC16A14 6.8979289 8.572895e-04 7.2070774 1.558555e-03
328 5757 PTMA 0.7519147 1.387138e-04 0.7887292 8.998932e-05
329 3635 INPP5D -2.2579553 5.424084e-03 -2.8167141 7.994280e-04
330 2637 GBX2 -2.7009859 4.364859e-03 -6.2925642 1.466669e-04
331 2817 GPC1 -0.7017691 1.769125e-03 -0.6905752 3.943678e-03
332 57506 MAVS -0.9250269 4.940842e-05 -0.8589591 2.809631e-04
333 6238 RRBP1 -0.8686983 2.637579e-05 -0.9009048 3.285356e-04
334 2937 GSS -0.7443337 9.556314e-03 -1.0706544 1.119043e-03
335 4826 NNAT 3.1680251 7.373864e-17 5.3451431 5.973672e-20
336 84969 TOX2 -1.1459953 7.190124e-04 -1.3343938 1.161155e-03
337 51604 PIGT -0.4991703 3.478910e-02 -0.6856040 8.774860e-03
338 11065 UBE2C 1.3363669 3.139684e-03 1.8034193 2.587335e-04
339 5740 PTGIS -2.7612814 1.938607e-02 -7.0602674 1.050621e-04
340 655 BMP7 0.4510380 4.264469e-02 0.5359877 4.798275e-02
341 100462983 MTRNR2L3 -0.9819699 4.010125e-03 -1.1648154 2.074899e-03
342 56937 PMEPA1 -0.5230080 3.898929e-03 -0.9284119 2.782505e-07
343 3911 LAMA5 -0.6052878 9.972372e-03 -1.2493616 1.833221e-08
344 6227 RPS21 -0.5594318 2.996471e-06 -0.4096432 2.941618e-03
345 79144 PPDPF -0.6670751 4.327228e-06 -0.3941644 2.634767e-02
346 140578 CHODL 0.8632802 1.949047e-02 1.2636075 1.565239e-03
347 10600 USP16 1.2901756 4.427731e-09 0.8025704 1.271791e-02
348 56245 C21orf62 0.8932085 5.886905e-04 1.0088722 1.367254e-04
349 1827 RCAN1 1.6169413 1.891122e-32 1.8123293 2.115084e-33
350 54102 CLIC6 -2.1222556 1.120768e-02 -2.6977421 1.231036e-03
351 5211 PFKL -0.9994257 6.710778e-06 -0.7925855 2.421756e-03
352 80781 COL18A1 -0.6429705 4.185984e-03 -1.4012123 9.118276e-11
353 1291 COL6A1 -0.7334164 4.184865e-07 -0.6569213 4.337822e-05
354 1292 COL6A2 -1.3165699 8.946508e-04 -1.4529057 1.727819e-03
355 114044 MCM3AP-AS1 4.8433420 2.054490e-02 5.1556030 2.798496e-02
356 9993 DGCR2 -0.7393578 1.071439e-06 -0.4582500 2.169798e-02
357 652968 CASTOR1 -0.8958563 1.023060e-02 -1.0533783 7.355072e-03
358 6948 TCN2 -1.3719700 4.070087e-04 -1.2418938 6.518635e-03
359 8542 APOL1 -2.4345061 3.621806e-03 -2.7056263 4.636375e-04
360 4627 MYH9 -0.3688866 5.764094e-03 -0.6803203 6.846859e-05
361 29775 CARD10 -0.9401422 7.212982e-03 -1.2970562 2.494423e-02
362 27350 APOBEC3C -1.9626154 5.457109e-06 -2.5872219 1.910522e-06
363 6122 RPL3 -0.3961649 5.118954e-04 -0.4590053 4.374010e-04
364 468 ATF4 0.7341126 2.253227e-04 0.6362050 1.221214e-03
365 55964 SEPTIN3 0.6535855 1.198266e-04 0.9633868 2.581520e-08
366 112885 PHF21B 1.5549008 4.011849e-03 2.2449725 6.620180e-06
367 55007 FAM118A -1.0297339 1.087497e-02 -1.3182219 4.685516e-02
368 7508 XPC -0.7104169 8.209563e-03 -0.7223291 3.410553e-02
369 9975 NR1D2 -1.2910971 5.627615e-03 -1.1626472 3.409973e-02
370 7068 THRB 5.0150341 2.159653e-03 6.1027810 1.198285e-05
371 116135 LRRC3B 1.1558432 8.772857e-04 1.5001648 1.478949e-04
372 27303 RBMS3 -2.0662574 7.993136e-03 -1.9706185 4.640489e-02
373 91392 ZNF502 2.1416244 4.197156e-02 2.7973671 3.055488e-03
374 10675 CSPG5 0.8446787 1.466453e-02 1.5989512 8.664247e-07
375 2771 GNAI2 -0.4543327 1.350202e-04 -0.4763113 4.896779e-05
376 7869 SEMA3B -1.6475938 4.752961e-08 -1.4351780 2.091144e-04
377 440957 SMIM4 -0.5866022 6.939397e-04 -0.5266405 3.709768e-03
378 7086 TKT -0.8925533 9.251601e-12 -1.1219402 9.581386e-15
379 2317 FLNB -0.6756728 4.312458e-02 -1.1666722 3.634648e-03
380 11170 FAM107A 1.1104834 2.350922e-03 2.4518419 3.100573e-17
381 8618 CADPS 6.9087162 1.192958e-03 8.3390362 6.297620e-07
382 132204 SYNPR 3.1769801 5.309386e-13 3.1533266 5.348598e-12
383 9223 MAGI1 0.4978709 1.427483e-02 0.7145249 1.937158e-02
384 56650 CLDND1 1.0804913 1.666380e-04 0.8996633 3.238264e-02
385 961 CD47 0.8328868 1.615238e-05 0.8365725 9.139134e-05
386 55081 IFT57 1.0963751 3.073686e-05 1.0014533 7.706317e-03
387 29114 TAGLN3 0.8305021 3.258099e-03 1.9453827 4.168558e-15
388 79858 NEK11 1.3419328 1.850060e-02 1.4663334 3.003395e-02
389 7018 TF -3.3138202 4.725046e-35 -2.6201244 4.161319e-27
390 5217 PFN2 0.3523718 4.478066e-02 0.7112425 1.747127e-05
391 5806 PTX3 1.8589852 1.367837e-11 1.4264436 2.064758e-03
392 6657 SOX2 -0.5130097 8.535894e-05 -0.4483034 3.190897e-02
393 84002 B3GNT5 3.1771060 9.492418e-05 2.9853757 3.924664e-03
394 6480 ST6GAL1 5.2895695 3.498778e-02 6.4189671 2.152638e-03
395 53834 FGFRL1 -0.8519278 5.105178e-04 -1.0492044 1.422827e-04
396 10417 SPON2 -2.3091363 1.712572e-08 -4.2748445 1.430988e-13
397 285489 DOK7 -1.6758347 2.855885e-02 -3.3022258 2.754133e-03
398 4043 LRPAP1 -0.4685760 2.442153e-03 -0.6023743 4.082433e-04
399 27065 NSG1 1.4644396 1.376922e-03 2.5808441 2.241496e-12
400 57537 SORCS2 -0.7871403 4.529303e-02 -1.6736831 1.240910e-04
401 8532 CPZ -0.7725561 6.978706e-04 -1.1278406 7.929514e-06
402 79730 NSUN7 3.2018938 7.750809e-03 3.7341067 1.069240e-03
403 9662 CEP135 2.3818682 3.676748e-03 2.3832982 1.944258e-02
404 57482 CRACD 1.4143169 1.376922e-03 1.3546269 3.076361e-02
405 84525 HOPX 0.8006811 1.307252e-05 0.4927356 4.720265e-02
406 2926 GRSF1 0.6256449 3.584136e-02 0.7764957 1.534185e-02
407 9508 ADAMTS3 1.2581974 3.040421e-02 1.8256163 4.890966e-03
408 9987 HNRNPDL 0.7667476 8.001259e-10 0.8633976 2.346389e-11
409 8404 SPARCL1 3.4518746 1.810240e-02 5.7065746 1.283212e-09
410 6164 RPL34 -0.2726410 1.335382e-02 -0.4045876 1.809947e-03
411 5188 GATB -0.6342599 1.003106e-02 -1.0566535 4.213824e-04
412 84057 MND1 6.3689687 7.985926e-04 5.7875478 2.157803e-02
413 3148 HMGB2 1.5575692 3.399080e-07 1.8107787 1.042069e-06
414 79682 CENPU 1.7572873 2.248609e-02 2.0161924 1.497425e-02
415 134121 C5orf49 1.2891972 2.152868e-02 1.6567350 1.318342e-03
416 100505738 MIR4458HG -0.6736466 4.960015e-04 -0.9792733 3.598272e-07
417 56172 ANKH 1.9013191 4.952983e-02 2.3061208 2.360646e-02
418 3157 HMGCS1 0.8251998 1.308333e-04 1.2064166 3.304990e-09
419 5144 PDE4D 3.0265041 1.233264e-02 3.9901682 7.651789e-05
420 3796 KIF2A 0.7211634 2.075211e-02 0.9707886 7.986975e-03
421 4131 MAP1B 0.6501884 1.009581e-11 0.4458657 4.760952e-03
422 8507 ENC1 0.7194827 7.482311e-03 1.0252607 2.551228e-05
423 3156 HMGCR 1.1990674 1.438339e-06 1.0978114 2.295227e-04
424 7060 THBS4 1.6714202 3.470063e-03 1.9361751 1.656621e-03
425 7025 NR2F1 -0.5239516 5.301621e-03 -0.7285153 1.757675e-02
426 1946 EFNA5 1.8063297 3.037219e-03 2.3255291 1.182043e-04
427 324 APC 0.8329661 1.828265e-03 0.9625218 7.604579e-04
428 4001 LMNB1 0.9899708 4.791469e-02 1.3387424 5.396264e-03
429 441108 <NA> -0.7398561 3.240773e-02 -1.0710016 3.001819e-02
430 23338 JADE2 -2.1443131 1.923788e-03 -1.8656289 1.839096e-02
431 9879 DDX46 0.4407792 3.343420e-02 0.7063848 2.434675e-03
432 9547 CXCL14 -1.1500571 5.710320e-15 -2.3721417 1.503380e-08
433 6695 SPOCK1 -1.4233259 3.516658e-06 -1.7590654 2.210035e-08
434 51306 FAM13B 1.3277345 2.263273e-02 1.4046261 4.749983e-02
435 1958 EGR1 2.2672437 3.856617e-04 2.4980545 1.028471e-03
436 1839 HBEGF 0.7151459 2.979990e-02 1.3006599 1.528098e-04
437 2246 FGF1 0.6235654 2.880638e-02 1.0320918 1.282450e-03
438 11346 SYNPO -1.1973194 6.008782e-09 -1.1898142 1.055427e-05
439 10318 TNIP1 -1.1302281 4.470897e-08 -1.1870840 2.381835e-05
440 345630 FBLL1 1.0250079 3.462894e-02 1.4713806 1.983706e-03
441 64901 RANBP17 7.3286999 9.675490e-05 6.1899962 4.070817e-02
442 1843 DUSP1 2.3389517 1.273281e-03 2.5638656 2.811444e-03
443 8992 ATP6V0E1 -0.3377393 3.655155e-02 -0.6097685 4.551850e-04
444 51617 NSG2 1.8040947 9.266707e-03 4.1218627 7.584436e-19
445 10814 CPLX2 1.6274692 9.392387e-04 3.5137687 2.236009e-29
446 11282 MGAT4B -0.7511967 1.707978e-05 -0.7693371 1.528098e-04
447 54898 ELOVL2 0.7979453 5.119268e-03 1.1282748 2.853354e-04
448 84062 DTNBP1 -1.1375751 3.084289e-05 -0.7077513 4.978160e-02
449 3400 ID4 1.3234321 2.454648e-14 0.9597098 5.245563e-06
450 10590 SCGN 2.4339359 3.784284e-26 3.1883757 3.436960e-49
451 100129195 ZSCAN16-AS1 -0.9824939 2.547771e-03 -0.9671895 2.925943e-02
452 3105 HLA-A -0.7630050 2.536753e-02 -1.1830565 7.298879e-04
453 3106 HLA-B -0.6920167 4.768717e-04 -0.9450936 4.991740e-06
454 6892 TAPBP -1.0925660 2.103301e-09 -0.8757743 3.598272e-07
455 221491 SMIM29 -0.9126123 3.265619e-04 -0.7966399 2.300862e-02
456 6428 SRSF3 0.5907227 1.544322e-03 0.5518872 2.655440e-02
457 1026 CDKN1A -0.7352948 5.673328e-09 -1.0205134 3.102091e-12
458 221477 C6orf89 -0.4290711 3.655155e-02 -0.5997872 4.176082e-03
459 4188 MDFI -1.6946787 4.396140e-07 -1.3989779 7.206897e-03
460 6722 SRF 0.5035098 3.518799e-02 0.6710501 1.775345e-02
461 5429 POLH -0.8482561 5.598261e-03 -0.8333806 3.838590e-02
462 441151 TMEM151B 2.4813514 2.136663e-02 3.8733208 1.307363e-06
463 10492 SYNCRIP 0.6547845 9.106134e-05 0.6851036 3.292267e-04
464 10690 FUT9 1.6228542 4.295188e-02 2.0538863 1.353922e-02
465 100133941 CD24 0.7826658 3.352518e-04 1.0520556 2.959499e-06
466 2534 FYN 0.5754398 2.001797e-03 0.5222057 2.420192e-02
467 2697 GJA1 0.7860628 1.505593e-09 0.6923602 6.978354e-08
468 1490 CCN2 1.7669666 1.270529e-38 1.5007085 1.529090e-03
469 51696 HECA 2.2305072 7.866600e-03 2.0715526 4.712701e-02
470 168090 C6orf118 6.5459019 9.453808e-06 7.1323223 6.583826e-07
471 51660 MPC1 1.3861836 7.373864e-17 0.9075743 1.915919e-03
472 392617 ELFN1 -0.7633663 1.535848e-02 -0.9114103 1.418670e-02
473 29886 SNX8 -0.8103420 1.203993e-03 -0.7810089 7.506111e-03
474 2115 ETV1 1.6971077 2.762389e-28 1.7069240 2.317050e-16
475 8701 DNAH11 6.6153200 3.513273e-03 7.3569130 9.558770e-04
476 4852 NPY 1.3154757 1.750895e-06 2.3498901 2.565768e-28
477 79017 GGCT 1.2098831 3.979532e-06 1.0129868 1.365792e-02
478 64759 TNS3 -0.6654305 1.769450e-02 -1.2746450 1.839349e-05
479 7378 UPP1 -0.6755395 6.065297e-03 -0.9264188 1.027423e-03
480 64409 GALNT17 -1.0238278 7.558802e-06 -1.9099800 7.365375e-03
481 11257 <NA> -0.8753233 3.386687e-02 -1.2223439 1.452713e-02
482 5244 ABCB4 -0.7938959 1.121641e-02 -1.7461164 2.042689e-08
483 1750 DLX6 1.6967820 9.798325e-04 2.9795474 1.176582e-16
484 1749 DLX5 2.1487545 2.964895e-05 3.6472396 1.961173e-22
485 25798 BRI3 -0.5605655 3.962710e-05 -0.4798202 1.983706e-03
486 245812 CNPY4 -0.6789715 3.975321e-02 -1.0137840 1.705933e-02
487 7205 TRIP6 -0.5516844 1.432870e-03 -0.4696307 2.079177e-02
488 5054 SERPINE1 2.8513450 2.722428e-19 2.6376578 1.989264e-14
489 2861 GPR37 1.9616350 2.142435e-04 2.6412375 1.626127e-09
490 9296 ATP6V1F -0.6208547 4.185984e-03 -0.4916495 3.190897e-02
491 4232 MEST 1.1090253 7.948981e-06 1.8407917 6.282989e-16
492 26958 COPG2 2.0371199 2.372957e-08 2.3938799 4.276165e-09
493 155066 ATP6V0E2 -0.9295884 9.408256e-10 -0.5163034 8.652071e-04
494 54480 CHPF2 -0.6725414 3.776809e-02 -0.8143411 2.613487e-02
495 349136 WDR86 -1.1356266 2.235311e-09 -1.1158594 1.933368e-04
496 79660 PPP1R3B 1.4609627 1.402176e-03 1.8644080 4.269170e-05
497 10395 DLC1 1.8760039 1.172214e-16 1.7331700 2.605612e-11
498 4747 NEFL -1.9411917 8.790184e-07 -2.8226773 6.525063e-10
499 1808 DPYSL2 0.8030127 2.557717e-10 0.9879686 4.168558e-15
500 1191 CLU -0.3444037 1.107307e-02 -0.6555587 8.068107e-07
501 80223 RAB11FIP1 -1.7160464 1.990906e-05 -2.8548878 3.690210e-05
502 286183 NKAIN3 -0.9005197 2.312742e-06 -1.0762978 3.536824e-05
503 5569 PKIA 1.2397113 4.103217e-03 1.2258164 2.598470e-02
504 11075 STMN2 5.0289645 3.982030e-03 6.2206112 3.882748e-04
505 862 RUNX1T1 2.3617343 8.043349e-06 3.3261306 7.110959e-14
506 79870 BAALC 0.5247210 1.942507e-04 0.4522719 1.094306e-03
507 23237 ARC 2.3259726 4.013752e-02 2.5449994 4.898564e-02
508 4062 LY6H 2.3379504 8.390999e-04 3.5422444 1.865305e-10
509 1936 EEF1D -1.1027922 3.167177e-05 -0.6419713 1.207905e-02
510 6132 RPL8 -0.6916893 2.510775e-12 -0.5291093 8.363642e-06
511 6595 SMARCA2 -0.4913439 2.490699e-02 -0.6977376 2.043116e-03
512 11168 PSIP1 1.0403464 5.601800e-10 0.5164943 3.233658e-02
513 92949 ADAMTSL1 -1.5663012 1.242368e-02 -2.3008362 6.016343e-03
514 10210 TOPORS 0.8707731 2.075211e-02 0.9013817 4.070817e-02
515 2683 B4GALT1 -0.7018422 3.693915e-03 -1.5182056 1.759899e-05
516 7094 TLN1 -0.5504165 1.272024e-02 -0.6671978 2.021297e-03
517 389765 <NA> 5.4260040 2.076460e-02 5.9222574 1.505603e-02
518 8013 NR4A3 1.7014179 5.244930e-03 1.7403012 1.588166e-02
519 5998 RGS3 -0.4113436 4.968924e-02 -0.7189488 2.225797e-03
520 3371 TNC 1.4982035 2.413395e-02 2.0004229 2.211218e-03
521 5069 PAPPA -2.2700106 9.962078e-03 -3.7710131 3.117838e-03
522 1620 BRINP1 2.2574012 1.286098e-07 3.0310203 1.402464e-15
523 6136 RPL12 -0.3890133 4.301363e-03 -0.5832159 1.061134e-05
524 64855 NIBAN2 -0.5600645 3.062325e-02 -0.7750492 9.711585e-03
525 10044 SH2D3C 2.6970967 2.599771e-02 3.4350539 1.777636e-03
526 114789 SLC25A25 0.9656242 6.642499e-04 1.0071855 1.250459e-03
527 5524 PTPA -0.6425695 3.513537e-03 -0.7954230 1.068443e-03
528 100506119 LINC01503 -1.1175070 1.441011e-02 -1.7491243 5.914940e-04
529 1289 COL5A1 -1.2018514 4.170684e-03 -2.0049030 7.289041e-07
530 4851 NOTCH1 -0.5344426 3.411729e-02 -0.7501386 8.179276e-03
531 4558 <NA> -0.4298432 1.610129e-03 -0.3656401 1.571732e-02
532 4577 <NA> -1.3295590 1.169416e-14 -1.2308466 1.478113e-10
533 4567 <NA> -1.4148488 9.051147e-27 -0.5195176 2.424442e-03
534 4578 <NA> -0.6747855 3.804763e-02 -1.0236360 1.254247e-03
535 1654 DDX3X 1.1505064 8.327307e-07 0.8401308 5.341800e-03
536 4129 MAOB 0.9150234 1.210106e-04 1.4785002 8.754387e-13
537 57477 SHROOM4 -1.3830074 4.845572e-02 -1.9919367 2.274220e-02
538 170685 NUDT10 1.4147710 2.818500e-02 2.0014879 1.025794e-03
539 28986 MAGEH1 1.2649014 4.241764e-03 1.7982424 3.382692e-05
540 4675 NAP1L3 1.7743412 3.889017e-04 1.9082968 2.021297e-03
541 6173 RPL36A -0.7016506 8.268025e-03 -0.7838113 1.120698e-02
542 11013 TMSB15A 1.5374152 2.912308e-05 2.7447767 3.849016e-24
543 55859 BEX1 0.3872331 1.470021e-02 0.4061960 2.133427e-02
544 51186 TCEAL9 -0.3800982 5.096525e-03 -0.3282262 1.872563e-02
545 5063 PAK3 1.4667139 3.497466e-03 1.5683615 9.420164e-03
546 1641 DCX 1.7466251 1.152596e-19 3.0922910 3.506445e-68
547 3597 IL13RA1 -0.6760142 4.444139e-02 -1.0945962 2.155298e-02
548 6170 RPL39 -0.4799398 5.522891e-05 -0.4551620 2.043707e-03
549 3897 L1CAM 0.9404392 4.222256e-02 1.3354601 4.174608e-03
550 2316 FLNA -0.6232515 9.169656e-07 -0.5112294 1.768206e-03
551 8270 LAGE3 -0.5859079 2.326138e-02 -0.8720007 3.940762e-03
552 2539 G6PD -0.5578078 4.406929e-02 -0.8467634 9.657304e-03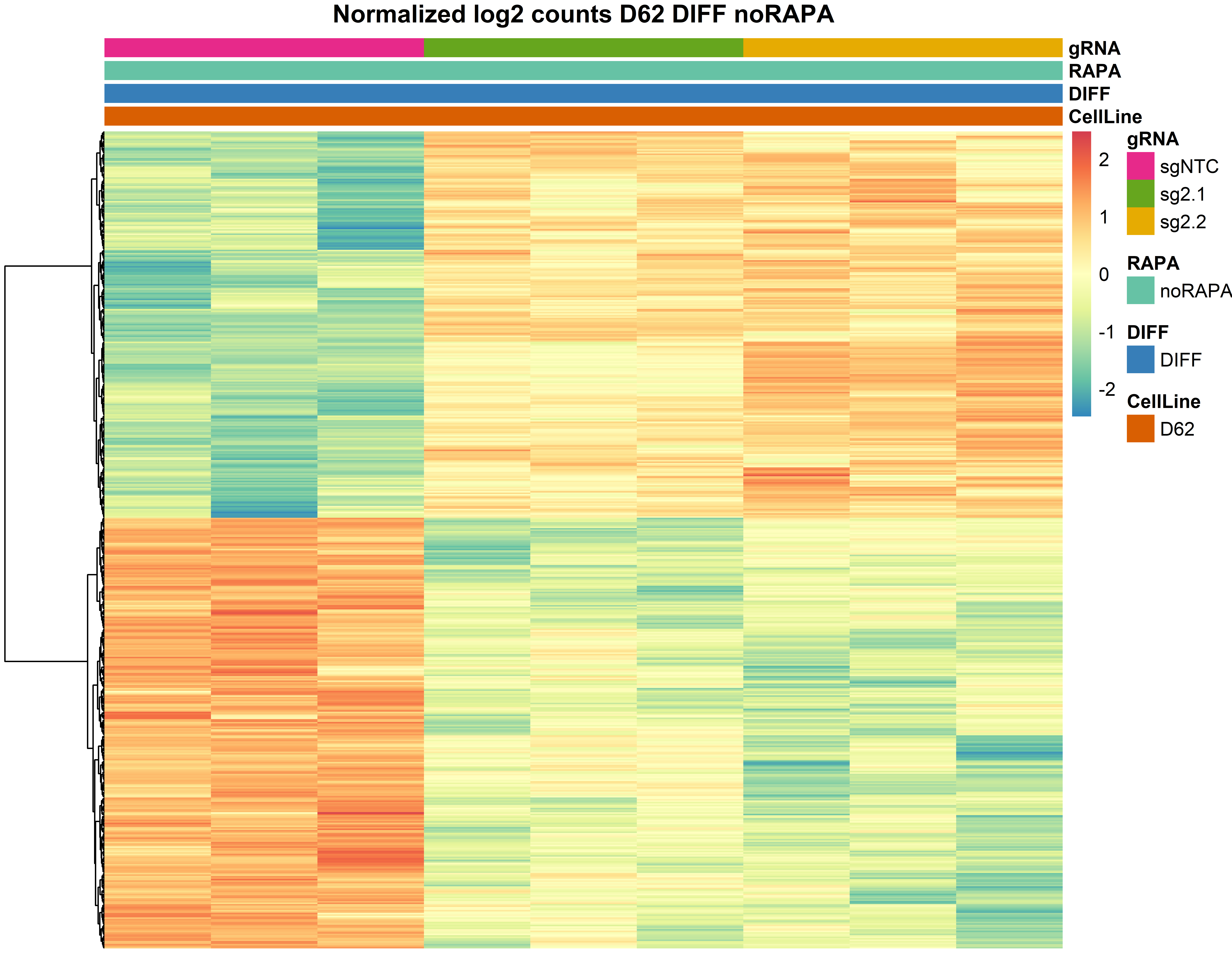
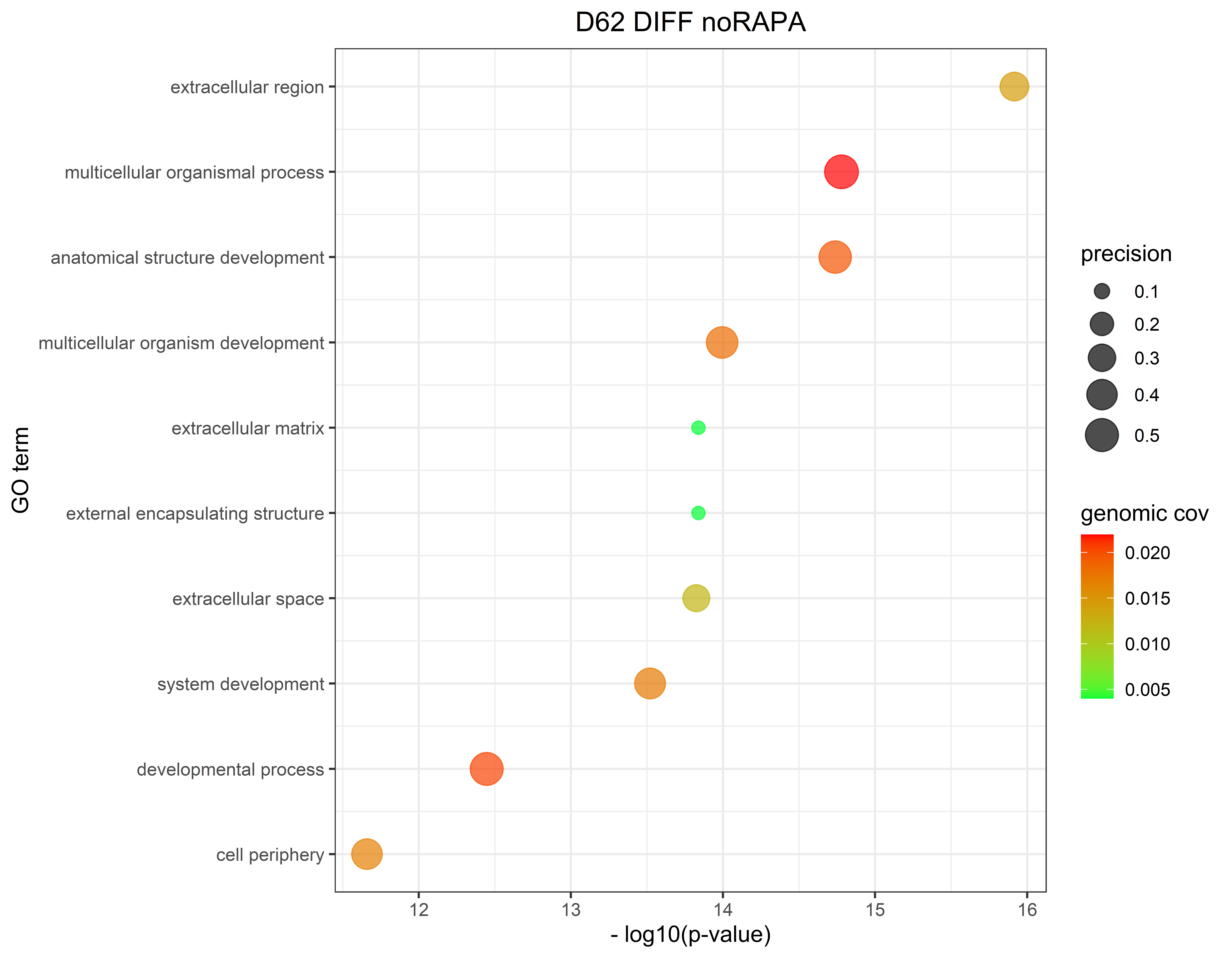
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.9583742 0.59708289 1.468663 1.000000e+00
02_TSC2_KO_Grabole_2016 1.8770766 1.57627048 2.235015 7.533614e-13
03_TSC2_Grabole-2016 1.9358721 1.61429855 2.327220 1.382478e-13
04_TSC2-tuber-Martin2017 3.4242840 1.79363318 6.073134 1.793370e-04
05_TSC2-SEN_SEGA-Martin2017 2.9945499 2.47096771 3.618615 4.881108e-27
06_all_EpilepsyGene 1.1121355 0.63584989 1.823161 6.910190e-01
07_high_conf_EpilepsyGene 1.4137717 0.55347557 3.026237 3.559290e-01
08_ASD_Epilepsy_EpilepsyGene 0.8396849 0.22405456 2.222993 1.000000e+00
09_Epi25_dataset 0.5945811 0.07059024 2.230521 7.744278e-01
10_SFARI_all 1.3814218 0.71934599 2.431774 2.442488e-01
11_SFARI_score4plus 1.4346896 0.67149147 2.726517 2.473882e-01
12_DeRubeis_RDNV 2.3715118 0.91541260 5.181357 3.707211e-02
13_Iossifov_RDNV 1.2888500 0.82533800 1.934140 2.076402e-01
14_FMRP_Darnell 1.3862891 0.97415992 1.926965 5.696057e-02
15_Voineagu_M12 1.0924633 0.60010072 1.845716 6.768491e-01
16_Voineagu_M16 3.2944950 2.28004905 4.654416 1.600300e-09
17_ID_genes_Neel 1.2296736 0.68894616 2.048181 3.933697e-01
18_ID_genes_pinto2014 1.1335593 0.55459533 2.083469 6.212855e-01
19_Fromer_SCZ_FunDN 0.8924430 0.52017976 1.440140 7.276071e-01
20_Cocchi_Pruning 4.0459242 2.04832572 7.391796 6.835179e-05
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -0.04251700 0.24141955 1.000000e+00
02_TSC2_KO_Grabole_2016 0.62971558 0.08910917 1.506723e-11
03_TSC2_Grabole-2016 0.66055790 0.09268233 2.764957e-12
04_TSC2-tuber-Martin2017 1.23089240 0.32992302 3.586739e-03
05_TSC2-SEN_SEGA-Martin2017 1.09679392 0.09805309 9.762216e-26
06_all_EpilepsyGene 0.10628204 0.28524225 1.000000e+00
07_high_conf_EpilepsyGene 0.34626111 0.47846876 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene -0.17472858 0.67404955 1.000000e+00
09_Epi25_dataset -0.51989815 1.08722718 1.000000e+00
10_SFARI_all 0.32311327 0.33292148 1.000000e+00
11_SFARI_score4plus 0.36094855 0.38734822 1.000000e+00
12_DeRubeis_RDNV 0.86352762 0.48566735 7.414422e-01
13_Iossifov_RDNV 0.25375031 0.22740438 1.000000e+00
14_FMRP_Darnell 0.32663043 0.18000522 1.000000e+00
15_Voineagu_M12 0.08843506 0.30565960 1.000000e+00
16_Voineagu_M16 1.19225290 0.18778364 3.200601e-08
17_ID_genes_Neel 0.20674874 0.29558209 1.000000e+00
18_ID_genes_pinto2014 0.12536248 0.36473421 1.000000e+00
19_Fromer_SCZ_FunDN -0.11379263 0.27540215 1.000000e+00
20_Cocchi_Pruning 1.39771001 0.34728943 1.367036e-03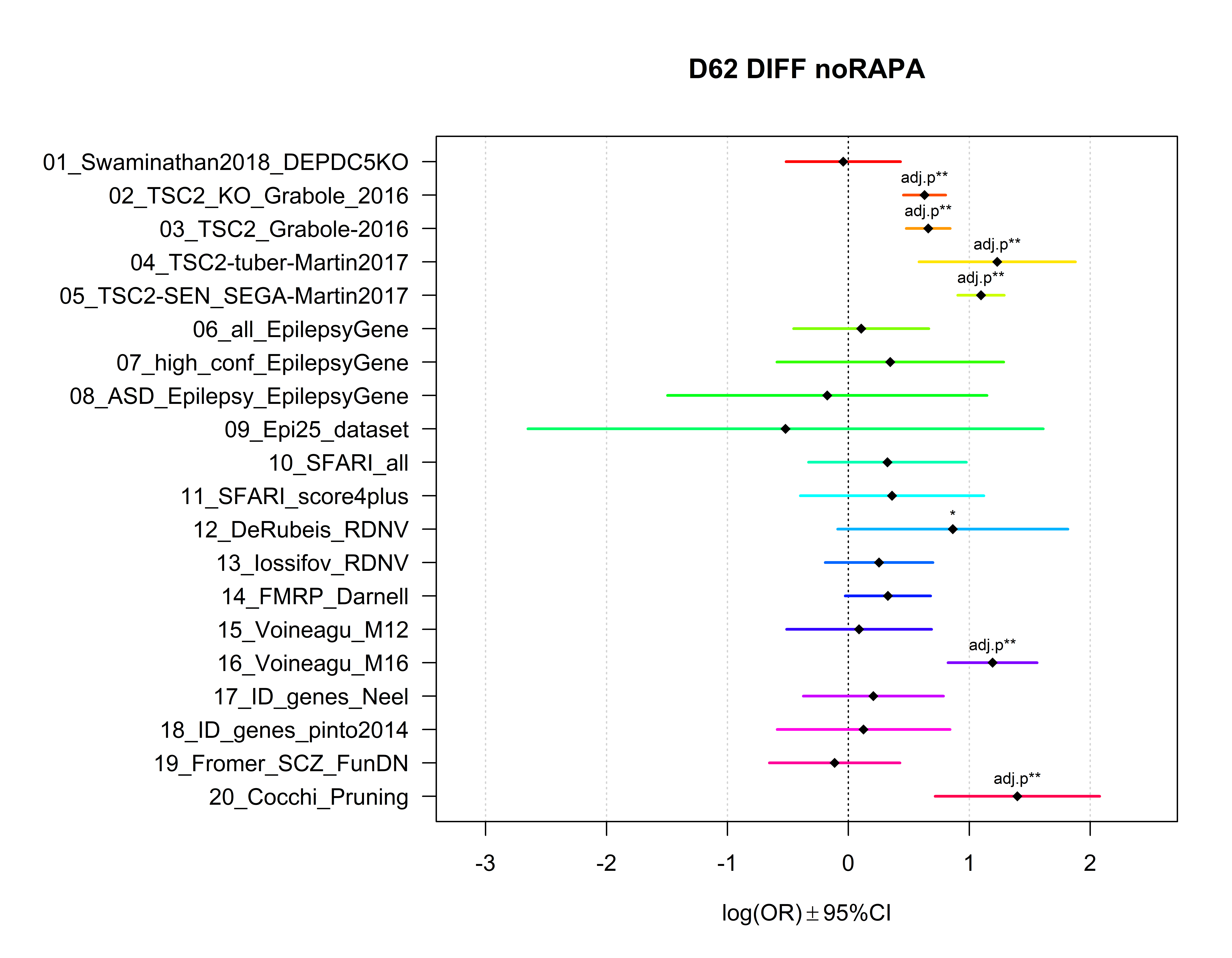
D244
getlinmodoutput(targetline = "D244",targetdiff = "DIFF",targetrapa = "noRAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 4237 MFAP2 -1.1043692 7.672135e-03 -1.2811560 2.995447e-02
2 57134 MAN1C1 0.5990502 7.965530e-03 0.9212493 1.543752e-03
3 6272 SORT1 0.4443084 1.768453e-02 0.5998305 4.210823e-02
4 5738 PTGFRN 0.9467220 3.518917e-04 1.0025640 9.050373e-03
5 6282 S100A11 0.6023700 2.653885e-05 0.5859466 4.404445e-02
6 10485 MIR9-1HG -0.8272271 9.664047e-04 -1.2643130 3.121650e-05
7 10763 NES -0.7704375 2.248345e-08 -0.7919072 4.336417e-04
8 3766 KCNJ10 -1.7938894 1.629161e-06 -1.4794653 4.628547e-03
9 477 ATP1A2 -1.0395969 8.462466e-08 -0.9120463 1.282985e-03
10 8490 RGS5 1.5556701 3.267739e-23 1.2211234 1.551601e-06
11 9019 MPZL1 0.7188379 3.194134e-04 0.6881345 4.555882e-02
12 25874 MPC2 1.1139096 6.543961e-06 1.0688515 1.180660e-03
13 116496 NIBAN1 0.7374131 4.703459e-03 1.0554679 1.282985e-03
14 25802 LMOD1 1.3562336 2.303687e-05 1.8496848 1.084917e-07
15 50486 G0S2 -1.4492217 3.370529e-32 -1.7927431 4.532054e-17
16 5629 PROX1 -1.5178846 1.675095e-02 -2.4556355 7.280961e-03
17 7044 LEFTY2 -1.6863840 1.444934e-19 -2.6934841 1.215175e-28
18 84886 C1orf198 1.2166944 7.326387e-13 0.9990460 1.949588e-05
19 84457 PHYHIPL -0.8575537 1.374903e-03 -0.9576670 2.428566e-02
20 9124 PDLIM1 0.6695783 7.704916e-06 0.9157777 6.404187e-05
21 6319 SCD 1.0653410 1.558967e-14 1.1186607 1.839053e-07
22 2674 GFRA1 -0.6523985 8.183490e-05 -0.8093946 1.203216e-03
23 8519 IFITM1 3.2595909 6.024803e-05 1.8815573 2.234507e-02
24 283298 OLFML1 1.7085130 3.680616e-24 1.8640153 5.795595e-17
25 9537 TP53I11 -1.7242136 7.206134e-08 -1.1792320 1.245941e-02
26 11098 PRSS23 1.0913154 9.842498e-20 0.8635884 5.530015e-06
27 4684 NCAM1 -0.5494706 1.026677e-03 -0.6331499 2.256772e-02
28 4837 NNMT 0.7842202 7.898614e-03 1.1464285 5.758824e-04
29 6876 TAGLN 0.4123892 9.665626e-03 0.7132672 4.250429e-03
30 53826 FXYD6 -1.7162372 1.433325e-34 -1.8416866 1.901516e-15
31 63876 PKNOX2 -1.7482105 1.168670e-04 -1.4982166 5.459458e-04
32 8076 MFAP5 2.3168338 8.651461e-20 2.3448004 2.273254e-14
33 347902 AMIGO2 -1.3842915 2.128170e-04 -2.1109406 1.299218e-06
34 1280 COL2A1 -2.1371974 3.797828e-07 -3.0302746 2.533087e-07
35 3875 KRT18 1.6064905 8.644783e-09 1.3205650 3.053862e-03
36 22877 MLXIP -0.9638468 5.810816e-05 -0.9796499 6.372486e-03
37 114795 TMEM132B -2.6346648 5.304330e-07 -2.3794077 2.797473e-04
38 1910 EDNRB -0.8195503 7.898614e-03 -1.1644740 4.006173e-03
39 1638 DCT -1.4330439 2.648927e-11 -1.7502418 9.909285e-10
40 2621 GAS6 0.5265338 3.923218e-02 0.7026356 1.951712e-02
41 652 BMP4 1.4247389 3.501929e-09 1.1752283 3.095265e-03
42 9369 NRXN3 -1.6621007 6.672684e-07 -2.4424245 1.695466e-04
43 3429 IFI27 4.2542529 6.279178e-06 3.6278061 1.376217e-09
44 12 SERPINA3 -1.6236056 2.521750e-24 -0.8631301 4.289578e-02
45 8788 DLK1 -0.9248486 4.651918e-04 -0.9471378 1.212670e-02
46 567 B2M 1.2077170 3.778239e-11 0.7984159 5.870878e-03
47 79875 THSD4 -2.6664291 6.734017e-08 -2.7334640 1.768468e-05
48 64220 STRA6 -1.0928913 3.537829e-05 -1.2826098 3.114309e-05
49 1512 CTSH 0.5571709 2.552339e-02 0.9196970 3.210155e-02
50 57214 CEMIP 1.8401544 5.074532e-32 1.3640633 2.369816e-10
51 30812 SOX8 -0.9468840 3.369214e-03 -1.2444795 1.103060e-03
52 6236 RRAD 0.8607933 9.125466e-03 1.3528494 4.718324e-04
53 55175 KLHL11 -1.1881721 1.259171e-02 -1.6699572 3.293354e-03
54 284119 CAVIN1 0.6081325 6.279660e-05 0.8970985 3.090207e-03
55 4804 NGFR -2.1767618 3.267739e-23 -2.3203327 2.270876e-09
56 81558 FAM117A 1.1920123 8.271636e-03 1.4175407 5.933378e-03
57 1277 COL1A1 -0.7832460 3.421826e-10 -0.5292790 4.809935e-02
58 114897 C1QTNF1 -2.0950386 5.044701e-12 -2.3070277 7.443622e-11
59 115701 ALPK2 1.0228813 4.356290e-11 1.0947289 4.170082e-07
60 1264 CNN1 0.8030666 7.729680e-05 0.8792943 7.015280e-03
61 10653 SPINT2 1.9402401 1.454866e-05 2.3186908 1.836859e-05
62 81606 LBH 1.5450729 7.986236e-08 1.1938272 1.245941e-02
63 4052 LTBP1 -0.6502580 3.452794e-02 -0.8744678 2.459597e-02
64 7039 TGFA -1.5478130 9.355259e-16 -1.8785682 5.052243e-14
65 2995 GYPC -2.4294636 1.874496e-11 -1.9199833 7.112116e-04
66 2487 FRZB -2.7856065 1.455250e-06 -4.1134175 3.895502e-09
67 23671 TMEFF2 -1.4201643 3.113925e-02 -2.1181155 1.599333e-02
68 8324 FZD7 -1.1339644 4.538585e-03 -1.7225282 3.199580e-04
69 2335 FN1 1.3432977 1.275635e-12 0.9941101 4.294325e-04
70 3485 IGFBP2 0.5823047 8.244139e-07 0.7080885 1.695466e-04
71 80303 EFHD1 -2.2240001 1.869882e-06 -1.6178733 1.377763e-03
72 10123 ARL4C -0.8157205 2.068082e-08 -0.6598818 1.004240e-02
73 3397 ID1 1.0968681 3.778573e-09 0.9615010 1.377763e-03
74 4826 NNAT -1.7863625 3.938824e-09 -1.0454326 2.778496e-02
75 7052 TGM2 1.0996047 1.172199e-04 1.1008156 7.015280e-03
76 56937 PMEPA1 -1.1739691 1.378185e-19 -1.1140883 6.627336e-09
77 1917 EEF1A2 -1.5125821 8.827770e-04 -1.6940719 3.053862e-03
78 6453 ITSN1 1.0065733 4.303701e-05 0.9960276 8.619861e-03
79 5121 PCP4 0.4446898 4.951707e-02 0.9254292 8.667160e-05
80 80781 COL18A1 -1.0190535 1.418955e-07 -1.0103524 6.341399e-05
81 7078 TIMP3 -0.6399207 2.209381e-07 -1.0523639 3.895502e-09
82 2847 MCHR1 -1.7383604 2.719626e-06 -1.7412994 9.279880e-04
83 2192 FBLN1 -1.6448627 1.282796e-08 -1.3138823 4.588621e-03
84 10752 CHL1 -0.6451113 4.538585e-03 -1.3138271 5.459458e-04
85 2199 FBLN2 -1.4094832 3.184372e-04 -1.7087813 1.467409e-03
86 7869 SEMA3B -1.3403981 3.060962e-05 -1.2301226 4.896891e-03
87 1295 COL8A1 -2.5537145 3.437208e-60 -2.5283334 5.898885e-25
88 100506621 <NA> 2.5230669 1.416933e-09 2.2736690 1.346570e-05
89 151887 CCDC80 0.9438908 9.829811e-11 1.2905848 3.240083e-10
90 84561 SLC12A8 -1.1198579 4.852200e-02 -1.6935275 2.118427e-02
91 7018 TF -1.7913566 9.804860e-08 -2.1881714 5.243374e-07
92 5947 RBP1 -1.0715058 1.418681e-15 -0.9284459 1.055017e-04
93 4071 TM4SF1 1.3518166 5.022271e-05 1.1842675 1.873053e-02
94 5010 CLDN11 1.8993934 1.201573e-29 1.8494500 3.978462e-13
95 1427 CRYGS 0.9755418 2.259854e-06 1.3402048 3.659429e-06
96 27065 NSG1 -1.3129250 1.061877e-02 -1.6624600 2.991588e-02
97 54360 CYTL1 -2.6337714 7.090266e-11 -2.1502290 1.855011e-05
98 55351 STK32B -1.5441350 2.153720e-03 -1.4961851 4.689396e-02
99 57537 SORCS2 -1.1432855 4.392572e-07 -1.3209447 1.875633e-06
100 8532 CPZ 0.7750727 1.514441e-03 1.2363550 1.434080e-06
101 9948 WDR1 0.4325543 5.175629e-03 0.7742839 1.377763e-03
102 83888 FGFBP2 -2.1483914 1.344610e-05 -4.2223052 7.674223e-15
103 201780 SLC10A4 0.7930787 1.766552e-03 0.7960404 4.902966e-02
104 65997 RASL11B -1.6303068 9.658775e-12 -2.0074814 1.152095e-05
105 25849 PARM1 -1.1715404 4.787249e-03 -1.9035644 2.824212e-03
106 1363 CPE 1.4185324 4.697313e-13 1.3283372 2.306290e-07
107 11341 SCRG1 0.9125289 8.199232e-15 0.6878051 1.216796e-03
108 100505738 MIR4458HG -0.9544000 2.870460e-05 -0.9170490 1.585138e-02
109 9037 SEMA5A -0.6428037 3.036736e-03 -0.8826364 2.219060e-02
110 1601 DAB2 0.7968959 1.540781e-03 0.7951780 3.414656e-02
111 5144 PDE4D -1.8089323 3.060405e-06 -1.3519833 2.899278e-02
112 645323 LINC00461 -0.8089834 7.729680e-05 -0.8159426 1.771747e-03
113 57556 SEMA6A -0.5829327 3.804354e-03 -0.8454463 3.402279e-02
114 972 CD74 1.0486764 4.082443e-05 1.9810723 8.203233e-12
115 11346 SYNPO -1.0320585 3.175806e-06 -0.8511356 7.015280e-03
116 2878 GPX3 -0.8307352 1.502602e-04 -1.4336932 4.108185e-06
117 23286 WWC1 1.5309623 5.957962e-03 1.9357560 1.936223e-03
118 1843 DUSP1 1.8428072 9.089214e-06 1.7754384 2.802331e-03
119 51617 NSG2 -2.0990520 1.175722e-03 -1.9440049 3.947452e-02
120 3107 HLA-C 0.7460155 1.928539e-03 1.1407880 3.121650e-05
121 3106 HLA-B 0.7080853 1.098278e-02 1.3086746 1.677737e-06
122 2697 GJA1 0.9786886 1.006861e-11 0.7103970 2.802331e-03
123 9053 MAP7 -0.9335155 3.404203e-03 -1.5664711 7.689786e-05
124 6648 SOD2 1.0135486 2.010532e-06 0.8424062 1.599333e-02
125 5154 PDGFA -1.3793439 1.064486e-02 -1.6068819 1.591965e-02
126 10457 GPNMB 1.1582452 3.810058e-08 1.3785261 4.578081e-08
127 358 AQP1 -1.8170423 2.468838e-04 -2.0168612 4.718324e-04
128 5244 ABCB4 -1.3752686 2.416092e-05 -2.2432327 1.296467e-05
129 5243 ABCB1 -1.1713101 3.427240e-05 -1.5823229 2.269751e-06
130 1278 COL1A2 -1.5956895 1.872763e-37 -2.2370695 1.682870e-38
131 1749 DLX5 -1.7044278 3.207583e-06 -2.6514489 9.565218e-07
132 51200 CPA4 3.3228930 6.365409e-05 3.1103410 8.470072e-03
133 28996 HIPK2 -0.8391336 1.550376e-09 -0.8387070 2.797473e-04
134 3084 NRG1 2.5200046 1.733589e-35 2.4862133 5.843487e-25
135 286 ANK1 1.8151747 1.147763e-03 1.7245348 3.953019e-02
136 4147 MATN2 2.1408080 8.995671e-10 1.9035397 1.636967e-04
137 114907 FBXO32 1.1939560 2.430979e-17 1.0565700 2.030369e-06
138 1030 CDKN2B 1.2067611 7.993677e-14 0.7141965 1.737711e-02
139 7169 TPM2 0.5224004 2.201374e-02 0.8922799 1.910044e-04
140 1164 CKS2 1.4049983 2.986016e-02 1.8050482 2.392489e-02
141 23446 SLC44A1 -0.6399794 5.854605e-03 -0.8460054 4.555882e-02
142 3371 TNC 1.4093005 5.957962e-03 1.4983126 4.016110e-02
143 1620 BRINP1 -0.8174476 3.918294e-02 -1.5059106 3.298460e-03
144 100506119 LINC01503 1.1047022 5.760934e-09 1.3491184 7.419941e-07
145 4577 <NA> 0.6482089 5.697536e-04 0.7339766 3.261869e-02
146 4564 <NA> 1.3438030 1.618919e-02 1.5611560 2.675819e-02
147 7102 TSPAN7 -0.7991686 3.697190e-02 -0.9151580 3.414206e-02
148 51186 TCEAL9 0.4947480 4.954149e-03 0.6425542 3.022561e-02
149 5354 PLP1 -1.7840245 1.631697e-04 -2.3895645 3.266449e-05
150 10857 PGRMC1 1.5465723 1.470735e-25 1.0576200 2.522925e-06
151 633 BGN -2.7382586 5.058178e-19 -1.7534538 9.522607e-05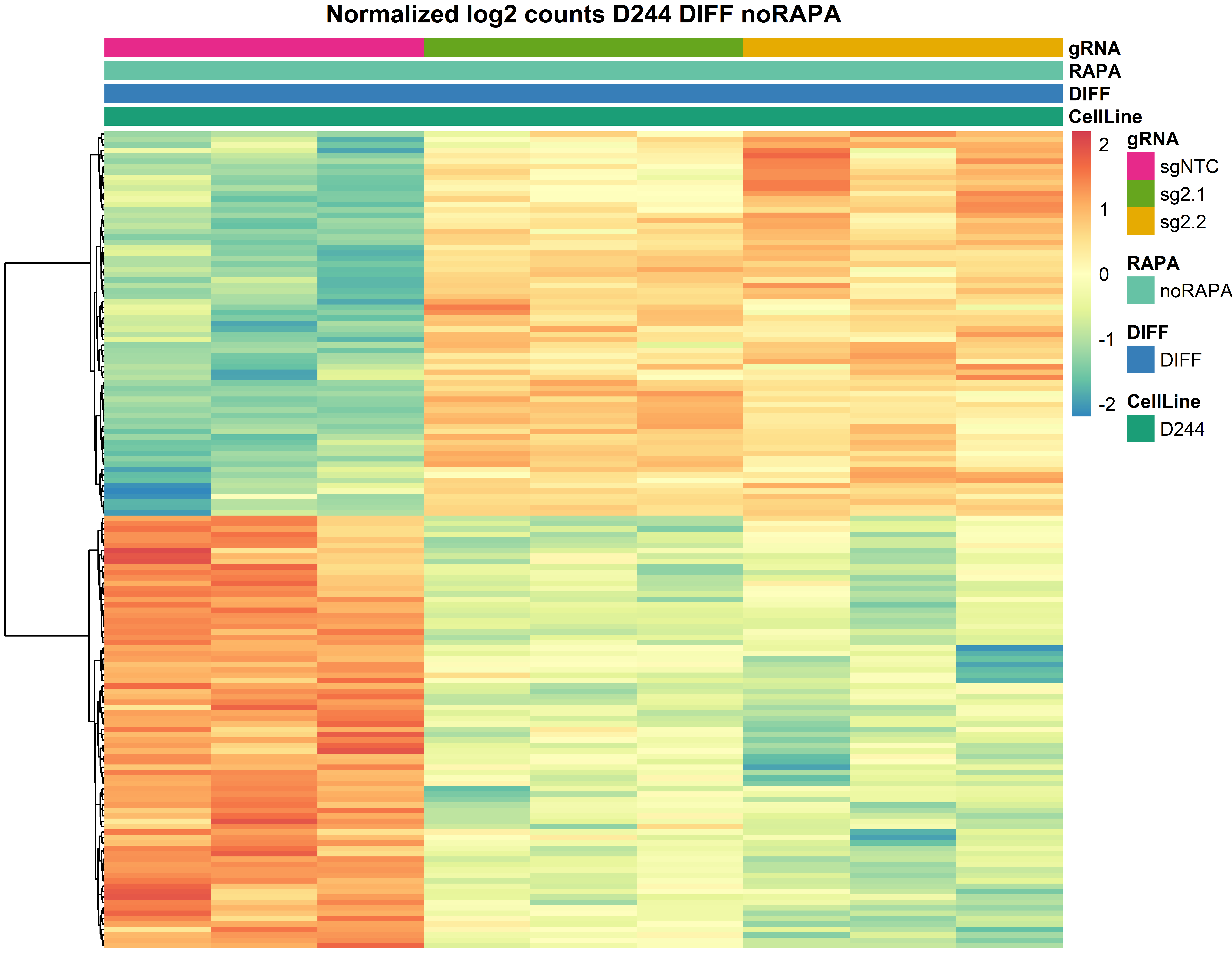
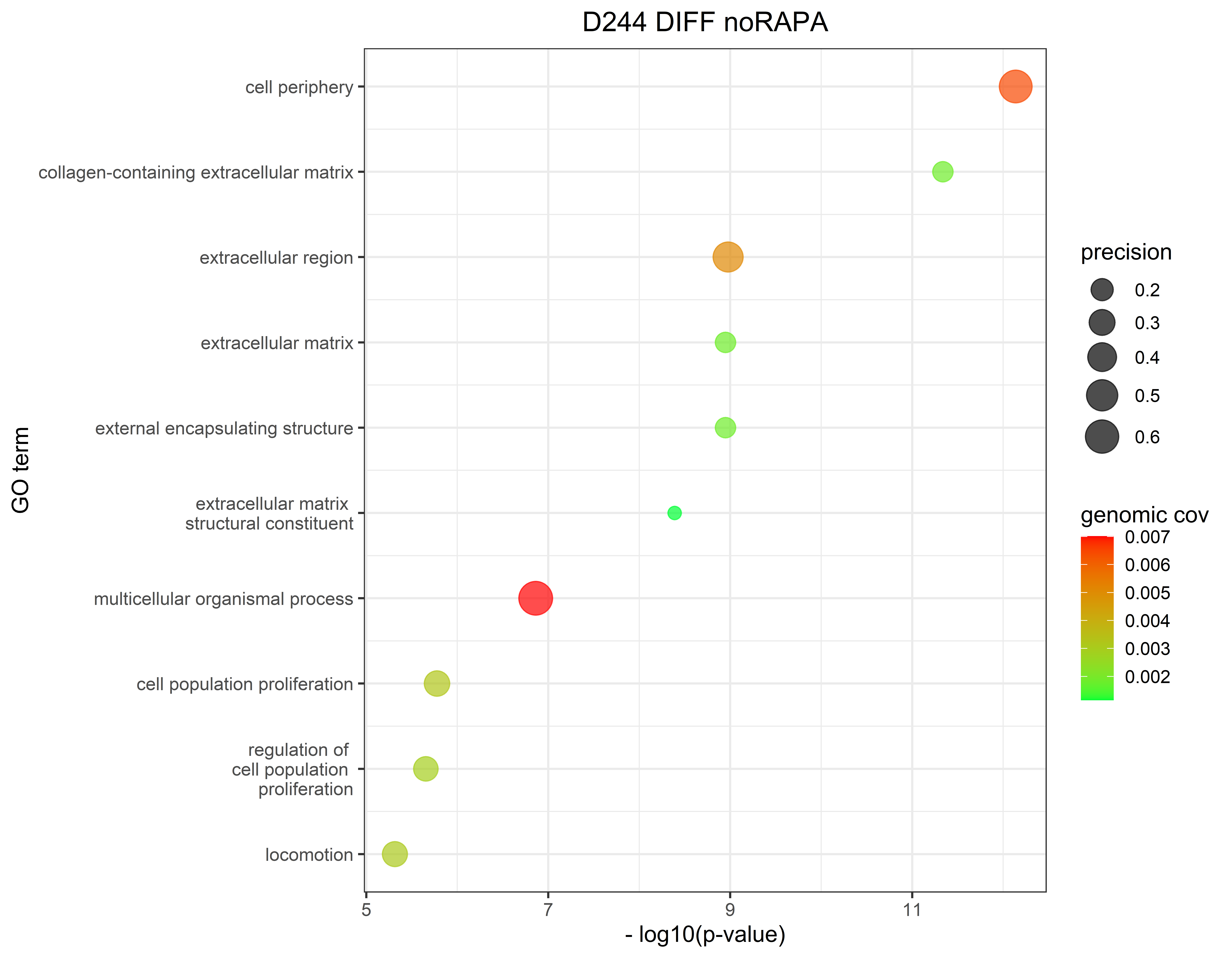
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.4056031 0.62654217 2.763592 3.109960e-01
02_TSC2_KO_Grabole_2016 1.9200002 1.37412094 2.683343 9.572006e-05
03_TSC2_Grabole-2016 1.7026951 1.20895133 2.417617 1.781611e-03
04_TSC2-tuber-Martin2017 4.2525902 1.33352593 10.463156 8.314656e-03
05_TSC2-SEN_SEGA-Martin2017 3.0531135 2.12290029 4.342801 2.595086e-09
06_all_EpilepsyGene 1.1957650 0.38028788 2.880076 6.162926e-01
07_high_conf_EpilepsyGene 2.2237529 0.44720014 6.782833 1.606467e-01
08_ASD_Epilepsy_EpilepsyGene 0.0000000 0.00000000 2.876627 6.426428e-01
09_Epi25_dataset 1.1097402 0.02757239 6.456619 5.977032e-01
10_SFARI_all 1.5440119 0.41169653 4.091552 3.387752e-01
11_SFARI_score4plus 2.1044631 0.55951964 5.602390 1.317950e-01
12_DeRubeis_RDNV 0.0000000 0.00000000 4.381144 1.000000e+00
13_Iossifov_RDNV 1.0672994 0.38346881 2.396791 8.276203e-01
14_FMRP_Darnell 1.6134753 0.83382001 2.868981 1.055826e-01
15_Voineagu_M12 0.5202004 0.06216787 1.927224 5.942408e-01
16_Voineagu_M16 3.7530167 1.92830724 6.721290 1.141028e-04
17_ID_genes_Neel 0.5450579 0.06511959 2.020046 5.900912e-01
18_ID_genes_pinto2014 1.1255446 0.22785602 3.394610 7.513912e-01
19_Fromer_SCZ_FunDN 1.1016149 0.39573027 2.474213 8.245434e-01
20_Cocchi_Pruning 0.9968485 0.02479799 5.784120 1.000000e+00
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.340466493 0.4122478 1.000000e+00
02_TSC2_KO_Grabole_2016 0.652325309 0.1706689 1.914401e-03
03_TSC2_Grabole-2016 0.532212368 0.1747240 3.563223e-02
04_TSC2-tuber-Martin2017 1.447528257 0.5916846 1.662931e-01
05_TSC2-SEN_SEGA-Martin2017 1.116161892 0.1853973 5.190172e-08
06_all_EpilepsyGene 0.178786188 0.5844964 1.000000e+00
07_high_conf_EpilepsyGene 0.799196280 0.8183395 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene -Inf NaN 1.000000e+00
09_Epi25_dataset 0.104125912 1.8852379 1.000000e+00
10_SFARI_all 0.434384148 0.6744148 1.000000e+00
11_SFARI_score4plus 0.744060353 0.6758862 1.000000e+00
12_DeRubeis_RDNV -Inf NaN 1.000000e+00
13_Iossifov_RDNV 0.065131557 0.5222595 1.000000e+00
14_FMRP_Darnell 0.478390416 0.3368001 1.000000e+00
15_Voineagu_M12 -0.653541225 1.0838652 1.000000e+00
16_Voineagu_M16 1.322559963 0.3397538 2.282057e-03
17_ID_genes_Neel -0.606863178 1.0840136 1.000000e+00
18_ID_genes_pinto2014 0.118266970 0.8149532 1.000000e+00
19_Fromer_SCZ_FunDN 0.096777155 0.5223467 1.000000e+00
20_Cocchi_Pruning -0.003156519 1.8846102 1.000000e+00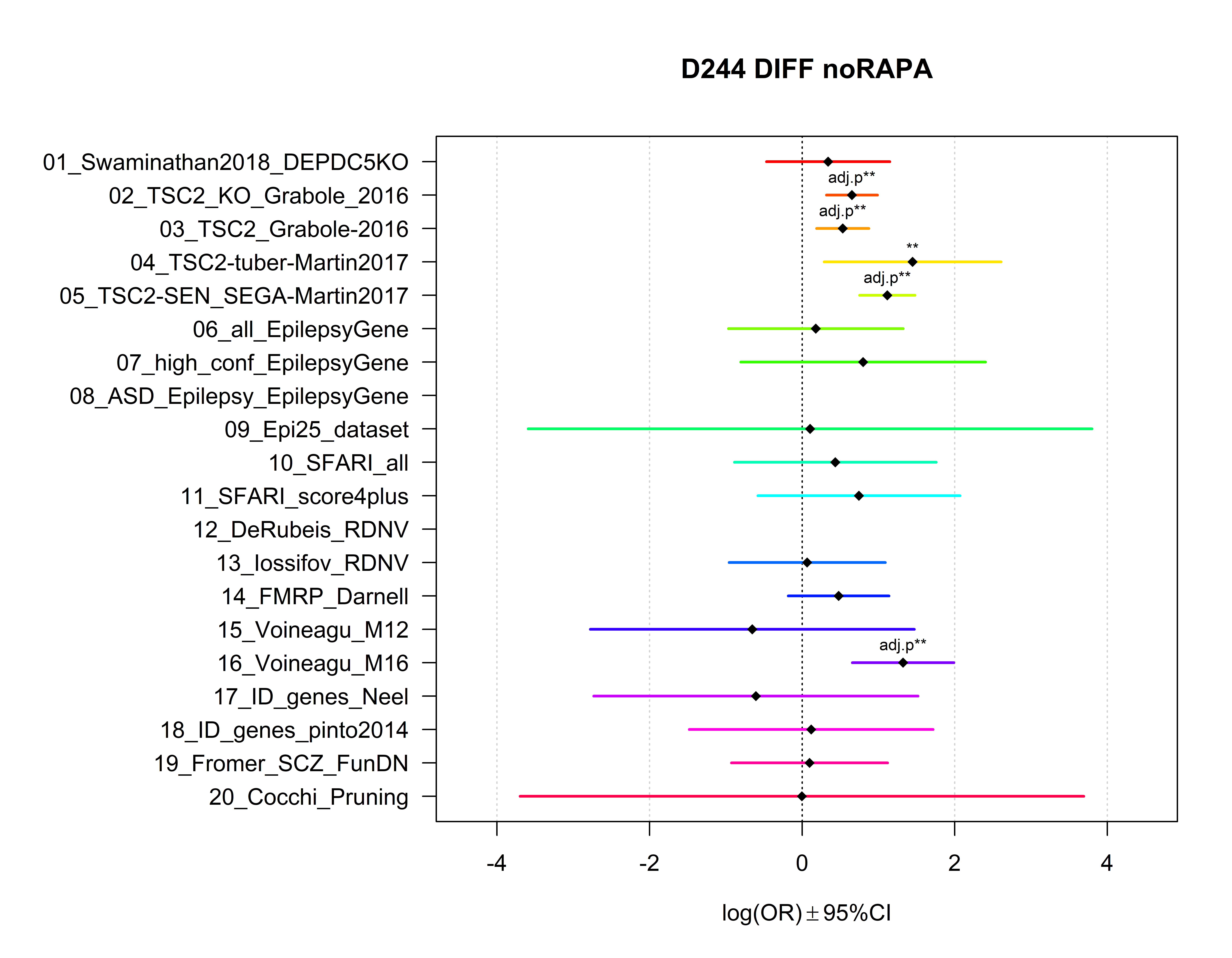
ReN
getlinmodoutput(targetline = "ReN",targetdiff = "DIFF",targetrapa = "noRAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 473 RERE -0.5118111 2.837468e-04 -0.5727503 1.267634e-02
2 56998 CTNNBIP1 -0.9224672 6.378644e-06 -0.6898244 2.600186e-02
3 57190 SELENON 0.7388231 2.242220e-03 0.9922930 3.648510e-03
4 93974 ATP5IF1 0.3750303 5.529278e-03 0.5328467 2.086976e-02
5 9672 SDC3 0.4200234 1.905774e-02 0.7255504 5.656066e-04
6 1296 COL8A2 -2.7649200 3.725687e-21 -1.0785453 6.224653e-03
7 8613 PLPP3 0.7599778 1.011492e-05 0.8241804 7.275014e-03
8 205 AK4 1.6580222 1.649947e-07 1.1798881 3.090197e-02
9 1266 CNN3 0.9730737 1.784179e-08 0.6234705 1.091323e-02
10 4853 NOTCH2 1.0229204 1.107394e-05 0.7966362 3.963774e-02
11 8991 SELENBP1 1.1278639 5.274970e-05 1.5364305 3.403004e-05
12 140576 S100A16 0.7930354 3.473825e-04 1.2748916 7.501518e-06
13 6284 S100A13 0.9651031 9.736536e-06 0.9486517 1.232686e-02
14 57863 CADM3 2.3186406 1.779588e-04 2.6663276 4.723515e-04
15 4921 DDR2 1.0420514 3.473825e-04 1.0152726 2.777144e-02
16 5087 PBX1 -1.0593261 4.907081e-28 -0.9601448 5.162137e-12
17 5396 PRRX1 -2.3101820 1.828671e-07 -1.5755696 3.261094e-03
18 23215 PRRC2C -0.4414583 1.981079e-02 -0.8365786 2.889939e-04
19 9588 PRDX6 0.8015208 2.449107e-13 0.6735934 7.244009e-05
20 93273 LEMD1 2.2784598 9.693402e-05 2.3308687 2.147669e-02
21 5629 PROX1 -1.7582585 2.840679e-10 -1.0251478 2.334237e-02
22 25782 RAB3GAP2 0.7103025 1.015203e-03 0.7609884 1.556170e-02
23 3142 HLX 7.3913395 4.742238e-05 7.9051559 2.131872e-04
24 375057 STUM -2.2381342 3.124981e-18 -1.5076900 7.360385e-04
25 183 AGT 1.4135071 7.255331e-03 1.8485512 3.789527e-03
26 5209 PFKFB3 1.2101232 4.928500e-06 0.9912340 1.573111e-02
27 7431 VIM 0.4836743 3.835456e-10 0.3223265 3.380892e-02
28 783 CACNB2 2.0236730 2.357647e-03 2.1401503 3.174840e-02
29 22852 ANKRD26 -0.6982284 3.945250e-02 -1.0848550 2.376198e-02
30 23522 KAT6B -1.1820263 7.110670e-08 -1.0116676 1.091323e-02
31 9124 PDLIM1 0.7066029 4.700066e-02 1.0994174 1.838462e-02
32 6319 SCD 0.5841642 2.539450e-07 0.5131367 3.524913e-02
33 57698 SHTN1 -2.2194186 2.081371e-15 -1.7826326 3.088823e-04
34 55366 LGR4 1.5362244 6.859844e-03 1.9709420 9.843683e-03
35 4192 MDK -1.5148613 3.682824e-06 -1.1201928 3.019314e-02
36 2785 GNG3 5.3650269 2.831130e-02 6.0132720 3.918578e-02
37 4718 NDUFC2 0.6489253 1.037384e-03 0.6959760 4.090037e-02
38 143686 SESN3 1.0484072 6.807509e-10 1.0902203 5.753519e-06
39 4684 NCAM1 -1.4661757 2.548571e-24 -0.9214890 1.984333e-06
40 4837 NNMT 1.7157895 7.167236e-04 1.5991146 2.853278e-02
41 6876 TAGLN -1.7986553 1.499050e-88 -0.4903330 9.931289e-03
42 220323 OAF 0.9612079 3.540122e-02 1.4388907 8.546162e-03
43 7132 TNFRSF1A 0.5667423 2.095919e-02 1.1179910 9.320462e-05
44 55885 LMO3 -1.9086464 1.936685e-05 -1.8756365 2.358171e-02
45 117177 RAB3IP -1.5583580 2.161168e-09 -0.8447957 3.317232e-02
46 1466 CSRP2 1.2824167 1.883552e-11 0.9813446 3.789527e-03
47 160335 TMTC2 0.9851093 1.735493e-10 0.7707206 7.161402e-03
48 283431 GAS2L3 -1.3843243 1.897922e-04 -1.3911010 6.647306e-03
49 51559 NT5DC3 -1.1395309 4.985118e-02 -1.5142001 3.111249e-02
50 84915 FAM222A -1.0703628 1.112791e-03 -1.1000597 2.376198e-02
51 54997 TESC 2.5229672 3.326932e-08 2.5729805 1.492068e-05
52 10186 LHFPL6 -0.9005337 1.673936e-10 -0.7186683 2.029774e-03
53 115207 KCTD12 -1.6539535 1.905866e-13 -1.2472397 9.558302e-03
54 10253 SPRY2 1.5622436 3.437285e-04 1.9049387 1.352112e-03
55 8660 IRS2 0.4989190 2.572734e-05 0.8149257 4.824424e-07
56 64067 NPAS3 0.8316952 2.113359e-03 0.9998032 8.258381e-03
57 3958 LGALS3 0.8192631 1.949086e-09 0.8413821 2.301044e-05
58 23002 DAAM1 -0.5669808 4.089591e-02 -0.8735874 1.988219e-02
59 6252 RTN1 0.7675965 1.783937e-03 1.0132506 1.990635e-03
60 123096 SLC25A29 -0.8758338 3.887469e-05 -0.8259506 4.038827e-02
61 113146 AHNAK2 -2.3453203 3.470782e-11 -1.9232024 1.388677e-02
62 4212 MEIS2 -0.4937972 1.523294e-02 -0.9262050 1.305772e-02
63 10099 TSPAN3 0.5119911 2.591050e-04 0.5529489 1.457608e-02
64 1512 CTSH 1.3023557 1.374873e-19 1.3941737 4.135188e-13
65 4828 NMB 1.1404205 5.594195e-07 1.8295736 1.045468e-15
66 56963 RGMA 0.5208479 6.670103e-05 0.9645057 7.845403e-10
67 80205 CHD9 -0.7636277 4.103199e-03 -0.7078110 4.253471e-02
68 643911 <NA> 1.9003473 1.146616e-08 1.4860851 1.919426e-02
69 221184 CPNE2 -1.5607056 1.524401e-15 -1.1346859 6.430378e-06
70 124491 TMEM170A 1.0445542 6.886734e-04 1.0648140 2.600186e-02
71 11345 GABARAPL2 0.6720920 2.394815e-05 0.6132839 2.309342e-02
72 57687 VAT1L 2.2804660 6.488897e-08 2.4309094 1.086892e-05
73 23199 GSE1 -1.1414934 1.486258e-09 -1.2333610 1.398099e-04
74 10381 TUBB3 -0.4585011 1.850568e-03 -0.6061171 4.102580e-03
75 9955 HS3ST3A1 2.2476161 1.085830e-04 2.3078288 9.843683e-03
76 6347 CCL2 1.0556954 8.383866e-09 1.5714356 4.135188e-13
77 91608 RASL10B 1.4627291 2.359389e-02 1.7165464 3.418162e-02
78 379 ARL4D -1.8159221 1.921646e-14 -1.5481257 1.602870e-08
79 11248 NXPH3 1.8474520 3.270374e-08 1.4162731 1.780658e-02
80 2186 BPTF -1.0095078 3.253859e-07 -1.0020383 2.484974e-04
81 23580 CDC42EP4 -1.2767945 3.626884e-11 -0.8325367 2.517962e-02
82 3021 H3-3B -0.6624998 3.241902e-12 -0.4133346 3.174840e-02
83 2302 FOXJ1 -1.1561841 2.045266e-09 -1.3793825 6.313723e-07
84 7077 TIMP2 -0.8077256 4.431683e-19 -0.3788819 2.148951e-02
85 1630 DCC -4.1789687 8.420107e-17 -2.9175898 3.789527e-03
86 23327 NEDD4L -0.5187851 8.363735e-05 -0.4976309 3.174840e-02
87 684 BST2 2.4194952 9.536273e-05 3.1164598 8.611552e-06
88 2014 EMP3 0.8953559 2.597256e-02 1.2974901 2.232260e-02
89 59284 CACNG7 1.7802245 3.679574e-08 1.7985133 2.937358e-05
90 59283 CACNG8 1.6369424 8.089671e-03 2.2351074 5.299233e-04
91 79042 TSEN34 6.6587376 7.209516e-03 7.6916724 5.188673e-04
92 10971 YWHAQ 0.6807554 5.214634e-09 0.5325181 2.358171e-02
93 55133 SRBD1 -0.9173926 2.311308e-02 -1.3623015 3.206683e-02
94 618 BCYRN1 0.8717516 1.901245e-06 1.3821674 6.620114e-09
95 6233 RPS27A 0.3229373 5.493640e-04 0.5275396 3.229242e-02
96 55704 CCDC88A -0.8204642 7.053404e-09 -1.2429623 6.536418e-11
97 53335 BCL11A -3.2405034 6.503446e-18 -2.0257391 9.931289e-03
98 7360 UGP2 1.0474353 1.016636e-02 1.3589884 8.860302e-03
99 72 ACTG2 -3.4080129 5.081336e-20 -1.3283225 3.174840e-02
100 200424 TET3 -1.0569728 4.651943e-02 -2.0848612 4.943602e-02
101 100506421 PANTR1 1.2313685 5.518367e-20 0.6304971 3.090197e-02
102 1622 DBI 0.4411729 1.371119e-04 0.6440859 2.541290e-02
103 56475 RPRM -4.6001121 1.157342e-02 -1.8142592 4.348741e-03
104 5937 RBMS1 -2.2493687 3.019080e-25 -1.3362943 2.341052e-04
105 1745 DLX1 -0.3424517 4.932440e-02 -0.5749909 7.304694e-03
106 165215 FAM171B 1.5448241 6.810704e-17 1.0039986 5.357695e-03
107 7855 FZD5 -2.2991055 1.419217e-04 -1.4463537 4.693355e-03
108 5270 SERPINE2 1.9587881 1.447870e-09 1.5305883 1.826175e-02
109 55502 HES6 1.7230597 2.643337e-38 0.8571782 3.531982e-03
110 1471 CST3 0.9847587 6.810704e-17 1.3108306 1.272181e-16
111 84532 ACSS1 -2.5583536 4.406397e-28 -0.8517395 4.475405e-02
112 128866 CHMP4B 0.6179510 2.898966e-03 0.7581324 1.702666e-02
113 4826 NNAT 0.6753798 1.008659e-06 0.5850898 1.246781e-03
114 23613 ZMYND8 -0.8587809 3.047943e-10 -0.5467463 3.165370e-02
115 1002 CDH4 2.2383921 1.002118e-05 1.9181880 3.151841e-02
116 3785 KCNQ2 -1.5671610 3.913126e-11 -1.5567066 1.229957e-05
117 7074 TIAM1 -1.6456508 8.638312e-07 -1.2257337 1.024657e-02
118 7267 TTC3 -0.4009573 1.301827e-02 -0.5714940 2.770684e-02
119 6285 S100B 0.4361850 3.958528e-03 0.6894344 3.418162e-02
120 23331 TTC28 -1.0914580 1.030457e-05 -1.0705977 8.621707e-04
121 29796 UQCR10 0.3755899 2.747706e-02 0.8975934 2.112227e-03
122 25817 TAFA5 -0.9044653 6.293602e-13 -0.6826780 3.380537e-03
123 23209 MLC1 0.4954270 1.746139e-03 1.0375064 1.001671e-10
124 10752 CHL1 -2.1227676 2.355915e-12 -1.3424921 9.843683e-03
125 79750 ZNF385D -1.4213989 1.596019e-19 -0.9925602 1.112720e-06
126 25907 TMEM158 0.7663621 8.892543e-04 1.0440572 2.484905e-04
127 10675 CSPG5 1.5547282 1.826144e-13 0.9684691 7.181364e-03
128 51246 SHISA5 0.6384714 2.468608e-05 0.9898423 4.570837e-03
129 11170 FAM107A 0.5347181 5.241972e-03 0.8459335 2.821248e-03
130 55079 FEZF2 2.7931010 3.433714e-21 1.7649628 3.084119e-02
131 132204 SYNPR -0.8516845 4.980919e-03 -1.3095218 1.100699e-03
132 23150 FRMD4B -2.1460348 9.954241e-19 -1.2760995 1.703790e-04
133 23024 PDZRN3 -0.6014417 2.158857e-03 -0.6640674 4.371370e-02
134 29114 TAGLN3 -1.5145174 4.800362e-12 -1.1764531 4.821974e-06
135 4638 MYLK -1.5925555 4.154974e-04 -1.4218626 3.100057e-02
136 3693 ITGB5 0.3997963 2.784764e-03 0.5681016 4.846183e-03
137 11343 MGLL 2.0283318 7.323354e-11 1.8218835 9.284187e-05
138 200916 RPL22L1 0.4708538 2.102961e-04 0.6575604 2.179467e-03
139 347689 SOX2-OT -1.0390614 2.900255e-04 -1.3448406 3.732726e-02
140 1974 EIF4A2 -0.5051331 2.784764e-03 -0.6192012 2.836337e-02
141 79572 ATP13A3 1.2381986 1.198998e-02 1.6383107 7.161402e-03
142 677770 SCARNA22 -0.5387333 1.540525e-02 -1.6310283 6.147798e-06
143 23231 SEL1L3 1.2038795 7.231155e-06 1.7878558 9.350406e-09
144 5156 PDGFRA 1.9689443 3.275954e-30 0.9211449 1.457608e-02
145 258 AMBN -3.6043640 1.144179e-10 -1.8675956 1.001671e-10
146 118429 ANTXR2 1.0746021 4.448427e-02 1.5519526 1.091323e-02
147 658 BMPR1B 1.4281673 1.988417e-03 1.5495631 1.457608e-02
148 5530 PPP3CA -0.4987347 2.427693e-02 -0.9842317 1.808248e-03
149 10252 SPRY1 1.4416377 9.053302e-03 2.0074464 7.624287e-04
150 166614 DCLK2 -0.7800887 1.516474e-13 -0.5514906 2.114357e-02
151 2891 GRIA2 -2.2004887 4.832502e-12 -1.0694494 4.719826e-02
152 1363 CPE 0.9929161 2.365879e-03 1.0805984 3.317232e-02
153 9037 SEMA5A -2.6665055 2.967264e-25 -1.1499589 1.078532e-03
154 10299 MARCHF6 -0.7717050 2.540553e-05 -0.5823373 4.572559e-02
155 10923 SUB1 0.6526205 1.541839e-05 0.6116682 3.418162e-02
156 6507 SLC1A3 2.0719378 2.013311e-90 1.2793758 2.431251e-13
157 6414 SELENOP 6.7692914 2.114500e-04 6.1650224 2.584376e-02
158 10769 PLK2 -1.8043835 1.077426e-04 -2.4574249 8.425086e-04
159 10788 IQGAP2 2.6721549 1.370323e-14 1.8300775 1.287786e-02
160 100462981 MTRNR2L12 -0.3435337 2.265463e-02 -0.4969972 4.023352e-02
161 5921 RASA1 -1.6779072 2.295906e-02 -3.0779635 3.019314e-02
162 4163 MCC -3.8365044 4.151600e-15 -1.9041806 2.744752e-03
163 1036 CDO1 1.1866761 6.076264e-06 1.2078375 4.161845e-03
164 57507 ZNF608 -1.3340790 2.232613e-06 -1.3623876 5.937990e-03
165 5307 PITX1 1.6884320 4.836088e-42 1.5022916 3.201550e-10
166 5097 PCDH1 5.5796435 2.755055e-03 6.3389812 3.095204e-03
167 6678 SPARC 0.3579895 5.944844e-04 0.6168653 1.435890e-05
168 55568 GALNT10 -1.9193496 4.431683e-19 -0.9553827 7.181364e-03
169 9750 RIPOR2 -1.7337824 3.508525e-13 -1.5673649 3.351622e-06
170 5754 PTK7 0.7098335 1.792818e-02 1.1872815 5.640193e-04
171 26799 <NA> -0.8880015 2.345857e-02 -1.0266831 4.356326e-02
172 6885 MAP3K7 -0.9445473 1.789151e-05 -0.8711201 4.177692e-02
173 2045 EPHA7 -9.0198174 2.012381e-09 -8.6471063 1.172572e-06
174 3910 LAMA4 1.4638053 1.185379e-05 1.4358070 1.306777e-02
175 7259 TSPYL1 0.9712468 3.991679e-04 1.0974665 4.472515e-03
176 2697 GJA1 -1.6674753 4.020441e-43 -1.0510888 2.784052e-08
177 2173 FABP7 1.2469020 4.198713e-03 0.7216714 4.472515e-03
178 221833 SP8 -1.9615685 1.132884e-16 -1.1588439 6.155514e-04
179 55536 CDCA7L -0.8393205 9.432644e-04 -1.1181122 2.137559e-04
180 5137 PDE1C -1.8102877 9.402767e-05 -1.5885147 1.710431e-02
181 6424 SFRP4 -1.8003394 1.038493e-04 -1.9072638 1.091323e-02
182 3315 HSPB1 0.6929614 2.545328e-08 0.5423033 2.541290e-02
183 6717 SRI 0.7564112 3.830091e-10 0.7037221 1.799961e-04
184 23089 PEG10 -0.7643596 1.273863e-19 -1.1453607 2.231584e-08
185 285987 <NA> -1.4335392 1.123079e-02 -1.7625131 2.597827e-02
186 1750 DLX6 -1.2595335 1.852040e-04 -1.3383312 7.275014e-03
187 1749 DLX5 -1.3065394 1.490941e-10 -1.1039911 1.569167e-05
188 5054 SERPINE1 0.7709491 4.408783e-02 1.3463896 1.255029e-03
189 692084 SNORD13 -0.8359273 1.105342e-02 -1.5020719 3.353433e-02
190 6422 SFRP1 1.3198774 6.856865e-05 1.2920121 4.008169e-02
191 11075 STMN2 -4.2789314 1.317360e-13 -2.1678468 4.038827e-02
192 79870 BAALC 1.5045546 1.133068e-24 1.0464921 5.683475e-07
193 4062 LY6H 0.9079039 2.116643e-02 1.3479402 1.070719e-02
194 4781 NFIB -0.4954804 9.717873e-06 -0.5378088 2.086595e-03
195 4915 NTRK2 1.7149831 1.179490e-17 1.4233780 1.129255e-04
196 1620 BRINP1 -1.7881517 2.753966e-27 -1.2716438 4.444840e-07
197 6418 SET -0.5442739 1.272150e-07 -0.5171314 1.966472e-02
198 4508 MT-ATP6 0.4249397 9.305886e-07 0.4145912 1.071062e-02
199 7114 TMSB4X 0.3710941 2.878931e-02 0.9514107 2.842344e-05
200 7102 TSPAN7 -2.3083989 1.147391e-16 -1.9063349 8.611552e-06
201 442454 <NA> 0.4602658 5.514531e-03 0.6320617 9.931289e-03
202 4435 CITED1 1.3956687 1.145260e-04 1.4024685 1.388677e-02
203 55859 BEX1 -0.5562600 7.400299e-08 -0.4944181 4.493398e-02
204 1641 DCX -0.9825873 1.428037e-16 -0.8743527 2.709261e-07
205 5358 PLS3 -1.4726799 1.578490e-24 -0.8123603 4.874364e-04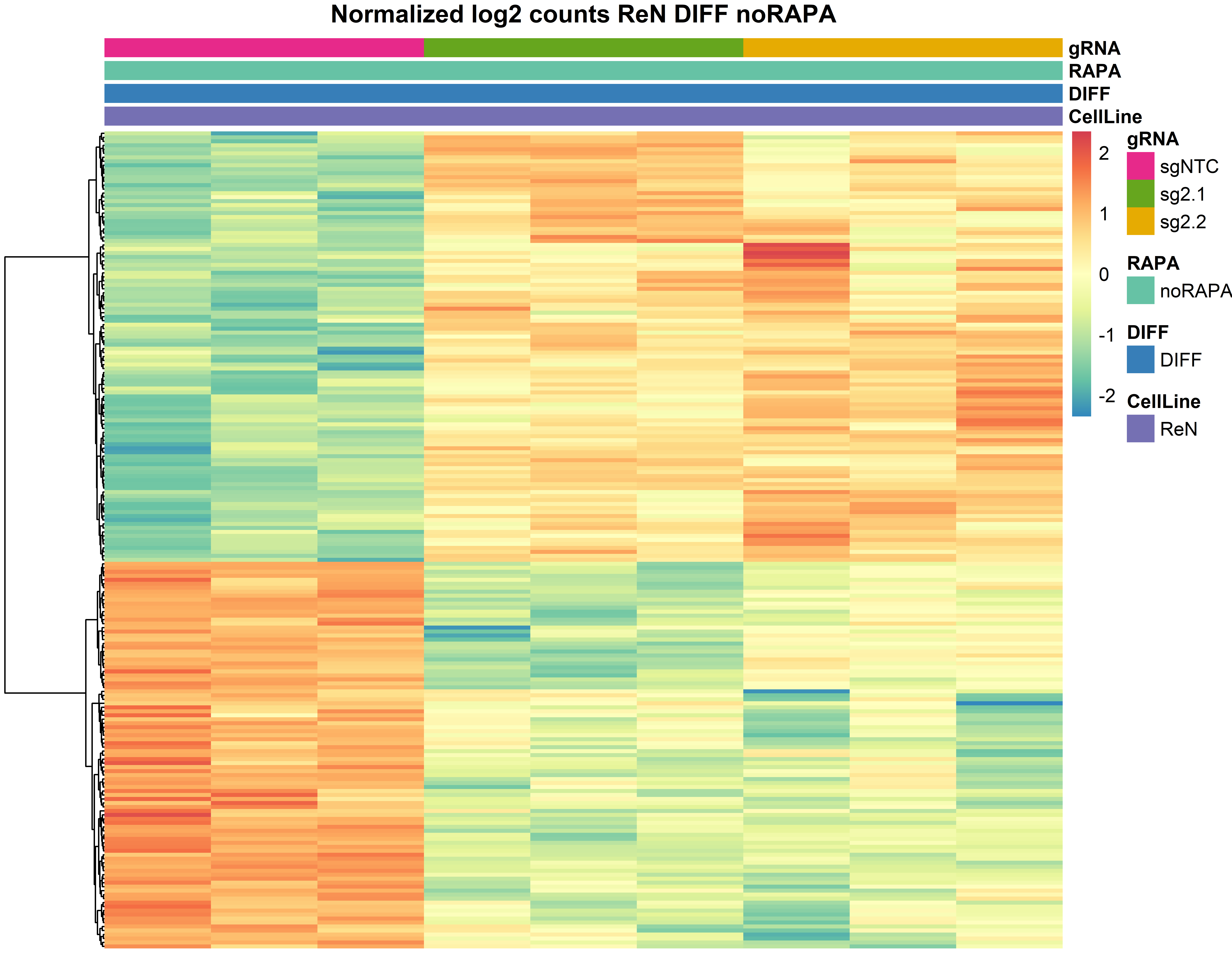
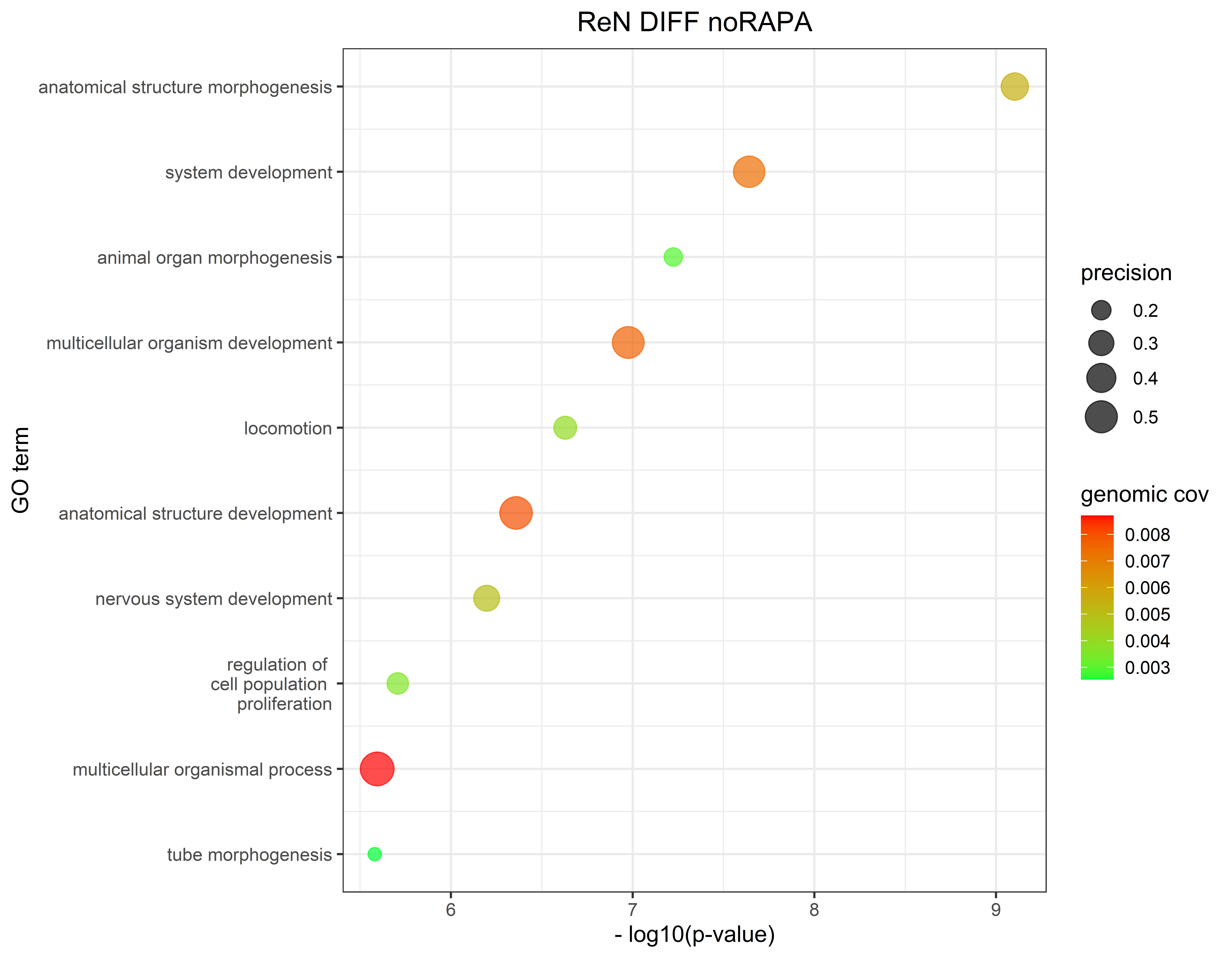
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.1345923 0.53266776 2.146972 6.061621e-01
02_TSC2_KO_Grabole_2016 1.9962929 1.49978294 2.658380 1.234092e-06
03_TSC2_Grabole-2016 1.9866254 1.47318893 2.699791 2.819082e-06
04_TSC2-tuber-Martin2017 3.0920126 0.97339012 7.563659 2.758485e-02
05_TSC2-SEN_SEGA-Martin2017 3.0120559 2.20576576 4.079449 1.050135e-11
06_all_EpilepsyGene 0.8693106 0.27755694 2.081081 1.000000e+00
07_high_conf_EpilepsyGene 2.1927566 0.58223629 5.849217 1.189359e-01
08_ASD_Epilepsy_EpilepsyGene 1.7387099 0.35036131 5.285237 2.566187e-01
09_Epi25_dataset 1.6540845 0.19554612 6.253977 3.474641e-01
10_SFARI_all 2.3356746 0.98254283 4.771893 2.729035e-02
11_SFARI_score4plus 2.3500704 0.84032329 5.313111 5.010660e-02
12_DeRubeis_RDNV 2.6626359 0.53244917 8.202205 1.111669e-01
13_Iossifov_RDNV 1.7647131 0.91614998 3.118571 6.005727e-02
14_FMRP_Darnell 1.4500183 0.80773723 2.433146 1.640624e-01
15_Voineagu_M12 0.3802491 0.04555281 1.402262 2.523069e-01
16_Voineagu_M16 4.9075984 2.95929549 7.786880 1.022341e-08
17_ID_genes_Neel 0.8098984 0.21720370 2.126622 1.000000e+00
18_ID_genes_pinto2014 1.1052273 0.29576687 2.912102 7.861544e-01
19_Fromer_SCZ_FunDN 0.9392830 0.37074278 1.986818 1.000000e+00
20_Cocchi_Pruning 3.0671837 0.80939785 8.262557 4.758133e-02
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.12627335 0.3857810 1.000000e+00
02_TSC2_KO_Grabole_2016 0.69129191 0.1459038 2.468184e-05
03_TSC2_Grabole-2016 0.68643744 0.1525551 5.638163e-05
04_TSC2-tuber-Martin2017 1.12882222 0.5896901 5.516971e-01
05_TSC2-SEN_SEGA-Martin2017 1.10262286 0.1589531 2.100271e-10
06_all_EpilepsyGene -0.14005479 0.5824869 1.000000e+00
07_high_conf_EpilepsyGene 0.78515948 0.6765502 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 0.55314337 0.8173131 1.000000e+00
09_Epi25_dataset 0.50324766 1.0893912 1.000000e+00
10_SFARI_all 0.84830074 0.4417919 5.458070e-01
11_SFARI_score4plus 0.85444526 0.5247009 1.000000e+00
12_DeRubeis_RDNV 0.97931656 0.8212165 1.000000e+00
13_Iossifov_RDNV 0.56798810 0.3344711 1.000000e+00
14_FMRP_Darnell 0.37157619 0.2985177 1.000000e+00
15_Voineagu_M12 -0.96692876 1.0826296 1.000000e+00
16_Voineagu_M16 1.59078471 0.2580783 2.044683e-07
17_ID_genes_Neel -0.21084642 0.6714659 1.000000e+00
18_ID_genes_pinto2014 0.10005101 0.6725687 1.000000e+00
19_Fromer_SCZ_FunDN -0.06263842 0.4742900 1.000000e+00
20_Cocchi_Pruning 1.12075978 0.6797064 9.516267e-01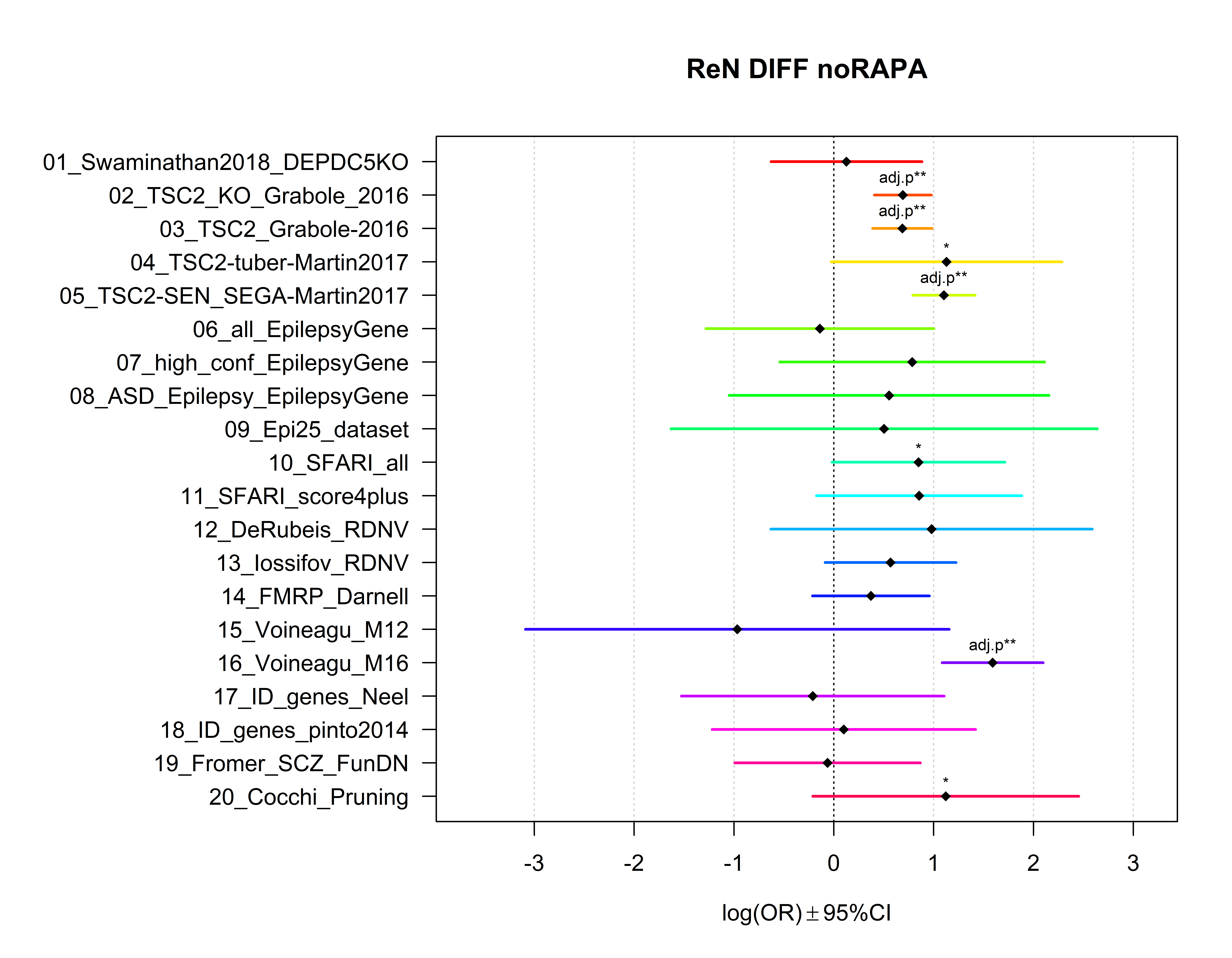
KO effect in noDIFF RAPA
D62
getlinmodoutput(targetline = "D62",targetdiff = "noDIFF",targetrapa = "RAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 64856 VWA1 0.7961187 1.591361e-02 0.8468392 2.620357e-02
2 3925 STMN1 -0.5685577 5.368831e-03 -0.6224295 1.905328e-02
3 9077 DIRAS3 1.9775503 6.473752e-17 1.5404231 2.797022e-07
4 64123 ADGRL4 1.8538504 7.722135e-05 1.6497307 3.911791e-03
5 6004 RGS16 1.1197917 1.317082e-06 1.0596972 2.861369e-03
6 29089 UBE2T -1.1211661 1.817676e-02 -1.2182221 4.945532e-02
7 149111 CNIH3 1.5083815 4.669966e-05 1.3495441 3.911791e-03
8 642938 INSYN2A 0.8617151 8.516245e-03 1.0006299 3.911791e-03
9 10581 IFITM2 -1.3249897 6.006754e-04 -1.5999599 8.700211e-05
10 4004 LMO1 -2.2672120 8.536899e-05 -2.2934700 8.213942e-09
11 960 CD44 1.4720618 1.480635e-07 1.3079443 6.140437e-05
12 1410 CRYAB 0.6463602 8.516245e-03 0.6999128 3.911791e-03
13 3709 ITPR2 1.4682329 7.883463e-05 1.4707635 7.064746e-04
14 84790 TUBA1C 0.8289466 1.522844e-05 0.7701332 1.404306e-02
15 160335 TMTC2 -0.9849648 8.069505e-03 -1.2868846 3.851921e-03
16 28526 <NA> 1.7946615 3.389494e-05 1.7259641 2.481597e-04
17 771 CA12 -0.8261845 2.555665e-04 -0.8015783 2.684880e-03
18 57611 ISLR2 -1.4138448 2.195926e-02 -1.3579845 3.754547e-02
19 56963 RGMA 1.1669346 1.509525e-08 1.0344198 3.851921e-03
20 2013 EMP2 -1.0920225 1.088237e-05 -0.7586095 3.222339e-02
21 643911 <NA> 2.7408935 2.461214e-02 2.9639647 4.974950e-03
22 2775 GNAO1 -1.1596849 6.876785e-06 -0.9286509 3.851921e-03
23 11031 RAB31 0.4783287 1.893394e-02 0.5147462 2.436953e-02
24 361 AQP4 0.7543128 2.751680e-03 0.8155313 5.746483e-03
25 1995 ELAVL3 -0.7800473 5.335671e-03 -0.7905853 2.436953e-02
26 348 APOE 1.0971097 6.637560e-05 1.0393426 2.965233e-03
27 79784 MYH14 -3.8557202 2.709801e-07 -2.3165157 2.436953e-02
28 151354 LRATD1 0.9410834 2.909158e-05 0.9934844 1.512676e-05
29 10267 RAMP1 1.2135869 1.056177e-06 1.0155836 3.681457e-04
30 1827 RCAN1 0.6625347 1.075336e-02 0.7270640 1.100868e-02
31 1826 DSCAM 1.3402666 8.069505e-03 1.7780242 6.401270e-06
32 2596 GAP43 1.3179970 1.456465e-22 0.9193596 2.388836e-08
33 5947 RBP1 -0.6459721 3.514077e-02 -0.8069445 9.568643e-03
34 4071 TM4SF1 1.8373599 6.909949e-07 1.4492589 3.890591e-03
35 3280 HES1 -0.7541376 8.973097e-04 -0.7161149 1.212638e-02
36 3490 IGFBP7 2.0277479 7.733560e-30 2.1251337 8.495963e-29
37 79966 SCD5 0.5668715 1.837778e-03 0.6470040 3.082466e-03
38 8404 SPARCL1 2.6711596 3.366738e-08 1.6962771 5.746483e-03
39 401207 C5orf63 0.6732523 3.217736e-02 0.8698764 1.422380e-03
40 85027 SMIM3 0.9707553 2.751680e-03 0.7790661 2.907467e-02
41 1026 CDKN1A 1.0793028 2.660019e-14 0.8376862 1.782308e-06
42 4188 MDFI -0.9944602 4.224909e-02 -1.2025531 2.447782e-02
43 4852 NPY -1.9241887 7.017330e-08 -1.5819410 2.956766e-06
44 23089 PEG10 0.7098065 2.053550e-03 0.7282390 8.417096e-03
45 1749 DLX5 -2.2913973 7.064968e-05 -1.8533389 8.417096e-03
46 25798 BRI3 -0.4962410 8.093883e-03 -0.5961950 3.911791e-03
47 5054 SERPINE1 1.9244309 3.434909e-05 2.1022473 1.782308e-06
48 4747 NEFL 1.1956686 1.505759e-05 1.2813013 1.832464e-05
49 2185 PTK2B 1.6672078 4.711653e-11 1.3930835 1.273791e-05
50 5327 PLAT 0.5886363 7.283803e-03 0.5708704 2.436953e-02
51 11075 STMN2 -6.0400119 7.422811e-03 -7.7180162 1.422380e-03
52 2171 FABP5 1.5435853 2.826377e-06 1.0667232 4.910801e-02
53 9568 GABBR2 1.7674445 1.682968e-16 1.6003234 2.740366e-09
54 56548 CHST7 1.0768094 4.713142e-04 1.1190497 2.965233e-03
55 5358 PLS3 1.0340243 6.595231e-04 1.2931225 1.535983e-06
56 10178 TENM1 -1.3363992 8.536899e-05 -1.3832123 7.013857e-03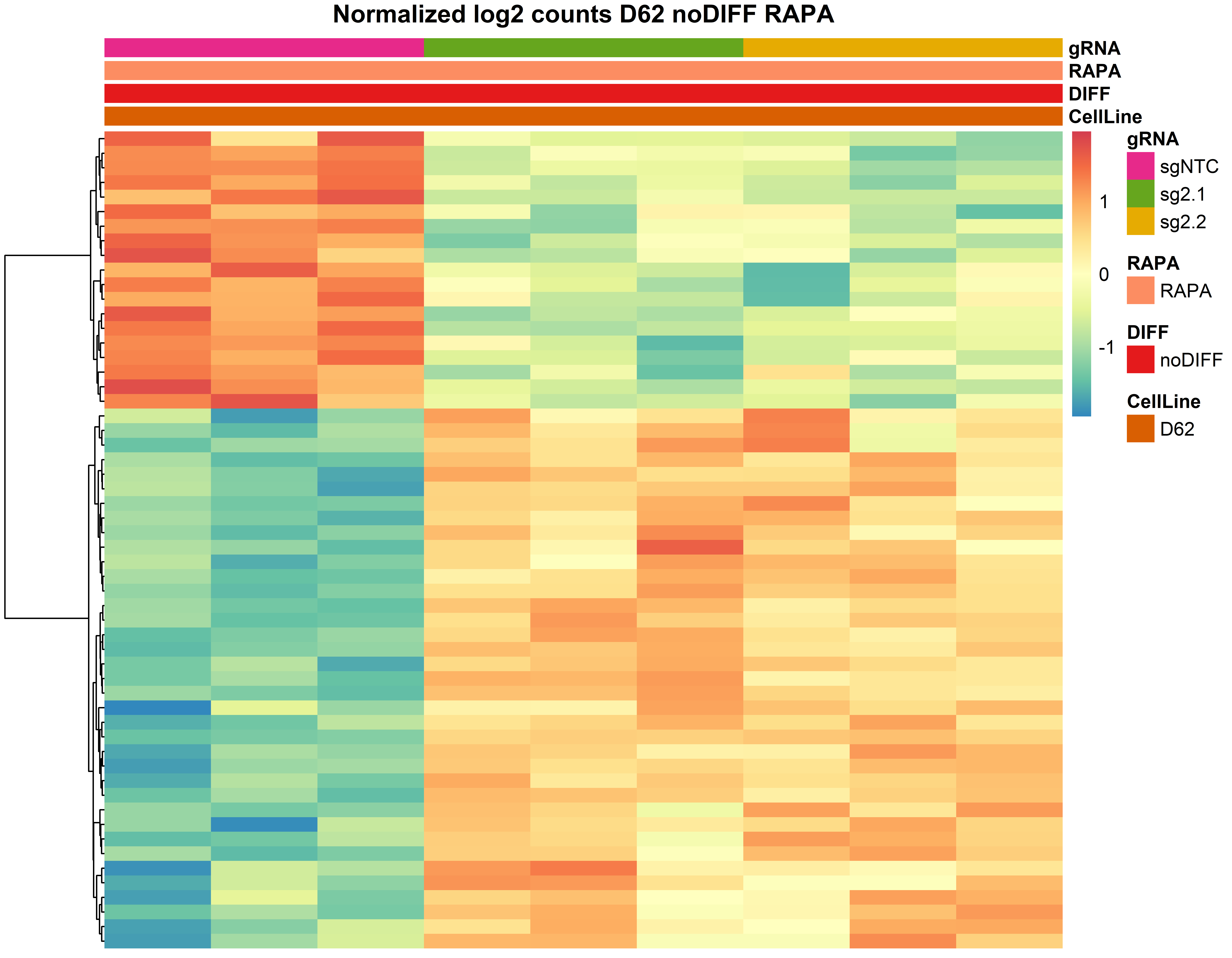
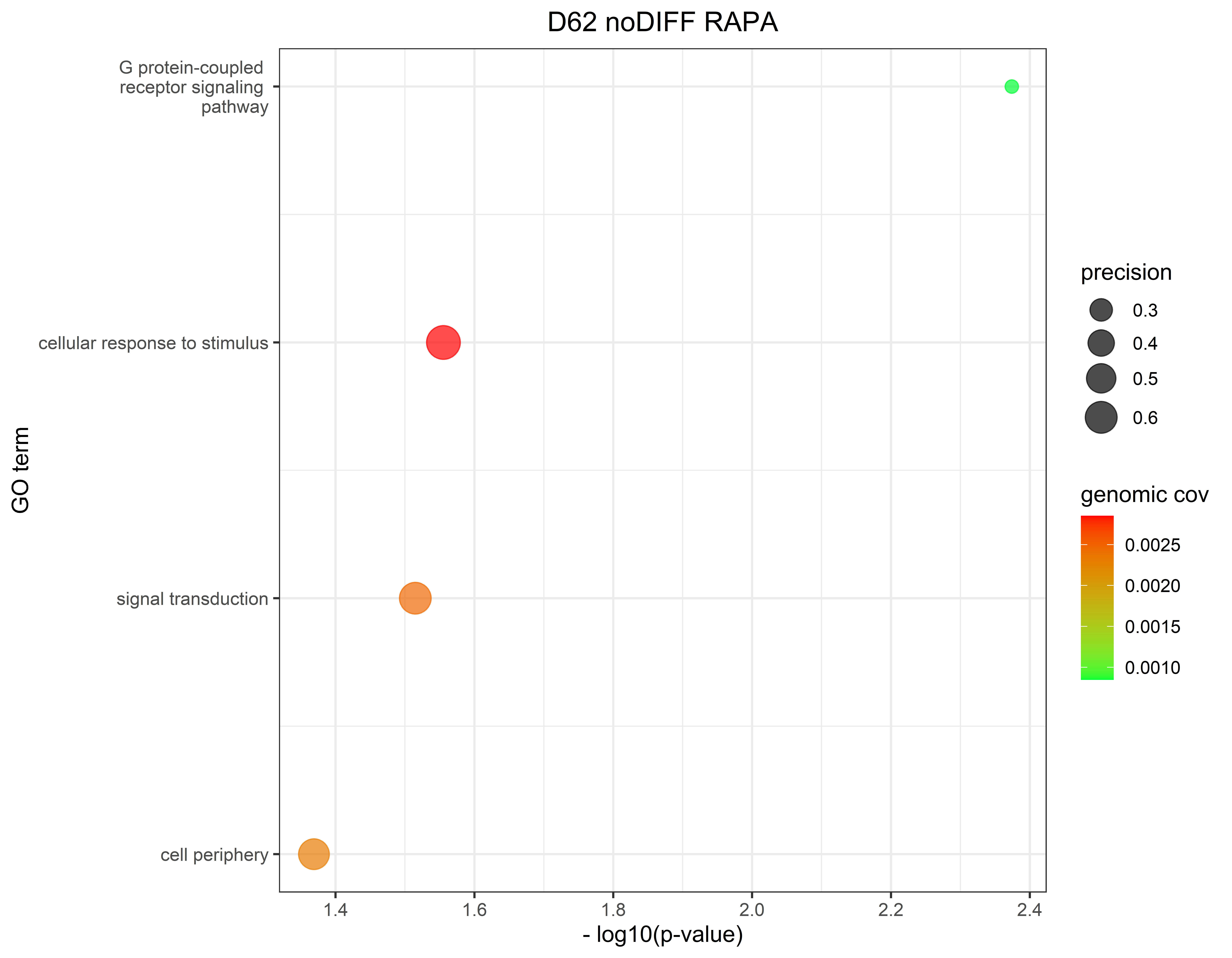
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.4004526 0.00994547 2.335242 5.191550e-01
02_TSC2_KO_Grabole_2016 2.3400242 1.33533983 4.138755 1.748419e-03
03_TSC2_Grabole-2016 2.2968387 1.26764835 4.334256 4.480912e-03
04_TSC2-tuber-Martin2017 6.9457439 1.36782496 21.963147 1.109565e-02
05_TSC2-SEN_SEGA-Martin2017 4.2623212 2.38392107 7.499436 7.408461e-07
06_all_EpilepsyGene 1.9797878 0.39393523 6.152519 2.051171e-01
07_high_conf_EpilepsyGene 1.9760607 0.04875929 11.678747 4.030732e-01
08_ASD_Epilepsy_EpilepsyGene 4.3453375 0.50713556 16.830829 8.305153e-02
09_Epi25_dataset 0.0000000 0.00000000 11.527050 1.000000e+00
10_SFARI_all 3.2205256 0.63913194 10.052591 7.383975e-02
11_SFARI_score4plus 2.8510008 0.33406010 10.963980 1.631103e-01
12_DeRubeis_RDNV 0.0000000 0.00000000 12.144245 1.000000e+00
13_Iossifov_RDNV 0.9544417 0.11240197 3.636546 1.000000e+00
14_FMRP_Darnell 2.0501461 0.71597593 4.804695 1.294979e-01
15_Voineagu_M12 0.0000000 0.00000000 2.656146 4.058569e-01
16_Voineagu_M16 4.7118897 1.63944077 11.100343 2.738157e-03
17_ID_genes_Neel 0.0000000 0.00000000 2.782932 6.457777e-01
18_ID_genes_pinto2014 0.0000000 0.00000000 3.788141 6.280384e-01
19_Fromer_SCZ_FunDN 2.0540941 0.53733918 5.614999 1.445296e-01
20_Cocchi_Pruning 2.7376465 0.06735287 16.287285 3.120327e-01
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -0.91515984 1.8854481 1.000000e+00
02_TSC2_KO_Grabole_2016 0.85016126 0.2862120 3.496839e-02
03_TSC2_Grabole-2016 0.83153370 0.3032501 8.961824e-02
04_TSC2-tuber-Martin2017 1.93812908 0.8290343 2.219131e-01
05_TSC2-SEN_SEGA-Martin2017 1.44981388 0.2964629 1.481692e-05
06_all_EpilepsyGene 0.68298965 0.8237543 1.000000e+00
07_high_conf_EpilepsyGene 0.68110533 1.8887576 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 1.46910342 1.0959594 1.000000e+00
09_Epi25_dataset -Inf NaN 1.000000e+00
10_SFARI_all 1.16954459 0.8250964 1.000000e+00
11_SFARI_score4plus 1.04767008 1.0939308 1.000000e+00
12_DeRubeis_RDNV -Inf NaN 1.000000e+00
13_Iossifov_RDNV -0.04662876 1.0913495 1.000000e+00
14_FMRP_Darnell 0.71791106 0.5367448 1.000000e+00
15_Voineagu_M12 -Inf NaN 1.000000e+00
16_Voineagu_M16 1.55008903 0.5386397 5.476314e-02
17_ID_genes_Neel -Inf NaN 1.000000e+00
18_ID_genes_pinto2014 -Inf NaN 1.000000e+00
19_Fromer_SCZ_FunDN 0.71983494 0.6841636 1.000000e+00
20_Cocchi_Pruning 1.00709860 1.8902594 1.000000e+00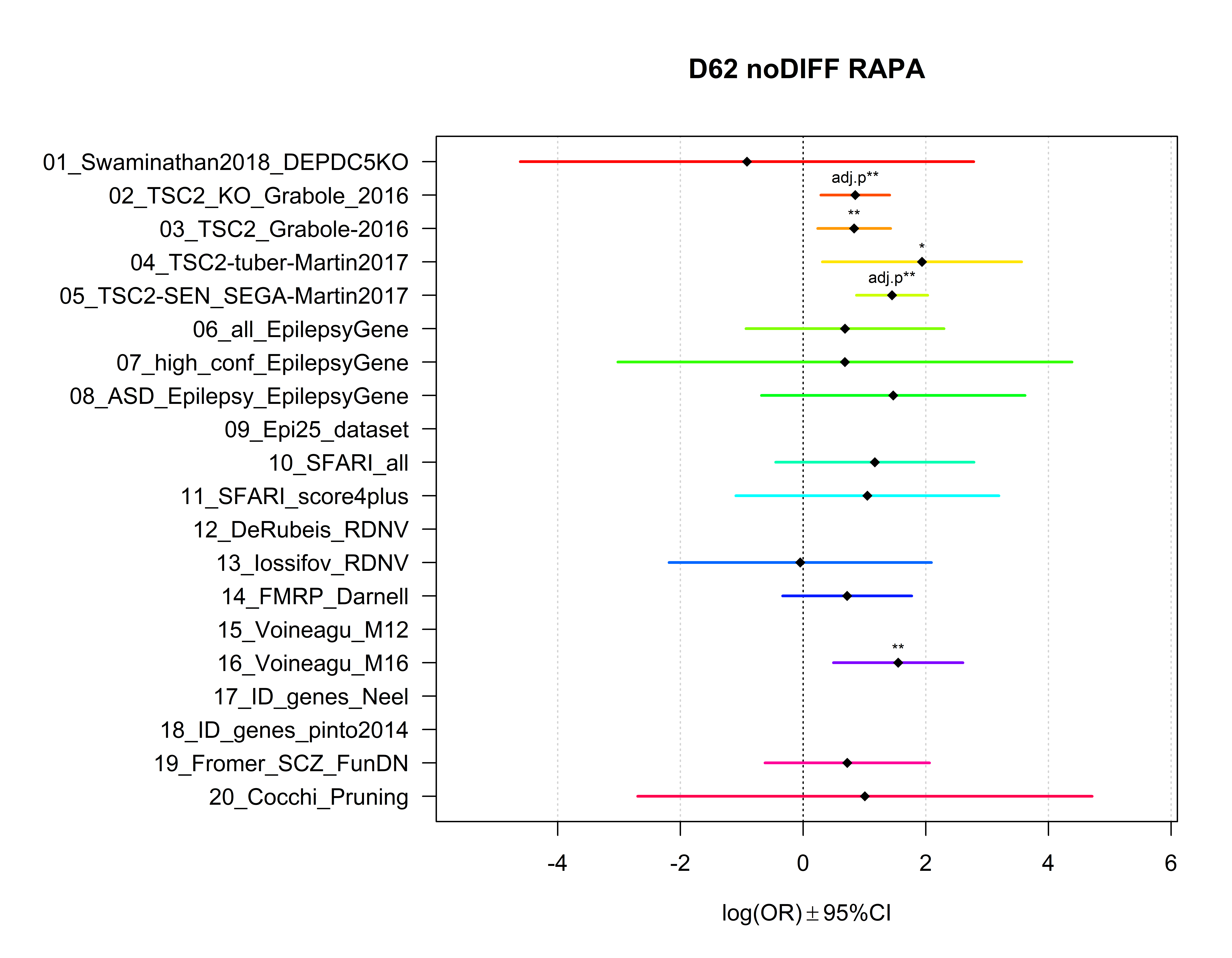
D244
getlinmodoutput(targetline = "D244",targetdiff = "noDIFF",targetrapa = "RAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 9636 ISG15 1.5824388 1.484743e-04 1.2829492 4.189665e-04
2 51150 SDF4 0.3756765 5.723901e-03 0.7747854 3.597778e-08
3 54587 MXRA8 0.5977314 1.411991e-03 0.7833115 7.754367e-07
4 440556 PRDM16-DT 1.1315687 1.249709e-04 0.8742095 7.287652e-03
5 63976 PRDM16 0.9034465 7.477217e-04 0.8046191 6.154955e-03
6 6146 RPL22 -0.4927361 7.363852e-04 -0.4927466 1.338800e-02
7 57449 PLEKHG5 1.6363155 3.230095e-04 1.3618997 2.095723e-02
8 23261 CAMTA1 -0.4277936 1.060105e-02 -1.0349325 2.432229e-08
9 54206 ERRFI1 -1.3165602 1.037519e-03 -1.4906598 7.650727e-04
10 5351 PLOD1 -1.1569610 1.240324e-06 -0.6079571 1.094670e-02
11 9927 MFN2 0.6205604 4.280046e-02 0.7025316 4.881864e-02
12 23254 KAZN 1.2345212 1.378131e-08 1.2945358 1.004445e-07
13 79180 EFHD2 0.5980518 6.568616e-03 0.6662879 5.180092e-03
14 440567 UQCRHL -1.2803222 8.776551e-04 -1.2369868 2.383955e-03
15 1969 EPHA2 -1.3444040 3.011003e-08 -1.1198203 6.815015e-08
16 51154 MRTO4 0.9001111 1.208548e-03 1.1304167 1.356098e-04
17 5322 PLA2G5 0.9577801 1.663496e-08 0.4856486 4.417485e-02
18 80818 ZNF436 -1.0371306 1.553439e-06 -1.1762577 9.841589e-08
19 3399 ID3 2.4835164 1.184258e-15 2.5010389 3.965480e-11
20 6135 RPL11 -0.9034285 1.834218e-07 -0.8115438 6.156107e-05
21 400746 NCMAP 2.9840207 1.064511e-04 2.3852290 1.763911e-02
22 10250 SRRM1 -0.3822801 3.181680e-02 -0.5304198 1.346275e-02
23 2537 IFI6 0.9741889 1.206228e-05 1.0236378 4.385758e-05
24 93974 ATP5IF1 -0.6261414 8.695858e-04 -0.6908658 2.192988e-03
25 9672 SDC3 0.3948315 1.958294e-02 0.4098725 3.427626e-02
26 3065 HDAC1 1.8390637 9.482389e-10 1.5314298 6.411181e-06
27 65108 MARCKSL1 -0.7073388 7.484675e-06 -0.6872964 9.988034e-05
28 79830 ZMYM1 1.8973526 1.712152e-02 2.0510684 9.972732e-03
29 9202 ZMYM4 0.8477035 1.770631e-03 0.8880819 1.951939e-03
30 54936 ADPRS 0.8709287 1.891722e-03 1.0841879 4.286178e-05
31 79647 AKIRIN1 -0.5293416 3.522515e-02 -0.6261662 3.241008e-02
32 4725 NDUFS5 -0.5830952 5.870251e-03 -0.6891822 8.998613e-03
33 643314 <NA> 2.1725309 4.898647e-07 2.3577943 1.002124e-07
34 4904 YBX1 -1.3904264 2.311743e-10 -1.2975221 2.323167e-09
35 6513 SLC2A1 -1.0617617 3.714706e-05 -0.7877181 3.154895e-04
36 26828 RNU5F-1 2.0302911 6.371631e-06 1.8065787 2.648764e-03
37 6202 RPS8 -0.5414825 1.190284e-06 -0.5575464 6.898967e-04
38 7388 UQCRH -0.6603621 1.640027e-04 -0.6892499 5.170060e-04
39 9528 TMEM59 -1.2316079 1.093312e-08 -0.6547585 1.963470e-02
40 205 AK4 -2.7310177 5.732523e-20 -3.0611435 4.595910e-13
41 54741 LEPROT -0.9389114 4.145418e-04 -1.6739560 3.367794e-11
42 1647 GADD45A -1.3406168 3.563966e-13 -1.2007802 1.082157e-10
43 9295 SRSF11 -0.4772421 3.689493e-03 -0.4496266 3.255072e-02
44 127254 ERICH3 4.1004930 2.151140e-02 4.3238701 2.009553e-02
45 5876 RABGGTB -1.4739387 2.272381e-02 -1.5220374 8.241496e-03
46 26289 AK5 -6.6003590 5.316610e-03 -6.4077026 9.311821e-03
47 2787 GNG5 -0.9030735 1.215658e-06 -0.7521400 1.000459e-04
48 3491 CCN1 -2.6405793 3.221564e-03 -2.0468527 4.443236e-04
49 9403 SELENOF -0.9302475 3.030301e-05 -1.1807101 1.279069e-06
50 2959 GTF2B -1.4583494 5.286202e-03 -1.1262157 2.822474e-02
51 6125 RPL5 -0.6367997 1.399423e-07 -0.5317626 5.771616e-04
52 9411 ARHGAP29 1.1653946 4.932031e-04 1.1596250 1.250806e-03
53 199857 ALG14 -2.0099368 1.930902e-05 -1.2869056 4.066935e-02
54 55119 PRPF38B -1.0050981 1.969653e-04 -0.7558508 7.029130e-03
55 6884 TAF13 3.1432810 4.391786e-16 2.6409385 1.938146e-09
56 84722 PSRC1 0.6166813 3.679775e-04 0.5774528 4.135456e-03
57 10542 LAMTOR5 -0.6859886 1.315846e-03 -0.7935888 8.983314e-05
58 10390 CEPT1 1.7325630 5.842110e-07 1.1284440 2.812949e-02
59 9939 RBM8A 0.8053453 1.599023e-09 0.6485053 1.615114e-04
60 200030 NBPF11 -0.5821200 2.713890e-02 -0.6515514 2.316868e-02
61 8337 H2AC18 -1.2977402 9.473626e-04 -1.3058019 9.580993e-04
62 51177 PLEKHO1 1.1133671 1.758600e-03 1.0186346 1.308626e-02
63 51107 APH1A 0.9963791 2.961773e-09 0.8144306 1.691489e-05
64 2029 ENSA -0.7210669 5.069302e-04 -0.7396818 3.820409e-03
65 29956 CERS2 -1.0922118 1.102436e-04 -1.2926428 2.963420e-05
66 10962 MLLT11 -1.5728076 4.391871e-12 -1.2923854 4.545775e-09
67 6281 S100A10 -1.5542314 2.194388e-05 -0.9993398 2.000028e-02
68 5872 RAB13 -0.9625542 1.498239e-07 -0.7637741 6.898967e-04
69 3782 KCNN3 0.8972991 1.779152e-02 1.2051483 4.906984e-04
70 55974 SLC50A1 1.1288291 1.406689e-03 1.0502843 8.855696e-03
71 54344 DPM3 0.7244787 1.674644e-02 0.8202974 1.898751e-02
72 55870 ASH1L 1.0599562 1.035521e-07 0.9928743 4.238631e-06
73 10485 MIR9-1HG 0.8086786 4.139934e-05 1.6709057 4.208470e-15
74 63827 BCAN 1.5532471 1.982801e-02 1.6892838 1.090152e-02
75 10763 NES 0.4079433 5.273539e-03 0.5694218 6.267806e-05
76 79590 MRPL24 0.9905111 5.694319e-03 1.1756719 9.838201e-04
77 25874 MPC2 -0.7726272 1.249143e-02 -0.9510370 2.192665e-02
78 481 ATP1B1 0.6534559 4.259089e-04 0.6993252 4.454316e-05
79 149041 RC3H1 1.0483608 3.350527e-02 1.1865388 5.598056e-03
80 9910 RABGAP1L 1.2266867 1.835811e-03 1.0466286 1.689752e-02
81 27101 CACYBP -0.8082859 8.814786e-03 -1.2054734 5.533371e-03
82 9917 FAM20B -1.0196979 5.220750e-03 -0.7592172 4.727448e-02
83 2752 GLUL -0.6833173 6.529329e-08 -0.8773989 4.292386e-08
84 81563 C1orf21 -0.6054954 6.615147e-03 -0.7207867 2.987392e-03
85 116496 NIBAN1 1.7929732 2.822698e-24 2.2280988 2.531885e-45
86 259266 ASPM 3.1818923 2.066537e-07 2.2533168 4.184859e-02
87 149345 SHISA4 0.7005681 4.574101e-02 0.8535469 1.245922e-02
88 6635 SNRPE -0.9221128 1.895528e-03 -0.9144657 3.896508e-03
89 55224 ETNK2 2.5831462 1.576230e-05 1.8539789 2.111881e-02
90 64710 NUCKS1 1.1967881 3.459305e-32 0.5989236 9.262941e-03
91 574036 <NA> -7.4593516 9.291791e-05 -7.2666958 2.165543e-04
92 127018 LYPLAL1 -1.6393321 1.663246e-07 -1.3579740 1.579159e-03
93 6726 SRP9 -1.2122287 4.135278e-07 -1.1517048 8.324859e-05
94 2052 EPHX1 1.2047806 2.671130e-03 1.0828554 9.705714e-03
95 3020 H3-3A -0.7006252 4.155287e-03 -0.6832695 8.007593e-03
96 388753 COA6 -0.8285424 3.154998e-02 -0.8040808 4.987914e-02
97 116228 COX20 -1.3698346 8.983419e-04 -1.7037172 2.264260e-06
98 51097 SCCPDH -1.0334920 6.477480e-05 -0.7709995 1.454060e-02
99 5209 PFKFB3 -0.9669253 2.240351e-04 -0.6608066 1.170651e-02
100 10659 CELF2 0.8972197 2.674221e-06 0.8151819 1.695273e-05
101 7431 VIM -0.6208164 2.762412e-06 -0.4048857 1.686708e-02
102 2572 GAD2 -5.6271621 6.625798e-09 -2.6726735 5.108571e-03
103 8829 NRP1 -0.9261206 5.007966e-07 -1.9809501 3.375055e-22
104 219749 ZNF25 -1.8097228 3.504311e-03 -1.1713950 4.546404e-02
105 8031 NCOA4 -0.8523913 8.727930e-05 -0.5980325 1.589383e-02
106 642826 <NA> 4.1420140 2.105696e-14 2.7679408 1.440485e-03
107 220963 SLC16A9 1.1803065 1.506299e-02 1.2230316 2.271632e-02
108 288 ANK3 1.3402628 3.229147e-03 1.2772510 1.614208e-02
109 3189 HNRNPH3 -0.8461245 7.945592e-07 -1.1500266 2.162598e-05
110 55680 RUFY2 1.3700833 1.408189e-03 1.4173596 3.519114e-03
111 80312 TET1 2.1154125 4.306909e-11 1.8346565 2.448617e-07
112 55749 CCAR1 -1.1702983 1.426437e-12 -0.5809230 5.498438e-03
113 219699 UNC5B 0.7114188 1.259044e-03 0.5167184 3.691250e-02
114 9806 SPOCK2 1.3285234 5.251144e-07 1.5233192 7.626200e-09
115 5033 P4HA1 -1.3247173 1.261018e-08 -1.5640109 2.099629e-05
116 729096 <NA> 4.5350368 2.036222e-06 3.4345158 2.574665e-02
117 142891 SAMD8 -1.2187331 7.389393e-03 -1.5312218 5.448833e-03
118 7417 VDAC2 -0.7118831 1.329084e-03 -0.5028832 3.987163e-02
119 6229 RPS24 -0.7305259 1.947391e-10 -0.5999155 5.163469e-06
120 57178 ZMIZ1 -0.5721452 1.152404e-03 -0.5240230 1.023035e-02
121 5995 RGR -1.1575992 1.435723e-02 -1.3838349 1.260681e-02
122 2894 GRID1 -0.9744374 1.443265e-02 -1.1174950 3.348652e-02
123 80351 TNKS2 1.0660217 1.026909e-06 0.7203107 1.142187e-02
124 83742 MARVELD1 1.0373666 3.352741e-08 0.7662849 1.344835e-03
125 81894 SLC25A28 1.5280121 3.234247e-03 1.5473076 5.637033e-03
126 282991 BLOC1S2 -0.7557446 2.011331e-03 -0.6518919 1.571734e-02
127 55719 SLF2 1.1337798 1.374251e-14 0.9604407 3.685936e-08
128 84833 ATP5MK -0.6046461 1.922291e-03 -0.7493619 6.201621e-04
129 85450 ITPRIP -1.5196831 1.985031e-02 -1.5912083 2.716982e-02
130 114815 SORCS1 -2.3375328 4.094130e-02 -3.5886849 1.429855e-03
131 8036 SHOC2 0.7463604 2.604400e-02 0.7716231 4.803865e-02
132 6934 TCF7L2 -0.5296395 4.536440e-02 -0.6965345 1.356053e-02
133 404636 DENND10 -0.8040247 2.075991e-03 -0.8048053 4.809852e-03
134 10935 PRDX3 -0.4712721 1.126754e-02 -0.6925035 3.584173e-04
135 2869 GRK5 -0.6878115 3.951226e-05 -0.5107161 5.310234e-03
136 9531 BAG3 -1.1004282 1.702147e-07 -1.0401403 1.381167e-04
137 22876 INPP5F -1.1029570 5.729009e-04 -1.1945523 4.220962e-04
138 10579 TACC2 1.5234636 5.789860e-08 1.5309721 9.740075e-06
139 5654 HTRA1 1.2019532 1.108631e-36 0.9903916 2.522760e-15
140 56647 BCCIP 0.8417131 3.873630e-04 0.8018413 2.264114e-03
141 664 BNIP3 -2.8319867 5.823775e-35 -2.2786315 1.137152e-20
142 10581 IFITM2 1.5109988 6.549627e-07 1.3074970 1.169641e-03
143 283232 TMEM80 -1.3327746 1.403477e-02 -1.2226829 8.515564e-03
144 975 CD81 0.2942571 4.408413e-02 0.3556005 3.099488e-02
145 4004 LMO1 1.4433843 6.220350e-11 1.3258866 3.649893e-10
146 133 ADM -1.1558237 1.797575e-14 -0.6475969 2.839770e-04
147 100463486 MTRNR2L8 2.0318202 4.235834e-62 2.0634858 5.935490e-05
148 1982 EIF4G2 0.3458726 3.094287e-02 0.5788820 1.346036e-03
149 5682 PSMA1 -1.0615781 9.014012e-06 -1.1012916 2.171462e-05
150 6207 RPS13 -0.6355080 2.211871e-05 -0.5301279 1.175016e-03
151 3939 LDHA -3.4450725 3.555975e-57 -3.2270563 4.818517e-50
152 89797 NAV2 -1.2715516 1.672697e-03 -1.3518098 1.553398e-03
153 55366 LGR4 -0.4596623 1.389123e-02 -0.6599830 1.297946e-03
154 79832 QSER1 1.2732257 5.842110e-07 0.9493788 5.798319e-03
155 966 CD59 -0.4470639 1.942333e-03 -0.4298558 2.265781e-02
156 4005 LMO2 -0.9439280 9.795680e-09 -1.0353899 1.362535e-09
157 25841 ABTB2 -1.9023949 1.148220e-08 -2.1948899 1.217194e-12
158 24147 FJX1 -1.2795283 1.485935e-06 -1.3571886 6.311075e-08
159 392 ARHGAP1 0.5524057 4.696285e-03 0.7313479 1.196175e-03
160 9793 CKAP5 1.3498049 1.190752e-10 0.9206405 3.693917e-04
161 5795 PTPRJ -1.4944567 4.669593e-09 -0.7948589 7.489466e-03
162 710 SERPING1 1.5477979 7.141345e-06 1.5806734 1.367010e-07
163 5007 OSBP -0.7514993 1.969092e-02 -0.8450710 1.734554e-02
164 219988 PATL1 -1.0861330 4.750628e-02 -1.1569902 5.180092e-03
165 9415 FADS2 0.7626196 1.026452e-06 0.7990381 1.071382e-05
166 2495 FTH1 -1.1447389 2.122669e-18 -1.0032806 2.806385e-11
167 7423 VEGFB -1.4590412 9.749048e-10 -1.2675706 9.435057e-07
168 5331 PLCB3 1.3387600 3.379337e-04 1.3823730 9.252685e-03
169 9379 NRXN2 1.3666355 2.553474e-09 1.3997911 2.401544e-09
170 10263 CDK2AP2 0.6093632 1.868947e-03 0.5886891 2.627495e-02
171 1374 CPT1A 1.0164792 3.092289e-20 0.7679831 1.375191e-08
172 219927 MRPL21 -0.8550106 4.875423e-02 -0.8281652 2.943702e-02
173 1717 DHCR7 -0.9898568 1.991312e-02 -0.7107950 2.855912e-02
174 23201 FAM168A -0.7923228 1.514209e-02 -0.9133797 2.001961e-02
175 254225 RNF169 2.2320646 1.382485e-02 1.9892575 3.990335e-02
176 143570 XRRA1 2.6346674 8.162851e-04 2.3630977 4.325461e-03
177 408 ARRB1 2.2685181 5.698223e-05 1.6957982 5.741357e-03
178 6188 RPS3 0.6455032 2.996139e-05 0.8239249 4.454634e-07
179 4135 MAP6 0.6653568 1.655575e-04 0.5934970 3.255681e-03
180 5547 PRCP -1.4113064 3.924570e-25 -1.2391368 1.972074e-10
181 1740 DLG2 1.4715011 2.963450e-15 0.9855713 2.342898e-04
182 11098 PRSS23 -0.5314697 2.316156e-02 -0.6340091 1.669958e-02
183 619383 SCARNA9 4.8265782 1.045932e-45 3.6141656 1.013795e-13
184 79101 TAF1D -0.8279775 3.410271e-02 -1.1685785 4.576136e-03
185 4361 MRE11 2.2080192 1.123628e-02 2.1289689 1.864014e-02
186 51503 CWC15 -0.6569116 2.854221e-04 -0.5998060 4.108747e-03
187 143686 SESN3 1.5208110 3.365994e-24 0.6697407 1.018237e-03
188 26521 TIMM8B -0.7276418 2.773393e-05 -0.5900655 1.967217e-03
189 5805 PTS -0.8421404 1.796029e-02 -0.9672717 1.497561e-02
190 23705 CADM1 -1.2893882 2.186411e-27 -1.1917463 2.734871e-11
191 6330 SCN4B 1.2799326 2.515569e-03 1.5017509 1.895289e-04
192 372 ARCN1 -0.6163297 1.131600e-02 -1.0063294 8.744126e-06
193 6230 RPS25 -1.1879930 1.087308e-10 -1.1487088 1.142438e-06
194 10525 HYOU1 0.4801722 1.991312e-02 0.5439716 1.002803e-02
195 867 CBL 0.9862442 7.630371e-04 1.1159437 5.209139e-04
196 23365 ARHGEF12 0.7708840 4.472279e-03 0.6039591 4.546651e-02
197 6309 SC5D -1.2578982 1.703773e-06 -0.6704416 1.613515e-02
198 6653 SORL1 0.9160623 4.693998e-03 1.2360072 4.872692e-05
199 84959 UBASH3B 0.9669846 5.205697e-04 0.7461784 5.877956e-03
200 3312 HSPA8 -0.8107801 3.612595e-13 -0.5565886 3.026526e-04
201 50863 NTM 1.6180618 2.150137e-08 1.0551223 7.257588e-03
202 27087 B3GAT1 0.8320166 3.689493e-03 0.7956606 1.030918e-02
203 81029 WNT5B -5.4734885 4.353497e-03 -5.3365568 7.838853e-03
204 928 CD9 -1.4360451 3.469720e-19 -1.6913935 8.625945e-17
205 51258 MRPL51 -0.6967226 2.937988e-03 -0.6955322 7.820459e-03
206 7167 TPI1 -1.0327660 2.121892e-06 -0.7844075 9.813256e-04
207 2026 ENO2 -1.3155183 7.505891e-13 -0.6506850 3.896812e-03
208 1822 ATN1 -1.7528870 1.805554e-14 -1.2290057 6.425936e-07
209 51279 C1RL -1.8665671 2.192402e-03 -1.5744124 3.264559e-02
210 8531 YBX3 -0.7382606 7.061527e-03 -0.6371255 4.546566e-02
211 2012 EMP1 -1.3494119 1.068472e-02 -1.8602544 1.890182e-05
212 55729 ATF7IP 0.8866408 2.954535e-03 1.2658836 1.146903e-05
213 3945 LDHB -0.6605578 1.633804e-05 -0.5790951 1.177618e-03
214 3709 ITPR2 1.0144843 3.199205e-03 1.0716091 1.786538e-03
215 441631 TSPAN11 0.9463665 6.891792e-03 1.0200880 8.443121e-03
216 440093 H3-5 -0.9320423 7.719732e-03 -1.2309948 1.305595e-04
217 636 BICD1 1.0501165 2.724718e-06 0.7343712 4.282353e-02
218 83448 PUS7L 3.5264050 2.710122e-08 3.0685051 5.825696e-05
219 9416 DDX23 1.1615000 2.153836e-07 0.8378215 9.345609e-03
220 7009 TMBIM6 0.7060460 1.363229e-09 0.6572957 2.324081e-06
221 57609 DIP2B 0.9470451 4.047441e-05 0.6473065 3.660628e-02
222 9802 DAZAP2 -0.9437682 4.271968e-08 -0.6995231 6.898967e-04
223 5204 PFDN5 -0.8609163 3.095867e-04 -0.5454512 2.821849e-02
224 140465 MYL6B -0.8054253 4.745960e-03 -0.8814983 2.903229e-03
225 4637 MYL6 -0.6163122 5.981035e-05 -0.6635701 6.276546e-04
226 506 ATP5F1B -0.4563243 3.048455e-04 -0.5400958 3.254161e-04
227 10728 PTGES3 -0.6156502 8.630233e-03 -0.9159722 9.753096e-05
228 4666 NACA -1.1459737 2.315934e-12 -1.3366001 6.239144e-15
229 6472 SHMT2 0.8459188 1.973739e-02 1.2364375 6.898967e-04
230 3798 KIF5A 3.6143608 2.034501e-03 3.4889098 6.053031e-03
231 10106 CTDSP2 -0.9923990 1.305389e-05 -0.5869442 2.513168e-02
232 4673 NAP1L1 -0.7120084 1.324329e-05 -0.4246846 1.740220e-02
233 1466 CSRP2 -0.6920638 9.505580e-06 -0.4917198 4.008592e-02
234 490 ATP2B1 0.9012531 2.614983e-05 1.0374740 1.108626e-06
235 6636 SNRPF -1.1324493 2.453012e-06 -1.0860560 3.082267e-06
236 79158 GNPTAB 1.1463324 2.173083e-05 0.6720679 4.565400e-02
237 7184 HSP90B1 -0.8634184 1.265523e-04 -0.7841444 2.164383e-03
238 728568 C12orf73 -1.5512684 3.242068e-03 -1.2469048 8.241496e-03
239 7296 TXNRD1 -0.6776423 3.615251e-02 -0.7882707 3.264559e-02
240 10970 CKAP4 -0.7690259 3.015689e-06 -0.9838818 1.017166e-08
241 51228 GLTP -0.4821875 4.618272e-02 -0.7599732 4.884879e-03
242 10094 ARPC3 -0.6102782 2.972650e-03 -0.9412420 6.729471e-06
243 84329 HVCN1 3.3747724 4.295913e-02 3.3686669 3.811470e-02
244 5501 PPP1CC -0.8128626 1.220379e-02 -0.8080460 1.116728e-02
245 643246 MAP1LC3B2 -1.4663643 1.267301e-04 -1.2280103 1.113274e-03
246 8655 DYNLL1 -0.5477195 3.986710e-04 -0.6919491 7.018900e-05
247 9761 MLEC 0.6722115 9.546733e-07 0.6169386 1.050568e-04
248 605 BCL7A 1.1315003 1.560655e-03 1.1728135 7.523336e-03
249 8099 CDK2AP1 -0.4694600 3.904163e-03 -0.4886292 1.635987e-02
250 7316 UBC 0.4997596 4.954409e-06 0.4822442 2.174564e-03
251 5901 RAN -0.3936993 3.107234e-02 -0.5350856 2.365134e-03
252 80324 PUS1 1.0286966 4.280046e-02 1.1286737 3.099488e-02
253 26278 SACS 1.5636743 7.664811e-07 1.1165926 6.584897e-03
254 221178 SPATA13 -0.8902048 6.035418e-04 -1.1657566 6.508979e-06
255 219287 AMER2 0.8461887 7.236179e-11 0.5680272 1.033381e-03
256 432369 <NA> -0.3692138 3.504351e-02 -0.4453423 2.397161e-02
257 51371 POMP -0.4804148 2.331793e-03 -0.6964847 2.260392e-05
258 9201 DCLK1 1.3179092 6.796682e-19 0.9142153 2.072691e-06
259 341640 FREM2 1.6954165 8.713935e-11 0.8982423 1.337545e-02
260 10186 LHFPL6 -0.6086307 1.863760e-02 -0.7115927 1.872176e-03
261 11193 WBP4 -0.6874295 2.487734e-02 -0.7996704 3.492492e-03
262 11215 AKAP11 1.4253930 3.820775e-11 1.1710854 7.201902e-06
263 387923 SERP2 2.4517691 3.534834e-03 2.6807674 3.979238e-04
264 2963 GTF2F2 -0.8249755 2.772878e-05 -0.5434308 2.988918e-02
265 7178 TPT1 -1.0059231 2.061131e-30 -0.8601923 2.276152e-11
266 23091 ZC3H13 -0.5529994 4.665459e-04 -0.5310758 5.180092e-03
267 9445 ITM2B -0.5220673 1.929989e-04 -0.6285540 4.805853e-05
268 51131 PHF11 1.4321384 1.437404e-03 1.5329898 1.737613e-03
269 27253 PCDH17 -0.9751132 2.876048e-06 -1.4640379 1.303741e-09
270 23077 MYCBP2 0.6255073 5.145461e-03 0.5997358 2.598748e-02
271 1638 DCT 1.0870598 1.498000e-02 0.9205481 4.979406e-02
272 5095 PCCA 2.0733463 3.953563e-08 1.2003783 3.959115e-02
273 1284 COL4A2 0.3695904 3.615251e-02 0.8082727 4.638740e-05
274 55002 TMCO3 -1.0332278 3.342980e-03 -1.0746997 3.079478e-03
275 6038 RNASE4 -0.9164290 2.077805e-02 -1.2163044 2.873368e-02
276 57680 CHD8 0.8968184 3.372939e-05 0.6991214 1.329218e-02
277 4323 MMP14 0.9850565 5.177454e-06 1.2786677 2.131713e-09
278 599 BCL2L2 -1.5865893 4.665920e-04 -1.7070956 3.857687e-04
279 51310 SLC22A17 0.5154397 1.814580e-04 0.4934902 1.966518e-03
280 10278 EFS 2.4883883 2.147077e-12 1.8752328 1.809868e-05
281 85446 ZFHX2 -1.6625580 9.933235e-08 -1.5340958 2.013072e-06
282 9985 REC8 0.7033157 1.748692e-04 0.5771245 2.657893e-02
283 26277 TINF2 0.7029526 1.197424e-02 0.7484572 2.706049e-02
284 29966 STRN3 1.5508301 8.057865e-08 1.2769628 9.613083e-04
285 25831 HECTD1 0.5459785 3.746719e-02 0.6708643 9.880774e-03
286 55012 PPP2R3C 0.5799324 4.782572e-02 0.6691687 1.549309e-02
287 5411 PNN -0.4168608 2.540171e-02 -0.7135687 1.201599e-02
288 23116 TOGARAM1 0.9600572 7.941597e-03 0.9851191 2.652754e-02
289 161357 MDGA2 1.6513023 8.923562e-11 0.9655134 3.225967e-03
290 6235 RPS29 0.4807624 3.455008e-04 0.6510355 2.116004e-05
291 6166 RPL36AL -0.9957680 9.082824e-08 -1.1016991 3.672404e-08
292 378706 RN7SL2 2.4805335 1.418235e-11 1.9179568 5.840687e-05
293 11183 MAP4K5 0.8785682 2.286557e-03 0.7507975 3.524523e-02
294 5684 PSMA3 -0.6584165 1.539840e-03 -0.8476550 1.354044e-03
295 9786 KIAA0586 1.6993680 2.599433e-04 1.4071980 1.440329e-02
296 51528 JKAMP -1.2023218 6.385736e-05 -1.3111579 3.396557e-06
297 6252 RTN1 -0.3639800 3.557987e-03 -0.6884447 1.770971e-08
298 51635 DHRS7 -0.6796524 5.647213e-04 -0.8371637 2.696340e-04
299 3091 HIF1A 2.1229447 3.487554e-16 1.6336410 3.065415e-07
300 23224 SYNE2 2.0670181 8.869231e-30 1.5662832 3.749292e-13
301 51109 RDH11 -0.5499171 2.484270e-03 -0.4760457 1.829743e-02
302 6430 SRSF5 0.5674659 2.224111e-03 0.5336157 1.077263e-02
303 64093 SMOC1 0.7132338 4.260506e-06 0.7084819 9.149357e-04
304 10965 ACOT2 0.6848311 1.900536e-02 0.7047440 3.214083e-02
305 4329 ALDH6A1 2.2418365 7.217714e-20 1.6061019 3.065415e-07
306 97 ACYP1 -1.3084894 4.216796e-03 -1.5900128 1.743211e-03
307 10972 TMED10 -0.8560095 8.713935e-11 -0.8328233 5.181240e-10
308 2353 FOS -1.6064915 1.243978e-03 -1.1608242 2.646130e-02
309 85457 CIPC -0.9313047 9.447552e-03 -0.8095020 1.789535e-02
310 9517 SPTLC2 1.1221050 7.990316e-06 1.1312278 2.118018e-05
311 9369 NRXN3 -1.6065716 3.449313e-02 -1.5887835 1.184699e-02
312 123016 TTC8 0.7142490 2.737574e-02 1.0099592 8.055467e-05
313 5700 PSMC1 0.7331662 2.158759e-02 0.8375600 8.052262e-03
314 9321 TRIP11 1.0443074 3.744096e-04 1.0147238 2.281472e-03
315 5641 LGMN -0.9730725 1.597545e-04 -0.7925332 6.240399e-03
316 23405 DICER1 0.8257131 2.331994e-02 1.1559007 5.170060e-04
317 8788 DLK1 -0.5609641 1.304295e-03 -1.7510012 1.281397e-17
318 3320 HSP90AA1 -1.1753867 9.781107e-29 -0.7131140 5.324325e-08
319 1983 EIF5 -0.6169436 1.244378e-02 -0.8321811 3.210971e-05
320 1152 CKB 1.1521331 1.734651e-16 0.7991025 1.815224e-04
321 9529 BAG5 0.8386522 7.607621e-04 0.6817342 2.821559e-02
322 64423 INF2 1.6891678 1.533649e-02 1.8987812 1.308795e-03
323 1397 CRIP2 1.1201174 9.734029e-09 0.9635236 8.096318e-06
324 646214 <NA> 1.2321069 9.224010e-04 0.9434233 1.500102e-02
325 400322 <NA> -1.5977707 1.410845e-02 -2.8006606 6.470850e-07
326 338429 SNORD109B 2.0097776 2.540171e-02 2.0919884 1.207255e-02
327 100505573 INAFM2 -1.4749615 1.319847e-02 -1.0051999 4.400500e-02
328 30844 EHD4 -0.6083114 2.158759e-02 -0.5089841 3.241008e-02
329 25963 TMEM87A -0.7257593 9.565968e-03 -0.6191638 1.354313e-02
330 10169 SERF2 0.4404454 2.170030e-03 0.6736692 6.526499e-05
331 2628 GATM 1.5643737 1.048643e-10 0.9616877 3.703696e-03
332 2200 FBN1 0.8607575 5.772459e-03 1.1847374 4.450604e-06
333 9728 SECISBP2L 1.0816217 1.255468e-03 0.8018022 4.377840e-02
334 55930 MYO5C 2.3785961 4.455859e-09 1.8100040 5.326105e-04
335 102 ADAM10 -1.0933933 6.246805e-06 -1.2182947 7.086852e-07
336 2958 GTF2A2 -0.8484710 2.716828e-03 -0.9048391 1.156914e-03
337 302 ANXA2 -0.5537088 1.866956e-04 -0.7113861 7.371241e-08
338 54832 VPS13C 1.1836564 4.451909e-06 0.8341882 1.007868e-02
339 4947 OAZ2 -0.4928508 4.484629e-03 -0.4782622 2.061321e-02
340 8766 RAB11A -0.9549842 1.054123e-06 -1.0568213 2.000083e-07
341 84465 MEGF11 -1.7186404 8.209982e-06 -1.1408627 4.145113e-03
342 6124 RPL4 -0.8771004 6.381839e-08 -0.5179205 2.001062e-02
343 10391 CORO2B 0.8589018 6.312711e-09 0.9243462 1.482286e-10
344 7090 TLE3 -6.7603617 2.978198e-03 -6.5677032 5.555131e-03
345 4756 NEO1 0.7737262 2.999006e-06 1.0788257 8.593413e-12
346 27020 NPTN -0.8818356 1.082249e-07 -0.6839712 1.133234e-02
347 9377 COX5A -0.6773013 9.758073e-05 -0.4486781 3.667230e-02
348 5780 PTPN9 -0.7547634 1.835840e-03 -1.2022981 3.903492e-07
349 257364 SNX33 1.6109347 1.408189e-03 1.8906458 1.031601e-04
350 5955 RCN2 -0.5281499 9.254131e-03 -0.9946384 2.552655e-04
351 10518 CIB2 1.5171911 4.633562e-04 1.4908279 1.120339e-03
352 1512 CTSH 2.0223758 3.355758e-19 1.2880767 2.118018e-05
353 54469 ZFAND6 -0.7238855 7.714215e-05 -0.6354142 2.675745e-04
354 100996492 CTXND1 -1.4610311 1.207149e-11 -1.0458236 3.462791e-06
355 57214 CEMIP -1.7258045 5.144530e-04 -1.2654567 1.334918e-02
356 9455 HOMER2 0.7921360 2.206341e-03 1.2020124 2.308369e-03
357 4828 NMB 1.3904482 1.209186e-03 1.7651951 4.082702e-05
358 23478 SEC11A -0.5071683 1.143399e-02 -0.7215752 1.530893e-03
359 11057 ABHD2 0.9867256 1.055214e-09 0.7448633 1.979386e-04
360 8826 IQGAP1 0.6887089 6.225407e-04 0.6857426 3.996255e-03
361 5045 FURIN 0.7288971 4.369405e-02 0.8065481 1.022489e-02
362 100507217 <NA> -0.6709382 1.434922e-05 -1.0526726 8.705861e-10
363 56963 RGMA 0.7056909 6.604404e-06 0.4876214 1.364992e-02
364 220 ALDH1A3 -3.6617546 2.321927e-02 -3.2939555 4.998178e-02
365 6627 SNRPA1 -1.4735138 6.025296e-06 -1.2717078 9.436098e-04
366 89941 RHOT2 0.6412986 2.211189e-02 0.6603700 2.526053e-02
367 146330 FBXL16 -1.3336109 1.052039e-03 -1.4302758 4.823952e-03
368 79006 METRN 0.5937960 2.511496e-04 0.5155136 1.360463e-02
369 4832 NME3 1.0266186 2.123065e-03 0.7703233 4.378514e-02
370 54442 KCTD5 -0.7554263 3.146392e-02 -0.9369020 2.344794e-02
371 23295 MGRN1 0.7979232 2.434262e-03 0.8281164 2.879230e-03
372 9665 MARF1 2.8542798 4.896935e-03 2.7023822 1.622654e-02
373 6210 RPS15A -0.7541864 9.596458e-06 -0.6651631 4.927704e-04
374 162073 ITPRIPL2 1.1724059 4.362093e-05 1.2406781 1.181085e-04
375 124152 IQCK -0.5529377 1.994773e-03 -0.8890497 2.921859e-06
376 51704 GPRC5B -0.5055507 1.513117e-03 -0.6900454 1.081939e-04
377 100132247 NPIPB5 -2.3215555 3.056852e-05 -1.8754768 1.802764e-02
378 4706 NDUFAB1 -0.5630920 8.843514e-04 -0.4566382 1.285059e-02
379 5930 RBBP6 -1.0066197 1.200452e-04 -0.7095351 2.843181e-02
380 728888 NPIPB11 -2.6355541 1.603400e-08 -1.9490314 8.570030e-05
381 283899 INO80E 2.9471199 9.759911e-07 2.1134615 9.468187e-03
382 226 ALDOA -1.3606083 2.798711e-17 -0.9728360 7.286096e-09
383 5595 MAPK3 0.6513239 3.661915e-02 0.7057474 3.446340e-02
384 11151 CORO1A 2.2349179 1.619569e-02 2.2342981 4.006712e-02
385 79077 DCTPP1 0.7113736 3.417531e-03 0.8565531 5.078489e-04
386 9274 BCL7C 1.5065615 1.943921e-08 1.1091868 6.498955e-04
387 6810 STX4 1.1925147 2.042837e-05 1.2388882 2.251825e-05
388 79001 VKORC1 -0.9601554 7.422678e-05 -0.6185071 1.877226e-02
389 2521 FUS 0.7410369 3.181030e-05 0.6204779 4.494913e-04
390 583 BBS2 -0.7482505 2.173867e-02 -0.8087094 6.589888e-03
391 4502 MT2A -1.2282267 1.664401e-22 -0.8229889 7.013897e-09
392 4493 MT1E 0.9300064 3.004138e-02 1.0751455 4.915824e-02
393 1783 DYNC1LI2 0.4974782 5.772459e-03 0.4532773 2.176852e-02
394 197258 FCSK 2.3984890 3.167946e-04 1.8808315 1.520342e-02
395 92154 MTSS2 0.5294757 1.914562e-02 1.0163654 3.065415e-07
396 5713 PSMD7 -1.0050699 2.112658e-04 -0.8860564 4.278111e-04
397 4166 CHST6 3.0645768 7.308068e-04 2.6810800 1.508054e-02
398 11345 GABARAPL2 -0.5831238 1.739853e-04 -0.5337674 8.088136e-03
399 54386 TERF2IP -0.6599059 6.545003e-06 -0.4920765 8.713752e-03
400 100130958 SYCE1L 1.8513710 6.371125e-05 1.9520282 5.917311e-05
401 4094 MAF 0.6601958 3.970478e-03 0.8544728 1.552264e-04
402 10200 MPHOSPH6 0.7186914 7.527866e-04 0.7569288 7.557718e-04
403 3281 HSBP1 -0.9791065 5.977306e-05 -0.8763515 2.071921e-05
404 23406 COTL1 -1.0468653 3.578360e-21 -0.7607910 6.151220e-07
405 81631 MAP1LC3B -0.7608972 1.625938e-04 -0.8909154 1.289561e-04
406 29123 ANKRD11 -0.6415374 5.192073e-07 -0.7459759 8.624795e-07
407 10381 TUBB3 -1.8799015 2.553474e-09 -1.5529605 8.242803e-08
408 7531 YWHAE -0.3729903 3.290109e-03 -0.3212018 2.610818e-02
409 5306 PITPNA 0.7210321 2.431354e-05 0.7542352 1.000956e-04
410 23302 WSCD1 1.0269661 2.545646e-09 0.8341752 3.197617e-05
411 1742 DLG4 1.0168062 1.241528e-06 1.0112762 5.170060e-04
412 37 ACADVL 0.6216676 3.701643e-03 0.6442395 5.809633e-03
413 11337 GABARAP -0.7586896 1.964721e-05 -0.5466087 1.233636e-02
414 23399 CTDNEP1 0.6708036 6.173454e-05 0.7816274 3.026526e-04
415 1984 EIF5A 1.1134736 3.154101e-08 1.3122690 6.205293e-09
416 8742 TNFSF12 1.0187220 1.459895e-02 1.1437520 6.790872e-03
417 6154 RPL26 -1.0460345 2.509745e-19 -0.8622822 2.463887e-11
418 4213 <NA> 1.5820650 6.035758e-03 1.6092586 1.622654e-02
419 9611 NCOR1 -0.6081892 1.695747e-02 -0.5495149 1.707968e-02
420 7314 UBB -0.4691251 1.061397e-04 -0.3483102 2.610818e-02
421 3996 LLGL1 1.4897518 7.828128e-12 1.1759520 6.767713e-06
422 22905 EPN2 1.4997465 4.376134e-20 1.2335509 1.938146e-09
423 5598 MAPK7 -1.7273884 1.782831e-02 -1.8507405 7.045468e-03
424 224 ALDH3A2 1.0707664 4.848732e-06 0.8587628 4.958752e-04
425 218 ALDH3A1 4.7649674 6.798268e-11 4.2115605 1.559989e-05
426 100462977 MTRNR2L1 0.5390923 4.998631e-04 0.4795306 3.597145e-02
427 26118 WSB1 -0.6667744 5.838890e-04 -1.0155676 3.164552e-06
428 6147 RPL23A -1.2495909 3.507351e-10 -1.4730771 3.905016e-14
429 147015 DHRS13 -1.5904096 4.436896e-03 -1.3580087 2.036057e-02
430 163 AP2B1 0.9170481 4.391871e-12 0.5561412 1.686939e-03
431 9349 RPL23 -0.9609142 1.448658e-14 -0.7499664 1.973976e-06
432 6143 RPL19 -0.6496690 7.202380e-07 -0.4333177 7.007300e-03
433 5709 PSMD3 0.4486873 3.803015e-02 0.4953493 4.337170e-02
434 10209 EIF1 -0.8655077 1.218459e-11 -0.7102267 1.207773e-05
435 60681 FKBP10 0.8066666 5.091359e-05 0.6908554 4.406530e-03
436 6155 RPL27 -0.5392102 4.821528e-04 -0.5422350 4.327176e-04
437 7343 UBTF 0.6029705 9.174393e-03 0.8177925 2.046263e-04
438 5164 PDK2 1.2152361 1.904797e-04 1.1907447 4.563305e-05
439 1277 COL1A1 -2.8471299 2.995600e-39 -1.8285507 2.040815e-13
440 51747 LUC7L3 -0.6610476 3.857106e-06 -0.9295326 1.304088e-07
441 400604 TOB1-AS1 -6.1036766 3.873388e-03 -6.8727609 1.644381e-03
442 404093 CUEDC1 1.0923463 2.369303e-09 1.2435518 2.771823e-09
443 6426 SRSF1 -0.5865971 9.832837e-03 -0.7513147 3.903825e-02
444 51651 PTRH2 -1.1640318 3.574377e-02 -1.5787622 5.267260e-03
445 653653 <NA> 1.4239438 3.467382e-09 1.7399199 6.011646e-18
446 342538 NACA2 -0.9300780 3.015689e-06 -1.1975437 1.017166e-08
447 9902 MRC2 1.1038193 1.638472e-20 1.1162576 8.625945e-17
448 1534 CYB561 2.4158674 1.680369e-05 1.6040817 4.933118e-02
449 57003 CCDC47 0.7906737 9.598831e-04 0.8245096 2.090641e-03
450 5705 PSMC5 -0.4908692 2.234183e-02 -0.5234272 2.177510e-02
451 5573 PRKAR1A -1.0014435 1.197939e-07 -0.8420927 6.534938e-05
452 388419 BTBD17 1.3664531 3.650656e-13 1.1790808 1.784528e-08
453 54868 TMEM104 1.3112915 6.952907e-03 1.3391409 4.117834e-03
454 23510 KCTD2 0.9061594 2.145458e-04 0.5868645 3.890927e-02
455 10476 ATP5PD -1.0698931 2.195591e-09 -1.1032170 1.084478e-08
456 3021 H3-3B -0.5430925 1.086129e-04 -0.7777642 2.435996e-08
457 91107 TRIM47 1.6792228 7.217060e-03 1.8933073 8.918467e-04
458 201292 TRIM65 1.1240151 2.665472e-02 1.4725577 8.160852e-04
459 51 ACOX1 1.1598918 1.702593e-13 0.5881737 9.993501e-03
460 10801 SEPTIN9 0.4981700 1.926090e-05 0.3542610 1.262411e-02
461 9021 SOCS3 -2.0406971 7.667438e-07 -0.9303624 2.637179e-02
462 3959 LGALS3BP 0.4278459 4.764323e-02 0.5390982 2.085401e-02
463 114897 C1QTNF1 1.1314203 1.006811e-07 0.7203363 1.501335e-02
464 284184 NDUFAF8 0.6478430 8.413831e-06 0.6534928 1.456353e-03
465 71 ACTG1 -0.4680908 2.715692e-06 -0.4227503 1.250806e-03
466 201254 CENPX 0.9654809 8.055192e-04 0.6960919 3.037249e-02
467 5881 RAC3 1.4383392 1.309392e-03 1.3111040 2.163096e-02
468 9123 SLC16A3 -7.4951724 1.577182e-08 -4.6801469 6.413758e-08
469 23347 SMCHD1 1.4355319 1.582610e-03 1.2198838 1.376472e-03
470 10627 MYL12A -0.8038030 5.724630e-05 -0.6752340 2.496280e-03
471 103910 MYL12B -0.8575369 1.621226e-10 -0.8117296 5.233679e-08
472 284217 LAMA1 -2.2257179 6.361789e-03 -2.0289047 7.975250e-03
473 201475 RAB12 1.3339036 5.817152e-07 1.1245542 1.084291e-04
474 23253 ANKRD12 -0.8681116 2.601905e-07 -0.9152023 3.065415e-07
475 114876 OSBPL1A 1.0766236 7.366692e-06 0.9044615 1.100923e-03
476 361 AQP4 1.5656176 1.853286e-21 1.4074809 5.825739e-17
477 498 ATP5F1A -0.6416771 4.356958e-06 -0.5865395 1.462442e-04
478 84064 HDHD2 -1.1724552 3.871622e-05 -0.8265877 3.960607e-02
479 6139 RPL17 -0.8860713 3.459308e-04 -0.8203003 5.170060e-04
480 10449 ACAA2 -0.9835198 1.748729e-04 -0.8956480 1.526013e-03
481 220164 DOK6 1.1034941 3.803015e-02 1.4893884 7.937452e-03
482 9306 SOCS6 0.6771815 1.511831e-03 0.6649241 1.376269e-02
483 9658 ZNF516 -1.3949158 6.602410e-03 -1.2481438 4.683662e-02
484 10907 TXNL4A -0.8554553 3.464622e-03 -0.8401470 3.249287e-03
485 57418 WDR18 0.9082210 1.322176e-02 0.8619321 2.884778e-02
486 6209 RPS15 -1.1655410 9.427873e-13 -0.7401840 8.264494e-04
487 4946 OAZ1 -0.9144420 1.035688e-06 -0.8295024 5.396696e-07
488 51690 LSM7 0.6851350 1.625938e-04 0.6592302 2.082811e-03
489 7064 THOP1 1.5633735 7.422678e-05 1.2915626 1.982972e-02
490 4782 NFIC -0.4955306 1.019937e-02 -0.6084822 2.238075e-03
491 10362 HMG20B 0.9104760 7.168929e-04 0.9698949 8.701801e-05
492 1613 DAPK3 1.1210634 8.583011e-03 1.4670027 5.149358e-05
493 51341 ZBTB7A -0.5329792 2.174952e-02 -0.6014640 1.680627e-02
494 10226 PLIN3 0.7468669 1.731812e-04 0.6718949 2.987392e-03
495 5802 PTPRS 0.5814723 3.792843e-02 0.5671801 4.497410e-02
496 9667 SAFB2 -0.6302106 1.991312e-02 -0.7619429 2.644829e-03
497 25873 RPL36 -0.5978185 3.267858e-03 -0.5381396 2.431132e-02
498 126328 NDUFA11 0.5444187 3.504072e-03 0.5460042 5.670675e-03
499 8570 KHSRP 0.4531844 3.890512e-03 0.6730349 1.215965e-04
500 5609 MAP2K7 0.8145074 2.515161e-02 1.0251983 6.875258e-03
501 59286 UBL5 -0.6634272 1.135321e-04 -0.5617379 5.769987e-03
502 11140 CDC37 0.6525490 2.916640e-04 0.9248559 1.478986e-08
503 6597 SMARCA4 -0.6388741 2.116413e-04 -0.4370694 3.788967e-02
504 664709 HNRNPA1P10 -0.8202349 1.319885e-03 -1.1368359 3.046410e-06
505 30000 TNPO2 0.9871910 2.879390e-12 1.0137554 1.123235e-10
506 811 CALR 0.5171798 1.055917e-04 0.5652609 2.200134e-04
507 4784 NFIX 1.3494924 3.877122e-10 1.1380430 5.245494e-06
508 4854 NOTCH3 -0.7984197 1.107035e-04 -0.5367607 3.290530e-02
509 23309 SIN3B 0.5376397 1.786169e-02 1.1559082 2.557046e-09
510 29086 BABAM1 0.5444527 3.192491e-02 0.5725905 4.870170e-02
511 684 BST2 4.1044688 2.448898e-07 2.4245661 4.133457e-02
512 23025 UNC13A -1.4931368 1.626257e-03 -1.2487069 2.646238e-02
513 6142 RPL18A -1.1924036 4.972920e-05 -0.8330070 2.756954e-02
514 54858 PGPEP1 0.5743370 2.946539e-02 0.6945012 5.880719e-03
515 9454 HOMER3 0.6580178 2.728967e-02 1.1169763 1.394607e-05
516 126382 NR2C2AP 1.7684018 6.227875e-03 2.0415365 1.297946e-03
517 1463 NCAN 2.0063903 2.093700e-39 1.5848372 1.802605e-19
518 199777 ZNF626 1.2503435 2.324914e-02 1.2250439 3.137160e-02
519 9141 PDCD5 -1.1924111 3.352741e-08 -1.5108428 1.043035e-10
520 2821 GPI -0.8182675 7.141345e-06 -0.6441403 9.803840e-03
521 57655 GRAMD1A 0.6170433 3.368466e-02 1.2095016 2.797167e-07
522 7392 USF2 0.8150802 7.818162e-04 0.8852666 2.003848e-03
523 55851 PSENEN 0.6441546 2.705447e-02 0.8293852 1.338893e-03
524 390927 ZNF793 1.1624545 5.507474e-03 1.1992339 7.007300e-03
525 81 ACTN4 -0.9711769 9.383044e-11 -0.8946542 1.902623e-04
526 1891 ECH1 -0.8841565 1.188934e-06 -0.7721249 7.751143e-06
527 2091 FBL 0.8150981 5.297385e-05 0.7676569 9.436098e-04
528 208 AKT2 0.6374496 6.928342e-05 0.5051990 1.813853e-02
529 6223 RPS19 -0.6876310 1.035756e-05 -0.6189867 2.820892e-03
530 348 APOE 1.1522867 2.550867e-12 1.1986122 8.038100e-11
531 27113 BBC3 0.7761841 5.367863e-05 0.7609061 5.485502e-05
532 6415 SELENOW 0.7770321 9.579005e-08 0.6348706 2.338289e-04
533 9266 CYTH2 0.6058005 2.732904e-02 0.7927179 1.934439e-03
534 114783 LMTK3 0.9480215 3.949142e-02 1.2953316 2.109213e-03
535 587 BCAT2 1.5592921 8.160051e-07 1.0495492 2.269406e-02
536 4924 NUCB1 0.7887961 1.329084e-03 0.9847285 2.839770e-04
537 2512 FTL -0.3556224 2.866145e-03 -0.4019268 5.246853e-03
538 23521 RPL13A -0.4194443 1.506299e-02 -0.4215042 2.365199e-02
539 6205 RPS11 -0.6614844 1.204615e-04 -0.4856142 1.551758e-02
540 284361 EMC10 0.6624030 4.771072e-04 0.5769863 1.183788e-02
541 94030 LRRC4B 0.4740838 2.088254e-02 0.6435874 1.955646e-03
542 22870 PPP6R1 1.2045892 2.694076e-05 1.2526980 2.118018e-05
543 3589 IL11 -2.3886842 2.449178e-02 -3.4881869 2.358497e-02
544 57106 NAT14 1.0891691 1.904718e-04 1.1249846 3.828627e-04
545 11338 U2AF2 1.0205563 2.423170e-08 1.2225093 1.970862e-11
546 10782 ZNF274 -2.3087036 2.034501e-03 -2.5598429 7.478455e-04
547 1 A1BG 1.0668549 1.135968e-02 1.0326944 4.966984e-02
548 27243 CHMP2A -0.9064416 7.832469e-07 -0.6666526 6.246455e-03
549 52 ACP1 -1.1377447 3.462058e-04 -0.8916591 2.090641e-03
550 7837 PXDN -0.3290889 3.408553e-02 -0.4502360 1.257367e-02
551 55256 ADI1 0.9224474 1.666662e-04 0.6410400 4.013384e-02
552 6664 SOX11 0.6912783 1.701194e-03 0.7174623 1.703667e-02
553 10971 YWHAQ -0.8210504 1.000003e-06 -0.6755020 1.226452e-04
554 9475 ROCK2 0.8346411 1.136763e-02 0.8095556 1.131041e-02
555 100506474 <NA> -1.5982889 2.440088e-02 -2.2156404 1.393295e-02
556 23369 PUM2 -0.8245376 1.197424e-02 -0.9152835 9.252685e-03
557 1788 DNMT3A 0.9579356 1.796029e-02 1.2639074 7.176007e-04
558 2498 <NA> -0.9087544 1.435723e-02 -1.1999027 1.444109e-03
559 29959 NRBP1 0.9624157 1.830496e-06 0.8847736 3.893351e-04
560 64838 FNDC4 0.9644681 8.342670e-04 0.9341919 4.833162e-03
561 23160 WDR43 1.2590444 2.095399e-03 1.2698739 5.741357e-03
562 57448 BIRC6 1.1606625 2.511496e-04 1.0147184 6.686043e-03
563 4052 LTBP1 -1.3724028 1.778799e-03 -1.7960557 3.974205e-08
564 5610 EIF2AK2 1.5877796 6.858212e-09 1.0966518 1.072710e-03
565 9167 COX7A2L -0.7887471 6.017926e-03 -0.9864544 3.965731e-03
566 9581 PREPL 1.4106356 1.490691e-08 1.1669508 6.670263e-05
567 90411 MCFD2 -0.8534024 6.212943e-04 -0.7653025 2.320449e-02
568 618 BCYRN1 1.1481897 4.687568e-11 0.7904782 1.743211e-03
569 6233 RPS27A -0.5869447 4.180878e-07 -0.4716746 2.436719e-03
570 2202 EFEMP1 -1.5040760 2.610553e-10 -1.4541560 4.618032e-10
571 150962 PUS10 -1.6267832 1.809134e-04 -1.1783226 1.738090e-02
572 9736 USP34 0.6629489 2.118902e-02 0.6043363 4.928176e-02
573 4190 MDH1 -1.3014972 3.011003e-08 -1.2366448 2.140939e-07
574 200734 SPRED2 -1.4266956 2.030167e-05 -1.6172656 3.995397e-06
575 27247 NFU1 -0.8449345 4.144806e-02 -1.2119069 2.533826e-04
576 22848 AAK1 0.5601755 2.011326e-02 0.7988644 6.392189e-04
577 5093 PCBP1 -0.8498054 2.218219e-06 -0.5342829 5.183356e-03
578 6637 SNRPG -0.8997136 3.242095e-09 -0.9151109 2.855369e-07
579 94097 SFXN5 0.9463019 1.681136e-05 0.9579809 4.220126e-05
580 116540 MRPL53 -1.0324314 5.395323e-05 -0.6913691 1.262411e-02
581 3099 HK2 -3.3309343 1.594738e-16 -4.2459586 1.346036e-03
582 9168 TMSB10 -0.8474615 2.767975e-10 -0.5227594 2.016952e-03
583 83439 TCF7L1 -2.0526037 4.683382e-03 -1.9991394 1.305595e-04
584 84913 ATOH8 1.1852155 5.269336e-09 1.0431946 8.096318e-06
585 55818 KDM3A -2.0335439 6.702358e-07 -2.0759424 1.607238e-06
586 729857 RGPD2 3.5272906 3.171255e-04 3.5276271 1.397300e-03
587 56910 STARD7 -1.0227191 1.006811e-07 -1.1204582 2.171022e-08
588 25972 UNC50 -0.8665031 3.410271e-02 -1.5261007 1.845494e-03
589 51263 MRPL30 0.8703242 5.947905e-03 0.8017773 2.918484e-02
590 9669 EIF5B -1.0963573 8.362389e-19 -0.8541908 5.292832e-08
591 4862 NPAS2 -5.7368942 3.130281e-03 -4.5240348 1.096030e-02
592 6160 RPL31 -0.9658469 2.797911e-17 -0.7099248 5.184623e-07
593 9448 MAP4K4 0.4100621 2.284218e-03 0.4202157 2.125937e-02
594 100506421 PANTR1 0.9780882 4.942034e-08 0.6353487 6.010093e-03
595 102682016 LINC01159 1.7227936 1.207016e-05 1.4002439 2.631022e-03
596 100288695 LIMS4 -1.6780865 1.324682e-06 -1.3833020 6.586686e-04
597 6574 SLC20A1 -0.6693197 2.565649e-03 -0.6163021 2.489354e-02
598 55240 STEAP3 -1.4419739 1.899507e-02 -1.8258206 1.253088e-03
599 5899 RALB 0.7628939 4.805134e-03 0.9772579 1.809868e-05
600 9394 HS6ST1 -0.9357036 1.382177e-04 -0.9316714 5.797352e-06
601 80097 MZT2B 0.6017073 3.355527e-03 0.6834076 3.508558e-03
602 653784 MZT2A 0.9549674 9.727316e-04 0.8347819 1.034728e-02
603 116372 LYPD1 -6.2885116 1.774920e-02 -6.0958524 2.657893e-02
604 23190 UBXN4 -0.7151932 7.489185e-05 -0.5442665 1.592498e-02
605 1615 DARS1 -0.4502497 4.357571e-02 -0.6777391 1.095256e-03
606 55777 MBD5 1.5335735 1.533649e-02 1.5209488 8.913731e-03
607 27249 MMADHC -1.0656269 1.362946e-02 -0.8307710 3.312198e-02
608 90 ACVR1 -2.0416431 5.843736e-03 -1.7937962 9.494858e-03
609 5937 RBMS1 -1.2392401 6.887964e-07 -2.4280154 2.528686e-18
610 9360 PPIG -0.6234780 9.426565e-04 -0.4444366 1.463157e-02
611 29081 METTL5 -1.4305197 2.031223e-03 -2.0690238 7.930938e-05
612 1745 DLX1 -0.7904554 4.379604e-02 -0.8927985 4.910555e-02
613 5163 PDK1 -2.1285998 2.115807e-02 -1.7678435 1.079832e-03
614 440926 <NA> -0.9154993 3.278871e-02 -1.0285757 3.559935e-02
615 518 ATP5MC3 -0.5910226 1.241528e-06 -0.4956253 6.875258e-03
616 10651 MTX2 -1.4065501 4.954487e-04 -1.4724389 5.725756e-03
617 220988 HNRNPA3 -0.7180730 2.563000e-03 -0.6992930 4.523637e-03
618 10787 NCKAP1 -0.9419657 5.004447e-04 -0.8625636 6.042247e-03
619 94101 ORMDL1 -0.9117302 2.497988e-03 -0.6406131 3.290530e-02
620 23671 TMEFF2 -6.9412488 9.263510e-10 -1.6671606 1.252755e-02
621 3336 HSPE1 -0.9361015 1.620163e-07 -0.5738337 2.943702e-02
622 4709 NDUFB3 -0.8599914 2.195084e-02 -1.5430307 1.972074e-10
623 66008 TRAK2 1.1469229 3.828445e-05 0.7988402 1.665171e-02
624 7341 SUMO1 -1.1692703 1.498417e-06 -1.0808101 3.724102e-06
625 4133 MAP2 0.9717932 3.904716e-15 0.8135220 6.413758e-08
626 2335 FN1 -0.6180557 7.870373e-03 -0.5129073 4.209339e-02
627 3488 IGFBP5 -1.0704516 8.624182e-21 -1.0255359 2.373062e-12
628 6508 SLC4A3 1.6170398 5.997335e-07 1.4806608 1.305595e-04
629 7857 SCG2 -0.9795131 8.076526e-13 -0.8821592 7.286096e-09
630 5757 PTMA -0.7980282 2.664439e-04 -0.5411530 2.348092e-02
631 26058 GIGYF2 0.7938093 7.465367e-05 0.6147934 9.513596e-03
632 2637 GBX2 -7.6628226 2.593849e-05 -7.4701666 6.396227e-05
633 547 KIF1A 0.5294785 1.624405e-03 0.4920022 1.093023e-02
634 55968 NSFL1C 0.5829945 3.826949e-02 0.7680300 2.534991e-03
635 6628 SNRPB 0.6229986 2.951031e-02 0.6490339 4.727499e-02
636 1059 CENPB 0.6440433 3.224106e-02 1.1292443 6.104656e-06
637 57506 MAVS 0.7851711 8.503194e-04 0.9669530 3.026305e-05
638 9770 RASSF2 0.7663205 1.857666e-03 0.7638085 5.221604e-03
639 29058 TMEM230 -0.6237705 4.614585e-03 -0.5474712 1.957347e-02
640 3642 INSM1 -1.9971556 2.843388e-05 -1.3653666 4.385264e-03
641 57186 RALGAPA2 1.0177461 4.768293e-03 1.1086594 1.949396e-03
642 55857 KIZ -3.2996821 9.426565e-04 -2.1092718 4.156827e-02
643 1471 CST3 1.2086087 3.743043e-26 1.3795657 6.923384e-27
644 57136 APMAP -0.6698550 1.185051e-03 -0.6812122 3.405905e-04
645 3397 ID1 2.3793819 8.169067e-03 2.7721296 1.739855e-04
646 9777 TM9SF4 -1.0423368 6.192435e-04 -1.4224951 4.563305e-05
647 84557 MAP1LC3A 1.2840689 2.218219e-06 0.8092823 2.802262e-02
648 58476 TP53INP2 -0.4969754 1.994346e-02 -0.8549743 6.390434e-04
649 26133 TRPC4AP 0.7202326 1.177626e-02 0.7369166 1.788840e-02
650 6185 RPN2 -0.4945573 5.046971e-03 -0.4877500 8.318659e-03
651 7150 TOP1 -0.4546108 8.589350e-04 -0.3807670 4.001298e-02
652 6385 SDC4 2.5582812 6.366611e-04 2.9459696 1.633773e-05
653 441951 ZFAS1 -0.7642309 3.630008e-05 -0.7115971 3.738045e-04
654 8813 DPM1 -1.3770244 5.706391e-03 -1.4120845 1.354480e-03
655 56937 PMEPA1 -1.3599819 1.250678e-09 -1.2412038 5.233679e-08
656 2778 GNAS -0.9110424 1.234079e-14 -0.9728377 2.043935e-15
657 3785 KCNQ2 -1.7287824 1.964923e-08 -2.0442425 8.570030e-05
658 1917 EEF1A2 -1.3102373 2.013576e-02 -1.5331370 5.277554e-04
659 79144 PPDPF -1.0455302 1.803030e-13 -1.0205089 1.391263e-11
660 6919 TCEA2 0.9970302 5.239650e-05 0.8564869 2.124406e-03
661 8204 NRIP1 1.5223402 9.760142e-13 1.2837321 2.001283e-07
662 54148 MRPL39 -1.2028438 1.651603e-02 -2.0249979 6.498955e-04
663 58494 JAM2 -1.2866157 3.075335e-05 -0.8060453 4.242012e-03
664 522 ATP5PF -1.1047917 2.067912e-16 -1.1207650 5.651067e-13
665 6647 SOD1 -0.6594340 9.505580e-06 -0.6400151 5.498998e-04
666 6651 SON -0.7380577 3.450537e-06 -0.8452348 3.347372e-06
667 539 ATP5PO -0.5510416 1.963777e-03 -0.5600488 3.519114e-03
668 64968 MRPS6 -0.6109838 2.284218e-03 -0.9016343 1.234605e-03
669 53820 RIPPLY3 1.8166987 4.033829e-02 2.3301752 2.972342e-03
670 51227 PIGP -1.0641385 7.642985e-04 -1.5559019 1.207749e-07
671 7485 GET1 -0.9670272 4.485725e-04 -1.0659018 7.469235e-05
672 4599 MX1 6.5129709 1.178985e-13 4.7489981 2.307235e-05
673 7307 U2AF1 -0.8753253 5.273963e-05 -0.5883751 1.725673e-02
674 150094 SIK1 -1.9449168 3.276602e-03 -2.5082565 8.172906e-06
675 754 PTTG1IP -0.7092259 4.694368e-05 -0.5106379 6.385133e-03
676 3275 PRMT2 0.9041244 1.447797e-13 0.7347605 1.607238e-06
677 23783 <NA> -2.0535666 1.064511e-04 -1.2660121 3.922499e-02
678 7122 CLDN5 7.6882552 1.507084e-06 7.5326013 5.245494e-06
679 9127 P2RX6 -6.9475368 1.411991e-03 -4.9495952 2.596403e-02
680 100037417 DDTL;DDT -4.1173363 4.351619e-02 -5.8350477 9.564642e-03
681 2130 EWSR1 -0.5497894 5.182409e-03 -0.6566420 2.652517e-03
682 8508 NIPSNAP1 2.0854791 1.117597e-17 1.7238679 1.342090e-09
683 253143 PRR14L 1.0061306 4.480015e-03 0.9355993 3.828982e-03
684 25828 TXN2 0.6309463 2.122480e-02 0.7837703 1.951939e-03
685 11078 TRIOBP -0.6890856 3.496671e-03 -0.8728017 8.957536e-05
686 3005 H1-0 0.9685330 9.727316e-04 1.3186945 1.014313e-05
687 25776 CBY1 0.8826165 1.291386e-02 1.2055688 5.978783e-05
688 4248 MGAT3 -1.3154641 1.331357e-03 -1.2968380 6.327923e-05
689 23112 TNRC6B 1.0234879 3.449208e-11 0.5734776 1.811473e-03
690 158 ADSL -0.8027735 1.002866e-02 -1.1597245 2.212467e-02
691 9978 RBX1 -0.7886203 4.661877e-04 -0.9068143 7.699282e-05
692 84844 PHF5A 1.1243600 3.196933e-04 1.4350685 3.087719e-05
693 27351 DESI1 0.9025063 2.198545e-02 1.0721990 2.589638e-03
694 706 TSPO 0.6557583 1.507084e-06 0.8486172 9.770725e-09
695 25814 ATXN10 -1.0353980 5.925748e-07 -0.6120749 3.824003e-02
696 10752 CHL1 -0.7303162 1.437404e-03 -0.9860194 6.170626e-05
697 57633 LRRN1 -1.8102115 2.183632e-06 -1.6260312 8.324859e-05
698 285362 SUMF1 -1.1070522 6.235271e-04 -1.0245436 1.631457e-03
699 7428 VHL 0.7257850 1.331805e-03 0.7279293 1.220129e-02
700 9797 TATDN2 1.0210697 3.189095e-02 1.2458058 6.390434e-04
701 6396 SEC13 -0.9579656 1.118838e-04 -0.6826414 9.044477e-03
702 5894 RAF1 -2.5997129 5.820569e-04 -2.0650136 2.646130e-02
703 6161 RPL32 -1.0806325 1.189903e-21 -0.7779658 1.989364e-08
704 2199 FBLN2 -0.8459523 7.881683e-04 -1.8077219 5.456321e-12
705 27258 LSM3 -1.0773957 1.068273e-06 -1.2368542 4.450604e-06
706 85403 EAF1 -1.4396254 4.664552e-04 -1.5459180 8.797670e-05
707 79750 ZNF385D 0.6342609 1.659110e-02 0.8168542 9.222303e-03
708 6138 RPL15 -0.7440450 9.290416e-12 -0.5354022 6.923078e-05
709 9045 RPL14 -0.9462397 2.834396e-11 -0.7659313 4.106732e-05
710 55129 ANO10 1.5672085 1.557156e-13 1.3859490 7.572853e-11
711 29072 SETD2 1.3332470 1.014466e-05 0.7952285 4.783854e-02
712 4134 MAP4 0.5866986 4.791304e-06 0.4371847 9.399814e-03
713 51372 TMA7 -0.4770049 3.816624e-02 -0.5930463 1.973507e-02
714 51447 IP6K2 0.6464313 1.626257e-03 0.8612760 4.869847e-05
715 64925 CCDC71 2.0654711 2.039079e-06 1.4270082 1.763911e-02
716 2771 GNAI2 0.4746637 1.711679e-04 0.4473208 8.711497e-03
717 9136 RRP9 1.6565451 3.603634e-02 1.5104446 3.241008e-02
718 80335 WDR82 0.4126762 3.940433e-02 0.4139527 4.612069e-02
719 28972 SPCS1 -1.2087611 1.507814e-05 -1.2522218 8.015505e-07
720 378 ARF4 -0.9342524 1.236015e-06 -0.7150834 1.177618e-03
721 5162 PDHB -0.8891871 6.426932e-04 -0.7744018 1.104922e-02
722 11170 FAM107A 0.7108706 4.564289e-07 0.8438374 8.634626e-09
723 9861 PSMD6 -1.2606773 5.977117e-06 -0.6906518 4.287478e-02
724 26018 LRIG1 -0.9823859 7.287051e-04 -1.2744312 3.203586e-03
725 131544 CRYBG3 1.9507274 4.119533e-02 1.8024199 3.578587e-02
726 1295 COL8A1 -2.0689175 2.320414e-02 -1.8307207 2.177510e-02
727 11259 FILIP1L -2.4415449 1.312190e-04 -5.6215128 3.702945e-09
728 55076 TMEM45A -3.5116566 5.214947e-03 -2.7410971 1.990733e-02
729 6152 RPL24 -0.4054846 4.885973e-03 -0.6037308 6.751807e-05
730 29114 TAGLN3 -1.5838847 6.961514e-03 -1.5878162 5.935201e-03
731 151887 CCDC80 0.6139833 2.964007e-05 0.5925393 9.847078e-04
732 10063 COX17 -0.6716026 4.485725e-04 -0.8393306 3.859001e-05
733 26355 FAM162A -1.9328801 3.288821e-22 -1.7126726 3.905016e-14
734 11343 MGLL 0.7436404 1.131736e-05 0.7988030 1.146903e-05
735 7555 CNBP -0.8977343 1.331197e-03 -0.8510197 6.236408e-03
736 8971 H1-10 1.0615496 4.847150e-07 0.8496144 6.375121e-04
737 6259 RYK -1.2134416 2.967454e-04 -1.1672802 3.665206e-03
738 80723 SLC35G2 -1.5472348 3.091483e-02 -2.4589313 2.526053e-02
739 92370 PXYLP1 -1.3140189 1.802276e-06 -0.8730358 4.523637e-03
740 9616 RNF7 -0.9841381 2.569506e-04 -1.3306717 1.375191e-08
741 483 ATP1B3 -0.7743038 6.459443e-03 -0.6626562 1.455525e-02
742 7029 TFDP2 0.8795826 1.665012e-02 0.8136336 2.710507e-02
743 9435 CHST2 -1.5760160 8.357784e-14 -1.8614337 1.631966e-22
744 4071 TM4SF1 -1.1531120 3.455921e-09 -1.0081670 5.173272e-07
745 51714 SELENOT -0.8079909 5.387139e-03 -0.6699049 1.260624e-02
746 6747 SSR3 -0.6347800 7.354903e-05 -0.7382513 1.225101e-06
747 25976 TIPARP 1.5621679 9.482389e-10 1.1563866 8.934141e-05
748 647033 <NA> 1.7055114 4.981620e-04 1.2759773 3.241008e-02
749 5806 PTX3 -1.4627165 2.682797e-03 -1.5351188 1.338800e-02
750 101243545 LINC02067 -2.2377320 7.530395e-16 -2.6491210 9.324337e-19
751 7095 SEC62 -0.7644827 9.650125e-07 -0.8421825 2.019764e-05
752 64778 FNDC3B -0.6494456 2.697896e-03 -0.7890739 1.471416e-03
753 79718 TBL1XR1 -0.7898441 3.911199e-02 -0.9407823 2.369589e-02
754 5708 PSMD2 0.6876897 6.829008e-05 0.6827839 4.826572e-03
755 64110 MAGEF1 -0.8812192 1.065961e-03 -0.8000918 2.555704e-03
756 10644 IGF2BP2 -2.0440181 8.734998e-18 -1.3019720 3.763786e-10
757 1427 CRYGS 1.1346179 6.562101e-05 1.1290719 9.777012e-05
758 2257 FGF12 1.9695269 7.906879e-05 2.0731871 2.039721e-05
759 3280 HES1 1.0207300 1.426437e-12 0.8214888 1.819415e-06
760 7037 TFRC -1.0562573 6.582075e-06 -1.8171049 5.988919e-13
761 1739 DLG1 0.8252637 6.755750e-03 1.0083381 2.819406e-04
762 6165 RPL35A -1.2229325 3.109067e-21 -1.0521400 6.485709e-13
763 521 ATP5ME -0.5657451 9.194512e-03 -0.5906954 7.838853e-03
764 2580 GAK 0.7219336 1.932494e-02 1.1020172 7.888319e-04
765 677770 SCARNA22 3.5414447 7.851343e-25 2.8760672 2.264676e-09
766 401115 C4orf48 1.0552561 6.545003e-06 1.2896350 1.768084e-05
767 6002 RGS12 1.4972339 6.355291e-08 1.1265948 4.678387e-04
768 114932 MRFAP1L1 1.1541753 9.482389e-10 1.0281080 3.491332e-06
769 23216 TBC1D1 1.1957982 8.857641e-03 1.1346256 1.299349e-02
770 6133 RPL9 -0.6058636 6.173691e-04 -0.5020652 2.212467e-02
771 81608 FIP1L1 -0.7217850 1.221124e-02 -1.2176557 1.226960e-05
772 5156 PDGFRA -1.5433008 1.666080e-09 -1.8040781 1.141369e-08
773 10606 PAICS -1.0034386 3.063956e-07 -0.6216646 1.456774e-02
774 258 AMBN -4.9822265 5.424457e-05 -8.2440786 4.237262e-07
775 25849 PARM1 -0.6114839 6.769801e-05 -1.3062333 3.326232e-14
776 10983 CCNI -0.6852370 1.904664e-10 -0.7118929 9.927156e-09
777 79966 SCD5 -0.5322866 1.230578e-04 -0.4569213 1.393295e-02
778 55008 HERC6 7.0791383 1.617490e-03 6.9486918 1.031036e-02
779 10611 PDLIM5 -0.6004503 1.507507e-02 -0.8534016 2.701263e-03
780 4547 MTTP -1.0906775 8.703424e-04 -0.8974650 3.606036e-03
781 4126 MANBA -0.7938814 3.283708e-02 -1.1831670 1.252402e-02
782 6164 RPL34 -1.3921977 1.380424e-30 -1.0834473 1.567134e-13
783 58505 OSTC -0.9279881 2.719888e-04 -1.2227041 1.725856e-05
784 79071 ELOVL6 0.9133948 8.399396e-03 0.8068751 3.241008e-02
785 401152 C4orf3 -1.1621356 2.825190e-09 -1.0761642 1.658403e-07
786 84162 KIAA1109 1.2338722 6.525714e-04 0.9774658 1.284853e-02
787 10252 SPRY1 1.0429198 4.128846e-12 0.9471736 2.323167e-09
788 27152 INTU 1.5599481 2.136125e-04 1.1175906 2.638770e-02
789 4717 NDUFC1 -0.7003878 9.709341e-04 -0.9021109 3.350605e-04
790 80155 NAA15 1.0183967 3.245051e-02 1.0491582 4.289425e-02
791 80854 SETD7 -0.8750229 1.105187e-02 -1.2817550 3.224189e-05
792 166614 DCLK2 0.6223728 6.876281e-04 0.5551005 7.506696e-03
793 2891 GRIA2 0.8500305 1.527124e-05 0.9016344 6.278649e-05
794 6307 MSMO1 -1.2139955 5.296230e-12 -0.7628001 1.877933e-03
795 1363 CPE 1.0386038 1.014466e-05 0.7934292 3.263649e-03
796 9848 MFAP3L -1.1793134 4.523090e-03 -1.1844621 8.500805e-03
797 291 SLC25A4 -1.2086954 4.107730e-05 -1.1735844 1.543444e-05
798 57587 CFAP97 0.7427990 3.950588e-02 0.8450499 1.622654e-02
799 170690 ADAMTS16 1.6694763 6.490208e-03 2.9001721 5.193755e-09
800 23379 ICE1 1.3175326 5.299632e-04 1.0401525 3.350546e-03
801 100505738 MIR4458HG -0.5665383 1.955629e-02 -0.5996206 2.801906e-02
802 1008 CDH10 -1.9505643 3.648729e-08 -1.5448575 1.119362e-04
803 6507 SLC1A3 1.1820878 5.080933e-34 0.8743854 3.552351e-12
804 25836 NIPBL 1.0564073 3.948792e-04 0.9007622 2.177334e-03
805 9180 OSMR -1.9049220 2.152111e-11 -1.4239353 1.686645e-08
806 100506548 <NA> -0.9277715 1.114229e-02 -1.3409343 1.334918e-02
807 3157 HMGCS1 -1.1957172 2.234338e-17 -0.6594077 6.647511e-05
808 10605 PAIP1 -0.8532986 2.411425e-02 -1.0505537 1.124294e-02
809 4338 MOCS2 -0.7519809 3.491021e-02 -0.7886653 2.489895e-02
810 4724 NDUFS4 -0.6970723 6.520793e-04 -0.7350511 2.682046e-03
811 285671 RNF180 1.0996899 2.037693e-05 1.0858061 6.668474e-05
812 285672 SREK1IP1 -0.9533591 1.662485e-04 -0.9230718 1.718029e-02
813 5295 PIK3R1 0.9872231 7.594497e-04 0.9326371 2.879230e-03
814 55814 BDP1 1.5183886 2.224452e-02 1.5600369 2.883354e-02
815 3842 TNPO1 1.2807072 5.574322e-14 0.9004943 1.779785e-05
816 3074 HEXB -1.3173354 9.564588e-04 -0.6483662 4.378514e-02
817 3156 HMGCR -1.0292358 3.587951e-07 -0.9936596 6.977203e-05
818 10087 CERT1 1.4438973 9.558491e-16 1.0091891 5.797352e-06
819 6902 TBCA -0.4512944 9.215731e-04 -0.5152711 2.836696e-03
820 202333 CMYA5 4.8864830 7.520028e-05 4.7441964 1.871909e-04
821 256987 SERINC5 -0.6496750 1.073260e-03 -1.1003432 4.929951e-08
822 100462981 MTRNR2L12 2.3607249 1.166815e-166 2.0421868 2.970385e-27
823 5924 RASGRF2 0.7056930 9.553566e-03 0.9703039 5.552133e-05
824 6228 RPS23 -0.8739794 9.387213e-18 -0.7366332 9.357521e-08
825 1462 VCAN 2.1957441 1.976976e-34 1.7224678 2.043935e-15
826 5921 RASA1 2.5061754 1.390634e-04 3.1646447 9.526825e-10
827 1070 CETN3 0.5774513 6.476386e-03 0.4861433 3.987163e-02
828 345757 FAM174A -1.6038075 3.055840e-02 -2.0149687 4.405284e-03
829 2241 FER 2.6067226 1.337098e-10 2.3302244 6.075290e-07
830 114915 <NA> -1.6808780 2.080361e-05 -0.9968276 2.213953e-02
831 324 APC 1.1150494 1.820257e-14 0.6892972 1.134053e-04
832 6728 SRP19 -1.0821472 1.024343e-04 -0.8080873 2.669405e-02
833 7905 REEP5 -0.6127878 2.211377e-03 -0.8389722 1.262360e-04
834 51014 TMED7 -1.0144508 1.223899e-03 -0.8793472 9.861464e-03
835 57556 SEMA6A -0.7987993 5.679116e-09 -0.8176305 3.724102e-06
836 153443 SRFBP1 2.1337669 7.873206e-06 1.5922350 8.858092e-03
837 57507 ZNF608 1.5250971 2.519027e-07 1.0416429 2.934910e-03
838 3094 HINT1 -0.9455195 5.671458e-14 -0.8364669 6.174445e-10
839 56951 C5orf15 -1.2535332 4.102697e-09 -1.0819668 5.070085e-07
840 6500 SKP1 -0.6264521 4.485725e-04 -0.5715314 1.346036e-03
841 23338 JADE2 1.7238473 4.448263e-02 1.9515703 7.862124e-03
842 9879 DDX46 -0.9901795 4.951180e-04 -1.0515823 7.445233e-05
843 9555 MACROH2A1 -0.5119695 1.561228e-02 -0.4970320 3.290530e-02
844 7045 TGFBI -1.7922980 4.119533e-02 -2.2067195 3.033021e-04
845 6695 SPOCK1 -0.4388064 2.561841e-02 -0.8318303 6.363547e-06
846 995 CDC25C 4.3163710 2.345874e-02 4.1862802 3.248617e-02
847 56662 VTRNA1-3 6.4490102 3.630008e-05 5.7757103 2.021229e-03
848 26167 PCDHB5 -0.6978699 1.013941e-02 -0.9148200 3.617950e-03
849 56106 PCDHGA10 2.5049006 8.315254e-03 2.4517359 2.358497e-02
850 1729 DIAPH1 0.8882793 5.145461e-03 0.9129698 3.249669e-02
851 10007 GNPDA1 0.9408571 5.054616e-09 0.7923198 4.220126e-05
852 815 CAMK2A 1.6792011 2.479306e-03 1.8405059 3.560700e-04
853 972 CD74 2.1906950 3.072670e-11 1.7478005 5.877562e-05
854 3340 NDST1 0.5832947 8.169067e-03 0.6146485 6.541794e-03
855 309 ANXA6 0.8098381 7.506017e-06 0.5304214 2.943702e-02
856 57451 TENM2 -1.1494814 3.857776e-10 -1.2878565 3.717116e-13
857 8992 ATP6V0E1 -0.6906722 1.282110e-04 -0.4490359 2.313609e-02
858 662 BNIP1 1.2812599 8.169067e-03 1.0978521 2.089476e-02
859 1212 CLTB 0.8729299 3.053562e-03 0.9015231 3.307371e-03
860 64324 NSD1 1.7515088 2.505110e-14 1.2402096 6.363547e-06
861 80758 PRR7 0.6315226 4.665216e-02 0.9166882 1.222449e-03
862 54732 TMED9 0.5645326 7.783222e-06 0.4013908 2.188471e-02
863 91522 COL23A1 2.3112797 2.544098e-11 2.1266171 5.256011e-09
864 55819 RNF130 -1.0077006 1.605162e-02 -1.7980915 5.245494e-06
865 10399 RACK1 -0.5254117 6.385736e-05 -0.3793132 1.650399e-02
866 56897 WRNIP1 -1.5471129 3.628200e-05 -1.7437219 1.607238e-06
867 1992 SERPINB1 4.9146404 2.926616e-04 5.2335655 2.204739e-05
868 7280 TUBB2A -0.7089601 6.267252e-07 -0.6708941 1.635422e-05
869 347733 TUBB2B -0.7067559 1.888823e-07 -0.4985577 4.462794e-03
870 81853 TMEM14B -0.9179555 2.742442e-02 -0.9018269 7.723997e-03
871 54898 ELOVL2 0.9428945 7.828128e-12 0.6285030 4.018058e-04
872 51406 NOL7 -1.0009519 4.066968e-04 -0.8148895 2.946177e-03
873 3400 ID4 -0.9120412 2.154370e-03 -1.1700889 2.372486e-04
874 51473 DCDC2 1.2443843 7.564192e-03 1.3195920 9.513596e-03
875 55856 ACOT13 -0.9022622 2.324914e-02 -1.4228438 2.518596e-07
876 9750 RIPOR2 -5.8034728 1.393580e-02 -6.5726093 7.286063e-03
877 10590 SCGN -1.3158239 2.928310e-03 -2.8434388 3.120874e-08
878 11120 BTN2A1 -1.4164333 2.808964e-04 -1.8137714 2.571776e-05
879 3137 <NA> 1.0127683 1.204615e-04 0.9826226 4.770550e-04
880 3107 HLA-C 1.2931591 7.351098e-18 1.0994190 6.540541e-11
881 7922 SLC39A7 1.4996572 1.875790e-07 1.1195832 6.582692e-03
882 57699 CPNE5 1.3108553 2.016435e-20 1.3795464 8.625945e-17
883 2739 GLO1 0.5911196 1.437404e-03 0.4748510 3.290530e-02
884 285855 RPL7L1 -0.5822925 1.867945e-02 -0.5495086 3.792659e-02
885 116138 KLHDC3 0.7039657 9.291893e-04 1.0077716 3.176501e-06
886 10591 DNPH1 0.5807379 8.537075e-03 0.8159909 6.468157e-05
887 25844 YIPF3 0.5531543 3.105704e-02 0.7077815 5.262326e-03
888 7422 VEGFA -2.3750017 9.132820e-12 -2.1017346 1.970862e-11
889 9697 TRAM2 -1.1687443 8.035919e-05 -0.7628945 9.987876e-03
890 667 DST 1.3066882 3.953791e-17 1.3636002 5.344141e-15
891 55788 LMBRD1 -1.0727743 1.004953e-04 -1.0754706 1.631457e-03
892 79627 OGFRL1 1.1393962 1.275092e-07 0.7481483 6.820254e-03
893 79940 LINC00472 1.9153970 9.865105e-04 2.1763861 3.356981e-04
894 1915 EEF1A1 -0.8015550 5.133405e-09 -0.6658325 1.563890e-03
895 9324 HMGN3 -0.5533836 1.318382e-02 -1.0783521 3.858040e-07
896 7162 TPBG -1.0432333 9.671356e-03 -1.6963659 5.982751e-09
897 387066 <NA> -1.3933526 7.309480e-12 -1.0545530 3.066793e-05
898 26799 <NA> 4.4480807 3.645600e-10 3.3109946 2.124406e-03
899 154313 CFAP206 -6.4873816 1.231225e-02 -6.2947208 1.916278e-02
900 1268 CNR1 0.6703656 2.747521e-02 0.7638065 1.993997e-02
901 10690 FUT9 1.6760412 5.693455e-08 1.0993674 2.726542e-03
902 25957 PNISR -0.6338491 2.610510e-04 -0.5849137 4.378086e-03
903 10973 ASCC3 1.3190847 1.630044e-04 0.8665382 1.962129e-02
904 5550 PREP -1.5078120 1.985515e-02 -1.3018358 1.763911e-02
905 100133941 CD24 -7.8393997 8.144601e-06 -5.7783913 5.659621e-04
906 8763 CD164 -0.9132365 2.242736e-04 -1.0199432 3.970221e-05
907 10758 TRAF3IP2 0.6105654 3.831231e-02 0.7831279 2.244861e-02
908 2534 FYN -0.4887334 1.857858e-03 -0.6654093 3.210971e-05
909 3910 LAMA4 0.9816690 1.796029e-02 1.3614822 2.822612e-06
910 7259 TSPYL1 -1.0066554 2.030167e-05 -0.8455353 1.966518e-03
911 25842 ASF1A -1.0844678 1.108836e-02 -1.6052613 7.714980e-05
912 57515 SERINC1 -0.6740563 4.823290e-04 -0.7793831 2.108357e-05
913 2173 FABP7 -1.5750241 1.992080e-07 -1.6703756 3.164172e-06
914 7164 TPD52L1 -1.0054001 1.661418e-02 -1.1429760 5.809267e-03
915 51020 HDDC2 -0.6481942 1.011438e-02 -0.6375036 4.250877e-02
916 23493 HEY2 -3.8037514 5.106707e-03 -3.2360874 2.559698e-02
917 93663 ARHGAP18 0.8694748 1.256783e-02 0.8629630 1.494738e-02
918 1490 CCN2 -3.1554085 3.418038e-16 -4.2814332 1.976875e-28
919 6206 RPS12 -1.1491929 1.332521e-13 -0.7474210 7.475852e-05
920 7430 EZR 0.9648313 8.092332e-03 0.9838192 1.228025e-02
921 3482 IGF2R -0.5881522 6.885354e-03 -0.8972238 1.596718e-03
922 6196 RPS6KA2 0.3810064 3.340861e-02 0.7111698 7.352437e-05
923 101929523 <NA> 3.6390108 1.793939e-02 4.4095356 8.844827e-03
924 5689 PSMB1 -1.3603850 3.785619e-09 -1.3308572 6.759562e-11
925 23353 SUN1 -0.9938442 3.021516e-02 -0.9336274 4.249711e-02
926 29886 SNX8 0.5523896 2.770815e-02 0.7926513 1.291166e-03
927 6624 FSCN1 -1.0713334 9.331388e-21 -0.8325498 9.905933e-07
928 4697 NDUFA4 -0.6857134 4.144021e-08 -0.5453823 5.154806e-03
929 8701 DNAH11 2.6990566 7.281788e-04 2.4321317 7.862124e-03
930 10643 IGF2BP3 0.8694833 3.339777e-03 1.0047908 6.621297e-05
931 4852 NPY -2.5012943 4.831459e-03 -4.1586786 6.690797e-06
932 11335 CBX3 0.5547655 6.426932e-04 0.4707137 2.361762e-02
933 9865 TRIL -0.6869435 1.858073e-08 -0.6966268 4.731214e-06
934 54504 CPVL -0.9011170 9.629234e-03 -1.1181150 2.631022e-03
935 23658 LSM5 -0.9948036 5.824816e-04 -1.3722624 5.060173e-05
936 23333 DPY19L1 -2.0667730 4.509647e-07 -1.4044842 3.706986e-03
937 83930 STARD3NL -1.3486632 6.786786e-06 -0.7768161 2.883354e-02
938 2737 GLI3 0.9593370 2.258820e-02 0.9499567 3.450329e-02
939 5683 PSMA2 -1.1084040 7.267595e-08 -1.2717362 2.899835e-08
940 23386 NUDCD3 0.7902751 2.744749e-03 0.9103817 1.743062e-03
941 4967 OGDH 1.2931752 1.419791e-05 0.8833557 2.269406e-02
942 83605 CCM2 0.8886044 1.070896e-02 0.9034776 1.313529e-02
943 3486 IGFBP3 -2.2835324 1.617490e-03 -3.0909521 8.105970e-04
944 23480 SEC61G -1.7027271 1.904718e-04 -1.7535890 7.572853e-11
945 51142 CHCHD2 0.5992320 3.754695e-07 0.7204071 3.903492e-07
946 51119 SBDS -1.1144088 2.370369e-03 -1.9388407 7.013897e-09
947 2969 GTF2I 1.8731674 3.857776e-10 1.0795996 7.489466e-03
948 5447 POR 0.9526933 8.936179e-04 0.7024575 2.395150e-02
949 100505854 APTR -1.6188872 4.195847e-02 -3.1489749 4.284771e-05
950 222194 RSBN1L -1.0346992 2.210049e-02 -1.4653252 2.226998e-02
951 6717 SRI -0.7315985 5.258914e-08 -0.7194124 1.258595e-05
952 1595 CYP51A1 -0.7679455 5.040980e-05 -0.6896136 1.344835e-03
953 54467 ANKIB1 0.8988626 6.611509e-03 0.8496291 1.589383e-02
954 1278 COL1A2 -1.8044267 2.774879e-17 -2.6985465 2.861990e-51
955 8910 SGCE -1.1595413 1.284825e-03 -1.3681407 7.241593e-05
956 23089 PEG10 1.2813683 2.586992e-39 0.3492587 1.840892e-02
957 7979 SEM1 -1.0150038 3.106345e-10 -1.0726954 9.106799e-07
958 1749 DLX5 -1.8207920 5.149658e-03 -9.1287072 6.304782e-10
959 10552 ARPC1A -1.5581714 1.434742e-06 -1.6037044 1.099400e-08
960 9551 ATP5MF -0.9977073 3.079767e-11 -0.9526787 1.718309e-07
961 7205 TRIP6 0.5688503 1.582610e-03 0.8974429 9.949827e-07
962 51024 FIS1 0.7062099 2.013576e-02 0.9859722 1.949396e-03
963 3912 LAMB1 -1.6430576 1.897841e-07 -0.7314453 3.937134e-02
964 401397 SMIM30 -1.1314569 3.784807e-03 -0.9816881 1.332957e-02
965 5803 PTPRZ1 1.1501936 1.405664e-13 0.9114296 4.464529e-06
966 2861 GPR37 -1.2991912 3.622386e-06 -1.1075672 1.968864e-08
967 64101 LRRC4 2.0242364 2.548109e-10 1.1252903 2.194272e-02
968 813 CALU -0.7317704 2.585546e-06 -0.6162849 4.690217e-04
969 64753 CCDC136 1.5435761 8.370267e-03 1.4607115 1.455525e-02
970 800 CALD1 -0.9200718 6.811766e-16 -0.9520463 4.679431e-14
971 5764 PTN -0.6769084 8.651978e-15 -0.7300167 2.117855e-10
972 51650 MRPS33 -0.8949939 1.758600e-03 -1.0986688 5.699268e-04
973 26047 CNTNAP2 1.7279244 1.659437e-06 1.4111858 4.493978e-04
974 155066 ATP6V0E2 0.9072461 4.474290e-06 0.7210372 1.950412e-03
975 10922 FASTK 0.7641096 1.787291e-02 0.9882660 6.245499e-04
976 6009 RHEB -0.6150900 2.937988e-03 -1.1056928 7.864066e-08
977 3638 INSIG1 -1.4066187 7.994214e-06 -1.1205356 7.557718e-04
978 79660 PPP1R3B -1.1395033 8.720267e-05 -0.5734475 1.002911e-02
979 2222 FDFT1 -0.7811303 3.323915e-05 -0.5716171 9.705714e-03
980 4023 LPL -1.3515968 8.879865e-16 -1.9391683 4.328081e-30
981 64236 PDLIM2 0.4546158 2.540171e-02 0.4772553 3.141640e-02
982 4747 NEFL -1.4123499 4.378022e-14 -1.9807071 1.008475e-18
983 665 BNIP3L -1.4808545 4.451508e-21 -1.7334117 3.282031e-26
984 148 ADRA1A 1.2223872 3.928010e-11 0.9041357 8.957536e-05
985 1191 CLU 0.5675630 9.570125e-08 0.3585553 1.221455e-02
986 2137 EXTL3 -0.6455879 7.664811e-07 -0.7128163 3.065415e-07
987 51669 SARAF -0.9491485 2.131984e-09 -0.8062936 1.698814e-05
988 692084 SNORD13 3.5705896 9.558819e-15 2.1180819 4.256124e-02
989 1052 CEBPD -0.9768778 7.951942e-03 -1.3845761 6.276546e-04
990 5591 PRKDC 0.9434372 1.262898e-04 0.8360927 2.249249e-03
991 641638 <NA> -1.3612659 4.237594e-12 -1.2802754 6.933904e-10
992 80243 PREX2 3.3335727 6.465637e-08 2.6851231 2.844817e-04
993 23213 SULF1 -1.9847588 7.581180e-08 -2.9676716 1.348690e-15
994 6129 RPL7 -0.8711653 1.847115e-05 -0.4738742 4.495736e-02
995 27067 STAU2 1.8946808 1.263630e-25 1.3784939 1.750258e-08
996 6921 ELOC -0.6828490 3.310423e-03 -1.1015451 2.091900e-07
997 54968 TMEM70 -1.4282037 1.505530e-04 -0.9587612 3.372442e-02
998 83690 CRISPLD1 -0.8076399 5.694319e-03 -0.6913367 3.266529e-02
999 401466 RBIS -0.6109441 2.282347e-02 -1.1290735 4.758366e-05
1000 169200 TMEM64 -0.8474270 3.615251e-02 -1.0096307 3.489786e-02
1001 7381 UQCRB -0.9882168 2.208474e-12 -0.8163836 3.724102e-06
1002 6383 SDC2 -1.2380643 6.236589e-09 -0.7518041 2.365134e-03
1003 10404 CPQ 1.4222343 9.653782e-06 1.1597588 4.165623e-03
1004 6156 RPL30 -0.7376391 1.013553e-10 -0.7108959 7.449348e-06
1005 1345 COX6C -0.5777179 5.474002e-05 -0.5623486 6.308417e-04
1006 5440 POLR2K -0.5044544 3.651419e-02 -0.7176969 7.967493e-03
1007 26986 PABPC1 1.0519808 6.214647e-14 0.9567969 3.060342e-06
1008 83988 NCALD 1.6890694 2.529989e-03 1.9039998 3.089672e-03
1009 50484 RRM2B 1.0343173 3.689493e-03 0.7836091 4.091967e-02
1010 51582 AZIN1 -0.5209480 4.680108e-02 -0.7501651 1.048791e-02
1011 3646 EIF3E -1.2967221 3.621184e-11 -1.7926835 4.692596e-19
1012 8667 EIF3H -0.8789134 2.277908e-07 -0.7488252 3.343362e-04
1013 5168 ENPP2 -1.3575140 8.795334e-06 -1.4292122 1.180285e-04
1014 28998 MRPL13 -0.7511183 2.502752e-02 -0.6989666 3.516548e-02
1015 157378 TMEM65 1.5657262 7.365527e-09 1.1724731 1.668589e-02
1016 6713 SQLE -0.8748781 3.452117e-03 -0.9996655 1.479596e-04
1017 4062 LY6H 2.0404450 3.587951e-07 1.8806104 2.075791e-05
1018 5339 PLEC 0.5398222 4.635828e-02 0.7203850 7.044621e-03
1019 2907 GRINA 0.9810473 6.768665e-15 0.9564501 3.474366e-11
1020 8733 GPAA1 0.8408190 1.166590e-03 0.7240015 1.626651e-02
1021 84232 MAF1 1.0458379 6.859720e-03 1.1526630 1.331088e-03
1022 65265 C8orf33 1.2926181 8.743396e-08 1.0108476 7.888319e-04
1023 23189 KANK1 0.7144651 7.895217e-03 0.8721550 8.060539e-03
1024 4781 NFIB 0.8854179 7.913729e-09 0.9374237 1.854601e-08
1025 6194 RPS6 -0.4156980 1.249709e-04 -0.4208246 4.123543e-03
1026 55958 KLHL9 -1.0329553 1.095726e-03 -0.7951135 2.601431e-02
1027 1029 CDKN2A 1.1355662 8.895738e-07 0.8740444 1.569308e-03
1028 4712 NDUFB6 -0.8906200 7.354903e-05 -1.0624409 3.517011e-05
1029 7415 VCP -0.4336137 4.673478e-03 -0.5964534 1.219025e-03
1030 26267 FBXO10 0.8254536 1.904018e-03 0.7197069 3.307371e-03
1031 347252 IGFBPL1 1.4842951 6.600389e-06 0.7910572 4.881864e-02
1032 23137 SMC5 0.8298430 4.069353e-02 1.0222075 1.277603e-02
1033 23670 CEMIP2 -0.7164397 4.263387e-04 -0.6846670 2.310065e-03
1034 301 ANXA1 -1.2241328 9.482389e-10 -1.0624764 5.946754e-06
1035 2619 GAS1 -1.6908161 1.035109e-02 -2.1164596 7.888319e-04
1036 53358 SHC3 0.9146960 9.227041e-05 0.7323579 8.160467e-03
1037 10952 SEC61B -0.6878836 6.312036e-06 -0.8087133 5.059413e-07
1038 19 ABCA1 1.3942846 1.002036e-16 0.8831721 2.555704e-03
1039 7295 TXN -0.9919738 2.440240e-09 -0.9387855 3.226627e-06
1040 3371 TNC -0.9351474 1.384094e-06 -1.0540684 4.937183e-07
1041 7099 TLR4 -1.4742665 2.792422e-05 -1.5358692 2.356454e-03
1042 11224 RPL35 -0.7140837 5.328804e-04 -0.5636122 2.370230e-02
1043 3309 HSPA5 -0.3475594 2.052126e-02 -0.6009229 4.952359e-04
1044 6136 RPL12 -0.7408997 2.799860e-05 -0.5080363 1.740316e-02
1045 2801 GOLGA2 0.9408435 8.648658e-07 0.8556162 7.906252e-05
1046 81605 URM1 0.5044218 3.868969e-02 0.6566555 2.631022e-03
1047 51490 SPOUT1 0.8880710 2.170030e-03 0.9268879 3.644784e-03
1048 23048 FNBP1 0.8670697 7.171580e-10 0.4497187 2.716982e-02
1049 23064 SETX 1.2082170 2.852961e-06 0.7463572 2.726974e-02
1050 10422 UBAC1 1.3077029 3.031354e-09 1.0328132 2.266897e-04
1051 8721 EDF1 -1.2571777 1.808271e-11 -1.1580497 5.795139e-07
1052 26012 NSMF 0.7596866 3.070386e-02 0.9300300 1.767130e-03
1053 64975 MRPL41 0.4664101 3.570204e-02 0.5562549 2.244861e-02
1054 4549 <NA> 1.6331191 3.857776e-10 1.3996902 1.510058e-02
1055 4577 <NA> 1.5219784 2.037433e-12 1.2943451 2.151536e-09
1056 4550 <NA> 1.0065265 6.789721e-36 0.8732613 2.053751e-02
1057 4535 MT-ND1 -0.9737351 1.140679e-32 -0.8329473 2.488051e-10
1058 4572 <NA> 1.3321582 2.895981e-17 1.5378374 5.611052e-18
1059 4569 <NA> 2.3208192 4.721665e-12 1.7409576 1.262360e-04
1060 4536 MT-ND2 0.5406437 8.802362e-07 0.6937946 1.795972e-04
1061 4570 <NA> 0.4694520 3.107778e-02 0.6724287 5.809267e-03
1062 4579 <NA> 0.9286471 4.908604e-08 0.9849385 1.159320e-07
1063 4513 MT-CO2 -0.9285861 7.322731e-11 -0.8475787 7.042812e-07
1064 4509 MT-ATP8 1.5395524 1.276534e-06 1.7137389 6.507206e-04
1065 4508 MT-ATP6 -0.8548935 5.067631e-20 -0.9766476 4.574961e-18
1066 4514 MT-CO3 -0.5001743 9.867495e-09 -0.6062588 3.065415e-07
1067 4539 MT-ND4L 0.4098798 7.860423e-03 0.7061768 4.445112e-03
1068 4538 MT-ND4 -0.9778092 2.474374e-28 -0.8874018 8.200323e-13
1069 4564 <NA> 2.8968952 2.657613e-09 2.1367216 5.236779e-04
1070 4540 MT-ND5 1.2589533 5.499656e-43 1.3017628 3.543200e-03
1071 4519 MT-CYB -0.8936775 1.210009e-18 -0.9136131 3.552351e-12
1072 4571 <NA> 2.8741663 3.964011e-60 2.7148072 6.076172e-47
1073 9189 ZBED1 0.8756657 1.626257e-03 0.6715765 4.443677e-02
1074 1183 CLCN4 1.1987875 7.477217e-04 1.0632694 3.777914e-02
1075 8905 AP1S2 -1.0197074 2.045773e-23 -0.7491404 7.673449e-07
1076 94056 SYAP1 0.6401036 3.195817e-03 0.5801140 1.362446e-02
1077 30011 SH3KBP1 0.5176608 2.284218e-03 0.4838495 1.217034e-02
1078 256643 BCLAF3 1.4383765 3.978266e-03 1.1011397 3.824003e-02
1079 10549 PRDX4 -0.9963902 6.235271e-04 -0.8931776 1.921452e-02
1080 10159 ATP6AP2 -0.6904247 3.074659e-03 -0.6512225 5.161862e-03
1081 54539 NDUFB11 -1.0911680 4.618734e-05 -0.6999433 1.573848e-02
1082 56850 GRIPAP1 1.1956108 8.144601e-06 1.1150998 3.350605e-04
1083 57477 SHROOM4 1.1972732 1.141820e-02 1.1100956 3.290530e-02
1084 9500 MAGED1 -0.5999945 1.024625e-06 -0.6306059 5.400590e-05
1085 54552 GNL3L 1.1412892 1.913864e-04 1.0420340 4.939225e-03
1086 550643 NBDY -0.5743538 7.468132e-03 -0.8051838 4.776423e-04
1087 442454 <NA> -0.5148396 1.333396e-02 -0.5696300 9.315952e-03
1088 9843 HEPH -1.1594244 7.268861e-03 -1.9647934 2.024163e-06
1089 1947 EFNB1 0.8758498 9.956070e-04 0.8150692 1.174269e-02
1090 340527 NHSL2 2.6372096 1.669647e-15 2.4046029 3.809264e-10
1091 392490 NHSL2 -1.2470683 3.225254e-03 -1.1971654 2.782563e-02
1092 5303 PIN4 -0.8235796 7.974793e-03 -1.2192356 1.284309e-05
1093 6191 RPS4X -0.6470555 9.270311e-09 -0.5282855 3.102246e-04
1094 1349 COX7B -0.6093206 5.357676e-04 -0.6052488 3.134155e-03
1095 5230 PGK1 -1.8136626 4.994565e-57 -1.0856060 1.184684e-15
1096 2846 LPAR4 1.5527621 2.624884e-11 1.3030488 2.365932e-07
1097 79366 HMGN5 -0.9809284 6.168525e-03 -0.9729236 1.118223e-02
1098 90843 TCEAL8 0.7450006 6.752355e-05 0.4504685 4.112077e-02
1099 340543 TCEAL5 1.3703889 1.571670e-14 1.2069287 7.371241e-08
1100 51186 TCEAL9 0.3238067 4.584347e-02 0.5488991 2.252696e-04
1101 7319 UBE2A -0.8935808 3.098489e-03 -0.8697075 8.308968e-03
1102 6170 RPL39 -0.4626669 1.100917e-02 -0.6139391 4.103190e-03
1103 4694 NDUFA1 -1.0161937 1.277991e-09 -0.8558358 2.205322e-05
1104 55026 TMEM255A -1.4802818 1.484960e-02 -0.9209734 4.378176e-02
1105 63035 BCORL1 1.2572034 2.282347e-02 1.2992716 2.420042e-02
1106 159090 PABIR2 -1.2347532 2.705447e-02 -1.0495457 4.915824e-02
1107 27316 RBMX 1.4104813 5.734145e-20 0.8287942 5.485502e-05
1108 1038 CDR1 4.1675101 2.089264e-70 3.9732007 7.747275e-33
1109 2334 AFF2 1.8926920 3.325676e-03 2.3471570 1.588911e-05
1110 1069 CETN2 0.4761454 3.698264e-02 0.5823596 4.184859e-02
1111 84968 PNMA6A 1.4555371 1.348505e-02 1.7028966 2.170595e-04
1112 2010 EMD 0.6855411 4.443434e-03 0.7763575 6.254644e-03
1113 537 ATP6AP1 -0.6037523 4.768119e-03 -0.6004954 1.650399e-02
1114 6192 RPS4Y1 -6.3220505 1.983734e-02 -6.1293881 2.943702e-02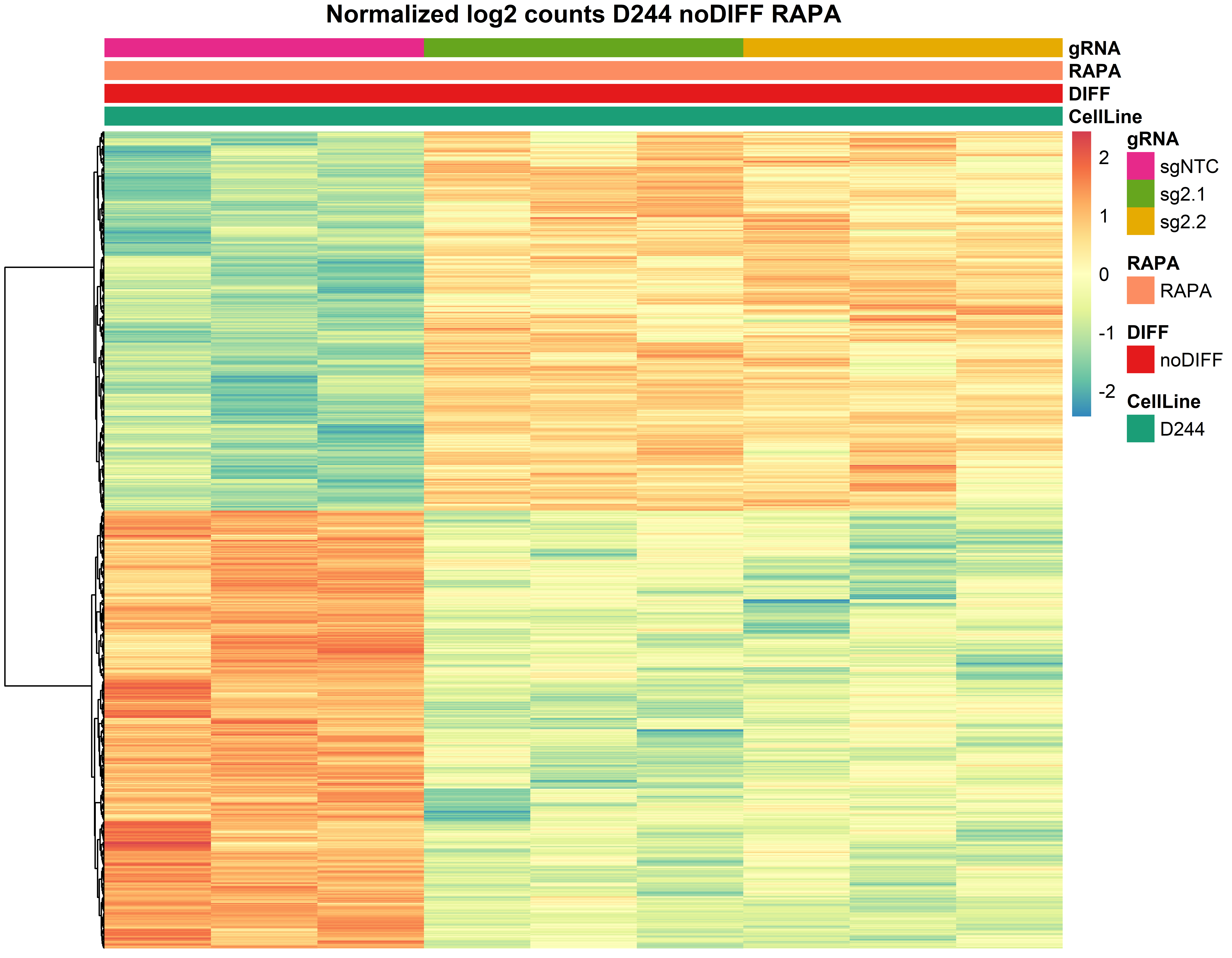
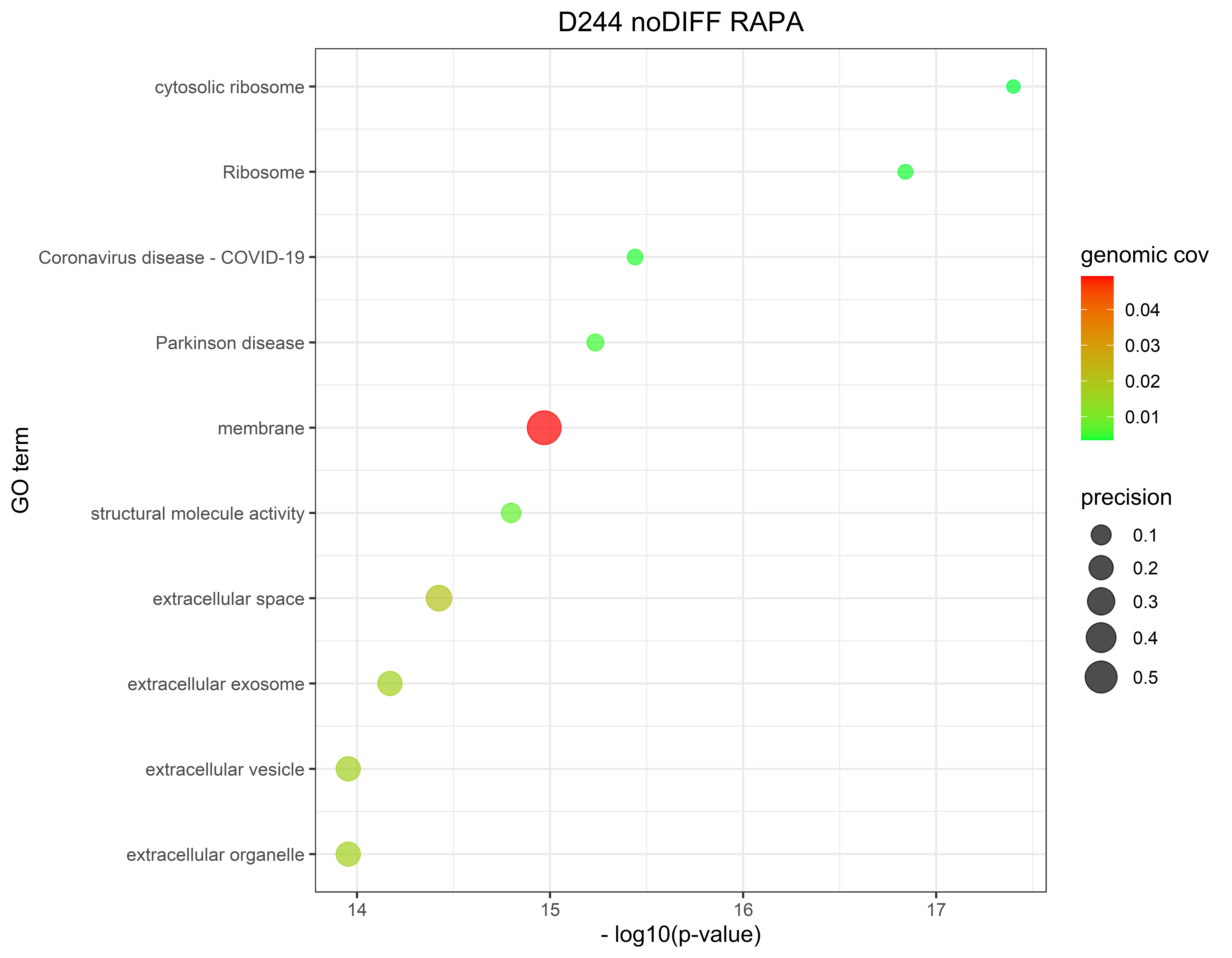
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.3368424 1.0028228 1.756475 3.802884e-02
02_TSC2_KO_Grabole_2016 1.3245406 1.1660937 1.503782 1.231542e-05
03_TSC2_Grabole-2016 1.3455116 1.1869565 1.525993 2.635266e-06
04_TSC2-tuber-Martin2017 2.1558355 1.2201462 3.612191 5.130354e-03
05_TSC2-SEN_SEGA-Martin2017 1.1765581 0.9891949 1.393715 5.959465e-02
06_all_EpilepsyGene 0.9611605 0.6354413 1.405705 9.243670e-01
07_high_conf_EpilepsyGene 0.7680892 0.3230734 1.570135 6.209493e-01
08_ASD_Epilepsy_EpilepsyGene 1.0534730 0.4895249 2.022132 8.645822e-01
09_Epi25_dataset 1.2247801 0.5083246 2.551856 5.419038e-01
10_SFARI_all 1.7074488 1.1248765 2.514414 8.321251e-03
11_SFARI_score4plus 1.8800000 1.1735809 2.899389 7.675800e-03
12_DeRubeis_RDNV 1.4810923 0.6477934 2.991091 2.926464e-01
13_Iossifov_RDNV 1.1774013 0.8500058 1.599312 2.837695e-01
14_FMRP_Darnell 1.6930496 1.3393722 2.121791 1.112072e-05
15_Voineagu_M12 0.7701221 0.4736596 1.193020 2.708067e-01
16_Voineagu_M16 2.7292144 2.0348510 3.616695 9.824988e-11
17_ID_genes_Neel 1.4083465 0.9615878 2.008817 6.450879e-02
18_ID_genes_pinto2014 1.1289230 0.6900336 1.762534 5.537765e-01
19_Fromer_SCZ_FunDN 1.2187890 0.8795088 1.656348 2.088175e-01
20_Cocchi_Pruning 1.5595925 0.7459834 2.956130 1.743610e-01
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.29031043 0.14667942 7.605768e-01
02_TSC2_KO_Grabole_2016 0.28106572 0.06500322 2.463085e-04
03_TSC2_Grabole-2016 0.29677435 0.06397035 5.270531e-05
04_TSC2-tuber-Martin2017 0.76817836 0.29041210 1.026071e-01
05_TSC2-SEN_SEGA-Martin2017 0.16259331 0.08849860 1.000000e+00
06_all_EpilepsyGene -0.03961384 0.21113355 1.000000e+00
07_high_conf_EpilepsyGene -0.26384934 0.44185022 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 0.05209231 0.39102668 1.000000e+00
09_Epi25_dataset 0.20276133 0.44867162 1.000000e+00
10_SFARI_all 0.53500035 0.21292200 1.664250e-01
11_SFARI_score4plus 0.63127177 0.24041432 1.535160e-01
12_DeRubeis_RDNV 0.39277985 0.42192008 1.000000e+00
13_Iossifov_RDNV 0.16330971 0.16623564 1.000000e+00
14_FMRP_Darnell 0.52653142 0.11955635 2.224144e-04
15_Voineagu_M12 -0.26120615 0.24798990 1.000000e+00
16_Voineagu_M16 1.00401382 0.14979144 1.964998e-09
17_ID_genes_Neel 0.34241635 0.19468660 1.000000e+00
18_ID_genes_pinto2014 0.12126405 0.25116280 1.000000e+00
19_Fromer_SCZ_FunDN 0.19785778 0.16645383 1.000000e+00
20_Cocchi_Pruning 0.44442455 0.37626348 1.000000e+00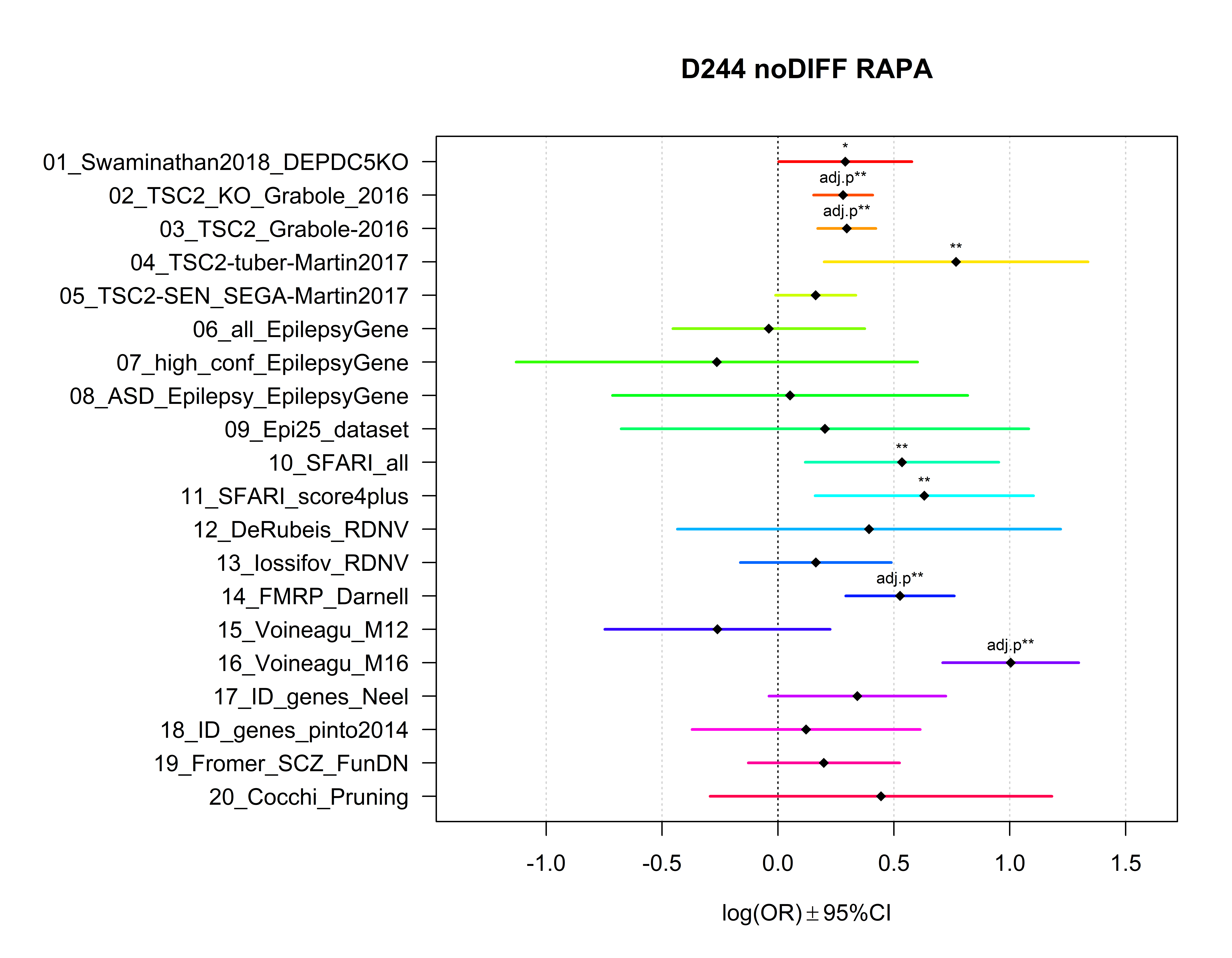
ReN
getlinmodoutput(targetline = "ReN",targetdiff = "noDIFF",targetrapa = "RAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 477 ATP1A2 1.220208 3.413380e-03 1.4114570 5.720993e-08
2 1116 CHI3L1 -8.526972 6.249375e-06 -3.4445448 3.594290e-04
3 283131 NEAT1 -3.020307 1.549094e-02 -2.6088409 2.202376e-03
4 100996492 CTXND1 2.154011 1.002823e-02 2.0926141 1.808240e-03
5 147495 APCDD1 3.498348 6.176015e-03 3.6529841 6.140606e-05
6 7837 PXDN 1.283920 7.440217e-04 0.8097423 2.805530e-04
7 6335 SCN9A -8.223257 1.521923e-04 -7.4694341 1.809116e-04
8 1292 COL6A2 -1.451055 4.091302e-02 -1.6813575 1.603615e-05
9 3693 ITGB5 1.102633 4.887867e-02 0.9037382 6.302982e-04
10 55752 SEPTIN11 1.161097 1.174152e-02 0.5291060 1.394031e-02
11 64065 PERP -1.162849 1.192027e-02 -0.8848644 8.653585e-03
12 148 ADRA1A 1.225387 2.497271e-02 1.6096956 3.510562e-08
13 793 CALB1 -1.195628 2.325343e-03 -1.4001215 7.414265e-13
14 9568 GABBR2 1.236450 1.521923e-04 0.8179580 5.197383e-05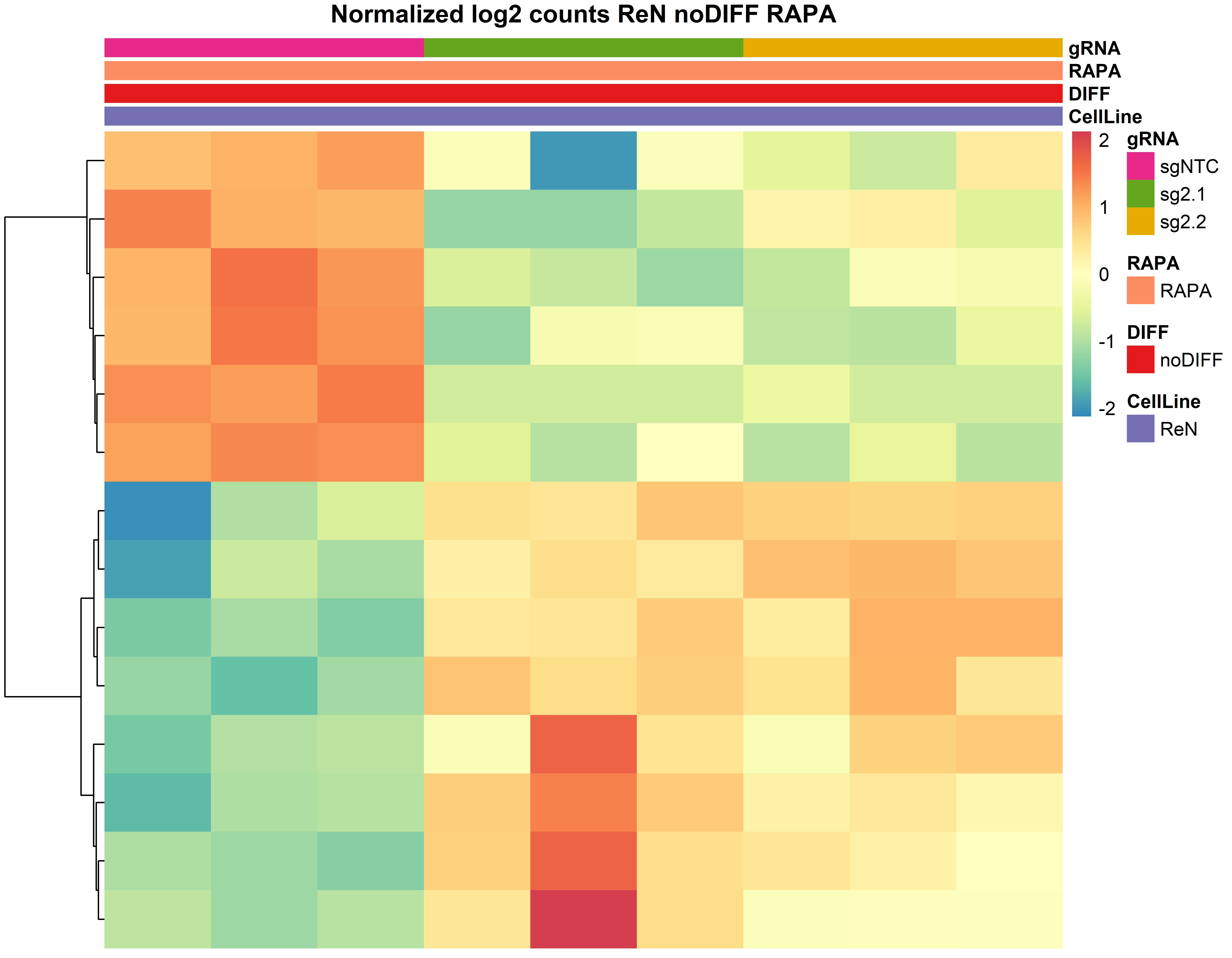
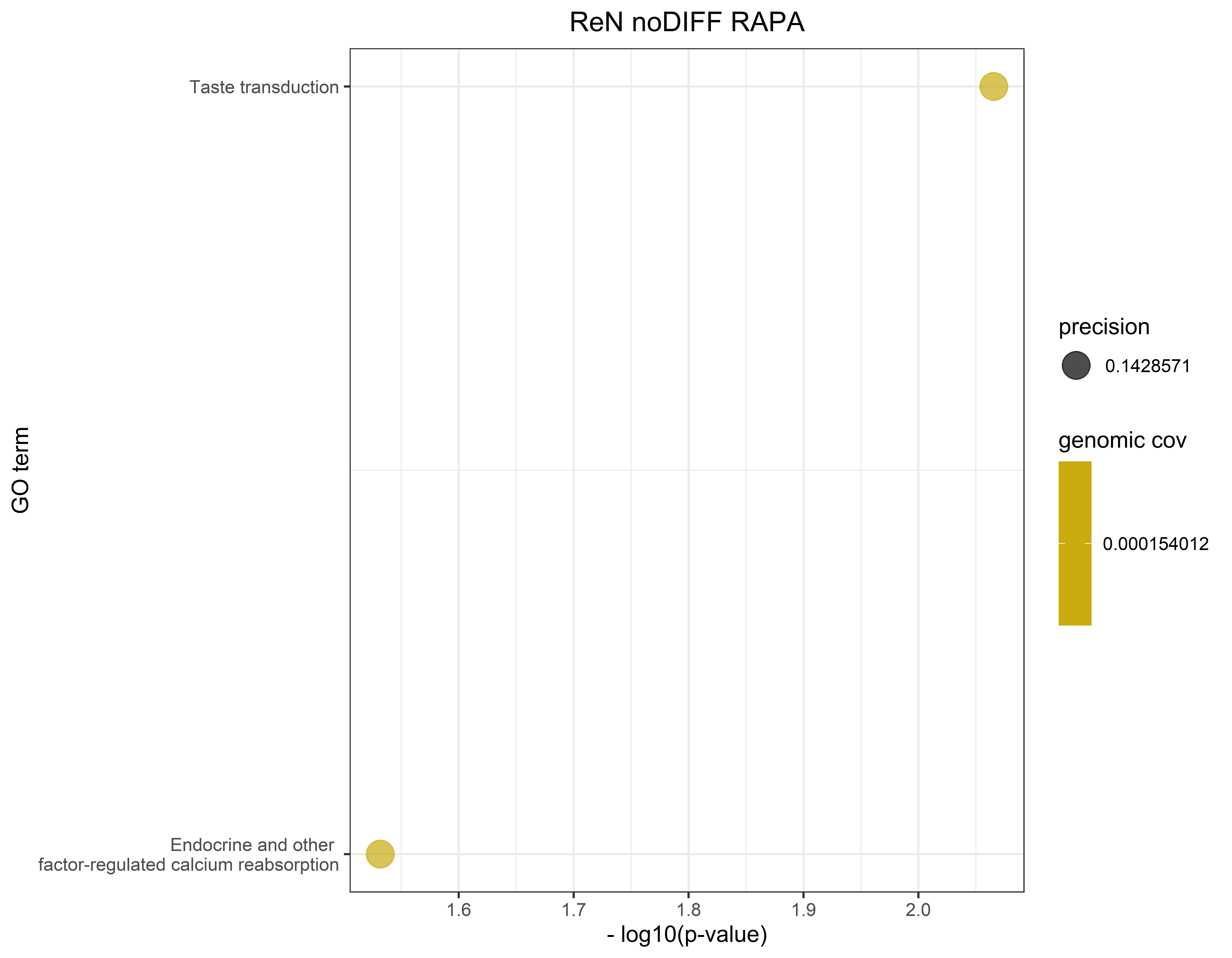
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.000000 0.00000000 6.668883 1.000000000
02_TSC2_KO_Grabole_2016 1.881390 0.56272516 6.289801 0.264620693
03_TSC2_Grabole-2016 1.330939 0.40472473 4.655797 0.790420601
04_TSC2-tuber-Martin2017 0.000000 0.00000000 36.590621 1.000000000
05_TSC2-SEN_SEGA-Martin2017 3.372386 0.88683771 11.219330 0.037502309
06_all_EpilepsyGene 9.565320 1.70653483 36.434078 0.006238071
07_high_conf_EpilepsyGene 18.301000 1.96933619 83.575682 0.007036976
08_ASD_Epilepsy_EpilepsyGene 0.000000 0.00000000 35.336315 1.000000000
09_Epi25_dataset 0.000000 0.00000000 51.036325 1.000000000
10_SFARI_all 0.000000 0.00000000 17.098391 1.000000000
11_SFARI_score4plus 0.000000 0.00000000 23.217073 1.000000000
12_DeRubeis_RDNV 0.000000 0.00000000 53.747406 1.000000000
13_Iossifov_RDNV 1.984480 0.04660814 13.257344 0.413250326
14_FMRP_Darnell 2.840533 0.30822586 12.795519 0.180510915
15_Voineagu_M12 0.000000 0.00000000 11.785713 1.000000000
16_Voineagu_M16 2.984612 0.07003711 19.976342 0.300270008
17_ID_genes_Neel 0.000000 0.00000000 12.348174 1.000000000
18_ID_genes_pinto2014 0.000000 0.00000000 16.806456 1.000000000
19_Fromer_SCZ_FunDN 2.047571 0.04808734 13.680127 0.403726846
20_Cocchi_Pruning 0.000000 0.00000000 45.842037 1.000000000
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -Inf NaN 1.0000000
02_TSC2_KO_Grabole_2016 0.6320107 0.6158034 1.0000000
03_TSC2_Grabole-2016 0.2858845 0.6073636 1.0000000
04_TSC2-tuber-Martin2017 -Inf NaN 1.0000000
05_TSC2-SEN_SEGA-Martin2017 1.2156204 0.6814866 0.7500462
06_all_EpilepsyGene 2.2581440 0.8794281 0.1247614
07_high_conf_EpilepsyGene 2.9069557 1.1373771 0.1407395
08_ASD_Epilepsy_EpilepsyGene -Inf NaN 1.0000000
09_Epi25_dataset -Inf NaN 1.0000000
10_SFARI_all -Inf NaN 1.0000000
11_SFARI_score4plus -Inf NaN 1.0000000
12_DeRubeis_RDNV -Inf NaN 1.0000000
13_Iossifov_RDNV 0.6853570 1.9139475 1.0000000
14_FMRP_Darnell 1.0439918 1.1331195 1.0000000
15_Voineagu_M12 -Inf NaN 1.0000000
16_Voineagu_M16 1.0934696 1.9143876 1.0000000
17_ID_genes_Neel -Inf NaN 1.0000000
18_ID_genes_pinto2014 -Inf NaN 1.0000000
19_Fromer_SCZ_FunDN 0.7166544 1.9139748 1.0000000
20_Cocchi_Pruning -Inf NaN 1.0000000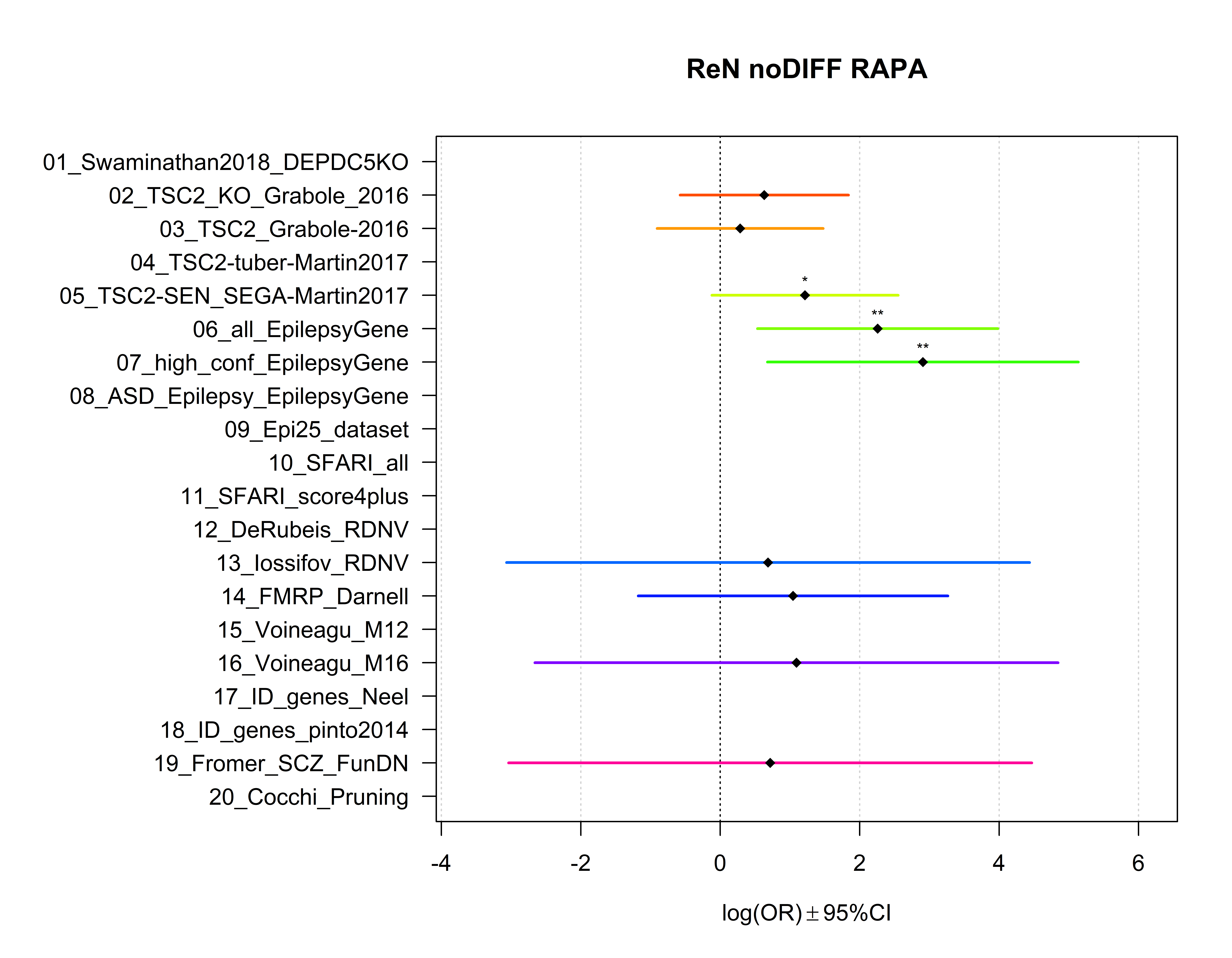
KO effect in DIFF RAPA
D62
getlinmodoutput(targetline = "D62",targetdiff = "DIFF",targetrapa = "RAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 2048 EPHB2 -0.5826101 1.678683e-02 -0.9117379 1.313340e-03
2 2537 IFI6 -0.6988523 7.989899e-03 -0.7770059 3.125215e-02
3 7802 DNALI1 1.1560264 7.989899e-03 1.1151275 2.698021e-02
4 656 BMP8B -4.3683019 9.578298e-05 -4.1372908 5.694587e-04
5 84879 MFSD2A 1.2289963 2.150229e-06 1.7582888 8.298043e-13
6 5865 RAB3B 1.6505809 4.937490e-03 2.4918745 6.877320e-08
7 3491 CCN1 0.9195214 7.581913e-06 0.7819251 2.641835e-02
8 2633 GBP1 1.8909950 1.759866e-03 1.7683986 4.240144e-03
9 127003 C1orf194 1.6879579 3.330240e-04 2.4605882 1.032009e-07
10 26227 PHGDH 0.8707624 5.898199e-03 0.9385243 4.829171e-02
11 10628 TXNIP -1.1214557 3.107157e-03 -1.1801245 6.734027e-03
12 63827 BCAN -1.1448836 9.790123e-07 -0.6860625 4.114286e-02
13 3766 KCNJ10 -1.2030614 1.487223e-04 -1.1249486 2.258695e-03
14 8490 RGS5 0.9423725 2.179796e-03 1.3297066 5.430918e-06
15 55220 KLHDC8A 0.6961260 4.835977e-02 1.0893545 7.903818e-04
16 50486 G0S2 -2.1437954 2.962521e-18 -2.7798517 3.914474e-23
17 7044 LEFTY2 -1.1461461 1.517838e-02 -2.5101270 2.217413e-06
18 2786 GNG4 2.3453579 4.858966e-03 3.1441825 1.403575e-07
19 1959 EGR2 3.8839588 1.874906e-14 3.0338531 5.266483e-05
20 84858 ZNF503 1.8061212 2.018034e-06 1.2808100 2.206113e-02
21 2674 GFRA1 -1.7237479 8.130698e-05 -1.9850405 7.413693e-05
22 5654 HTRA1 0.9285695 1.920731e-05 0.9326186 1.201306e-03
23 10418 SPON1 2.9484113 1.881502e-29 3.3355085 3.371835e-38
24 4038 LRP4 -0.7469557 9.813347e-03 -1.0170181 8.237349e-03
25 374393 FAM111B 3.2862486 3.272494e-02 3.5433728 1.519904e-02
26 79026 AHNAK 0.6672394 4.172050e-02 0.8083736 3.003533e-02
27 254263 CNIH2 1.8795923 1.019547e-04 2.0891824 2.361622e-06
28 2 A2M 1.5219410 1.637477e-12 1.4270188 9.741003e-07
29 79365 BHLHE41 1.2107203 7.466811e-04 1.2350832 4.063916e-03
30 6857 SYT1 1.6677230 1.075725e-05 1.8266367 2.280229e-06
31 101929005 <NA> -6.9715587 4.902298e-04 -4.6002948 9.128832e-03
32 114795 TMEM132B -2.0420745 9.180844e-15 -1.4932069 6.161726e-06
33 688 KLF5 1.6322523 1.076324e-03 1.2452259 4.025393e-02
34 1948 EFNB2 0.9663081 7.296640e-05 1.1138922 2.080333e-04
35 23002 DAAM1 0.7554336 2.883318e-02 0.8353463 2.729476e-02
36 6252 RTN1 1.7650685 1.637477e-12 1.5572796 2.559488e-04
37 89932 PAPLN -1.5716382 4.500273e-02 -1.9661083 1.608497e-02
38 10516 FBLN5 2.0034532 5.633136e-26 2.5520488 2.184210e-21
39 26153 KIF26A 4.9479081 5.215718e-03 5.7609605 3.792368e-05
40 7057 THBS1 1.3681011 2.335652e-03 2.0741402 4.016936e-06
41 4756 NEO1 0.9742518 3.129045e-08 1.1479467 1.474883e-05
42 4916 NTRK3 1.5317619 4.721754e-08 1.3041078 1.647761e-04
43 2903 GRIN2A 2.6808625 3.537752e-05 2.5409853 4.486943e-04
44 729993 SHISA9 -3.8900657 6.124761e-22 -1.3730096 3.597350e-03
45 51704 GPRC5B -0.6943593 1.607767e-02 -0.7158227 4.390799e-02
46 54768 HYDIN 3.1155151 3.786053e-07 3.4412476 6.679975e-10
47 794 CALB2 1.9745979 3.021138e-03 2.4744374 1.492341e-06
48 57687 VAT1L 1.0691442 3.043079e-09 1.1210515 7.707488e-06
49 5187 PER1 1.6211046 2.190498e-03 1.9721847 3.039329e-06
50 201161 CENPV 1.0748005 7.683185e-03 1.3627941 2.911996e-04
51 7153 TOP2A 0.8909203 2.167562e-03 0.9775318 6.734027e-03
52 10882 C1QL1 -1.2462122 5.655205e-06 -1.2834520 1.792262e-03
53 113026 PLCD3 -0.7125454 2.066817e-02 -0.8306542 3.008581e-02
54 4804 NGFR -2.2008093 1.158457e-10 -2.1406331 1.813272e-06
55 2302 FOXJ1 1.2523610 5.640820e-06 1.3486622 1.172300e-03
56 11322 TMC6 -4.5286641 5.215718e-03 -4.0938782 3.270737e-02
57 81035 COLEC12 -1.4481407 2.447676e-02 -2.1538079 1.443187e-04
58 361 AQP4 2.5636450 2.492154e-18 3.3509493 1.099340e-16
59 5596 MAPK4 1.9477813 6.272020e-04 2.2808623 7.694179e-08
60 51046 ST8SIA3 2.3046926 3.116301e-03 2.6061923 1.164931e-03
61 90701 SEC11C -0.6549366 3.435661e-02 -0.7731007 4.443954e-02
62 404217 CTXN1 1.0659448 8.423358e-05 1.0100041 9.810692e-04
63 1995 ELAVL3 1.3869316 2.022244e-05 2.2056761 2.475397e-13
64 9518 GDF15 -0.7403892 4.767716e-02 -1.6894578 3.955062e-05
65 79784 MYH14 -2.3279006 1.986276e-02 -3.3385941 1.061620e-03
66 92749 DRC1 2.1693851 3.021138e-03 2.6759469 3.653718e-06
67 57628 DPP10 -5.2195891 3.952380e-08 -3.4116486 3.792368e-05
68 3625 INHBB -0.9765568 9.918180e-03 -1.6587721 5.064936e-06
69 2995 GYPC -2.3370979 5.640820e-06 -2.1226178 2.620590e-06
70 129881 CCDC173 2.3618858 1.709086e-03 2.4161211 5.153394e-03
71 8324 FZD7 -1.3251978 1.958775e-04 -1.0401722 2.527407e-02
72 8609 KLF7 1.0357609 1.588294e-05 0.8513477 1.101567e-02
73 3488 IGFBP5 -1.2042255 1.457509e-03 -1.2840334 3.354261e-03
74 5270 SERPINE2 -1.3848242 2.877852e-07 -1.2444555 4.917038e-04
75 92737 DNER 1.1961221 4.937490e-03 1.4081602 9.810692e-04
76 3769 KCNJ13 1.7530569 2.673956e-02 2.4403902 3.280473e-05
77 4826 NNAT 3.6894568 3.488356e-26 4.2039042 2.362392e-28
78 55959 SULF2 -1.4277449 1.159590e-02 -1.7758706 2.595059e-02
79 56245 C21orf62 1.0646833 4.488374e-05 0.8807853 2.698021e-02
80 116448 OLIG1 -1.1515256 6.288787e-03 -1.1549137 1.146610e-02
81 1827 RCAN1 0.7193603 2.551274e-03 0.9394508 1.688169e-04
82 3708 ITPR1 1.9492299 8.682648e-03 1.9092677 3.573171e-02
83 116135 LRRC3B 1.9133409 1.961045e-03 1.7450856 1.695501e-02
84 114884 OSBPL10 -1.9426068 5.017110e-03 -1.7311338 3.270737e-02
85 79442 LRRC2 3.4398862 2.502129e-03 3.6516561 2.331951e-04
86 64147 KIF9 1.5020285 1.328423e-04 2.1385875 6.097979e-08
87 11170 FAM107A 1.4030083 3.471441e-02 1.4558105 4.169884e-02
88 132204 SYNPR 6.8513818 2.117888e-04 6.6455378 2.331951e-04
89 1295 COL8A1 -4.5471851 7.161381e-03 -3.7379338 2.815452e-02
90 151887 CCDC80 0.7317143 1.740585e-02 0.8723305 1.319498e-03
91 7018 TF -3.7162415 1.267988e-67 -2.8326531 2.204247e-30
92 5947 RBP1 -1.0910735 4.502025e-07 -0.7994095 4.109501e-02
93 5806 PTX3 1.5680761 1.391425e-04 1.7658383 1.403575e-07
94 10417 SPON2 -2.4640319 1.386561e-02 -2.8331591 2.289093e-03
95 83888 FGFBP2 -1.9562182 9.567416e-03 -2.3549128 2.393688e-04
96 201780 SLC10A4 1.0951238 6.731157e-06 1.0314890 4.917266e-03
97 23284 ADGRL3 -3.3747984 7.547937e-11 -1.7590050 1.454827e-03
98 9508 ADAMTS3 2.2334709 4.424914e-04 2.4774894 2.276936e-04
99 134121 C5orf49 1.9616110 1.942062e-04 2.7374028 1.875367e-10
100 8507 ENC1 0.9047363 2.522638e-02 1.0186176 3.003533e-02
101 1946 EFNA5 1.6677640 1.275805e-06 1.7088384 2.736112e-05
102 9547 CXCL14 -0.8064340 7.756694e-04 -1.5018443 1.611197e-09
103 6695 SPOCK1 -1.1943301 9.157880e-03 -2.2354822 7.206010e-08
104 1958 EGR1 2.4401660 4.817670e-35 2.5287577 7.564100e-19
105 2878 GPX3 -1.3013559 3.780855e-08 -2.0926990 6.557503e-11
106 23286 WWC1 1.0141019 4.480737e-03 1.2974451 1.326571e-04
107 1843 DUSP1 1.9108451 3.135363e-02 1.9266165 2.620078e-02
108 51617 NSG2 6.1426994 3.418688e-03 6.8884048 3.792368e-05
109 10814 CPLX2 5.4606577 4.901378e-04 6.4010760 1.032009e-07
110 221749 PXDC1 0.6438690 5.973334e-03 0.8543232 7.604554e-03
111 84062 DTNBP1 -0.9335939 1.619760e-02 -1.0706331 8.110300e-03
112 3400 ID4 1.0490761 1.704169e-04 1.0639650 3.249851e-03
113 10590 SCGN 3.4205650 6.377511e-11 4.5896530 4.348573e-26
114 285755 PPIL6 1.9279650 1.149142e-02 2.0954784 3.249945e-03
115 1490 CCN2 0.9956475 7.740168e-05 0.7663979 2.527407e-02
116 80129 CCDC170 2.1586226 4.715202e-02 2.8181988 8.319996e-03
117 10457 GPNMB -1.5291994 9.285904e-04 -2.1669377 1.004649e-07
118 4852 NPY 3.1522125 8.913224e-05 4.2222539 1.588167e-14
119 6424 SFRP4 -0.7815462 4.441654e-04 -1.3426635 1.190509e-07
120 81628 TSC22D4 -0.6157171 1.986276e-02 -0.7767061 3.021189e-02
121 5054 SERPINE1 1.6827552 1.689858e-08 1.2590846 3.249945e-03
122 2861 GPR37 1.3670610 2.441806e-03 1.8297795 1.213313e-04
123 10395 DLC1 1.6123746 1.080367e-09 1.4263211 1.924268e-06
124 1808 DPYSL2 1.0114199 5.825476e-08 1.0652386 2.958230e-06
125 23643 LY96 -2.5944597 2.432275e-03 -1.9480631 4.727642e-02
126 11075 STMN2 4.3069575 1.817215e-02 5.8540685 8.750900e-09
127 6383 SDC2 0.8744463 2.437254e-04 0.7889015 1.687242e-02
128 6674 SPAG1 2.0789941 2.964268e-03 2.0054451 1.090224e-02
129 1029 CDKN2A -1.0355011 1.361593e-03 -1.0444652 4.684678e-03
130 687 KLF9 1.4648767 5.328514e-09 1.6563008 6.155819e-08
131 9568 GABBR2 2.3415232 5.056054e-03 2.2372413 3.030523e-03
132 54566 EPB41L4B -1.0936684 2.311572e-03 -1.1537337 5.516218e-04
133 1620 BRINP1 2.9343779 2.911080e-02 3.7325356 2.891086e-05
134 83543 AIF1L -0.9344874 1.208559e-03 -0.9029015 2.547855e-02
135 8021 NUP214 -0.8740175 4.181539e-03 -0.9834048 5.828119e-03
136 5730 PTGDS -2.0109583 5.051669e-20 -1.9008593 7.330218e-09
137 4129 MAOB 1.2838784 2.040953e-05 1.0111052 1.111439e-02
138 1641 DCX 3.7333360 1.413819e-18 5.0056019 2.802915e-51
139 1069 CETN2 0.6735559 1.149142e-02 0.9817983 9.286896e-05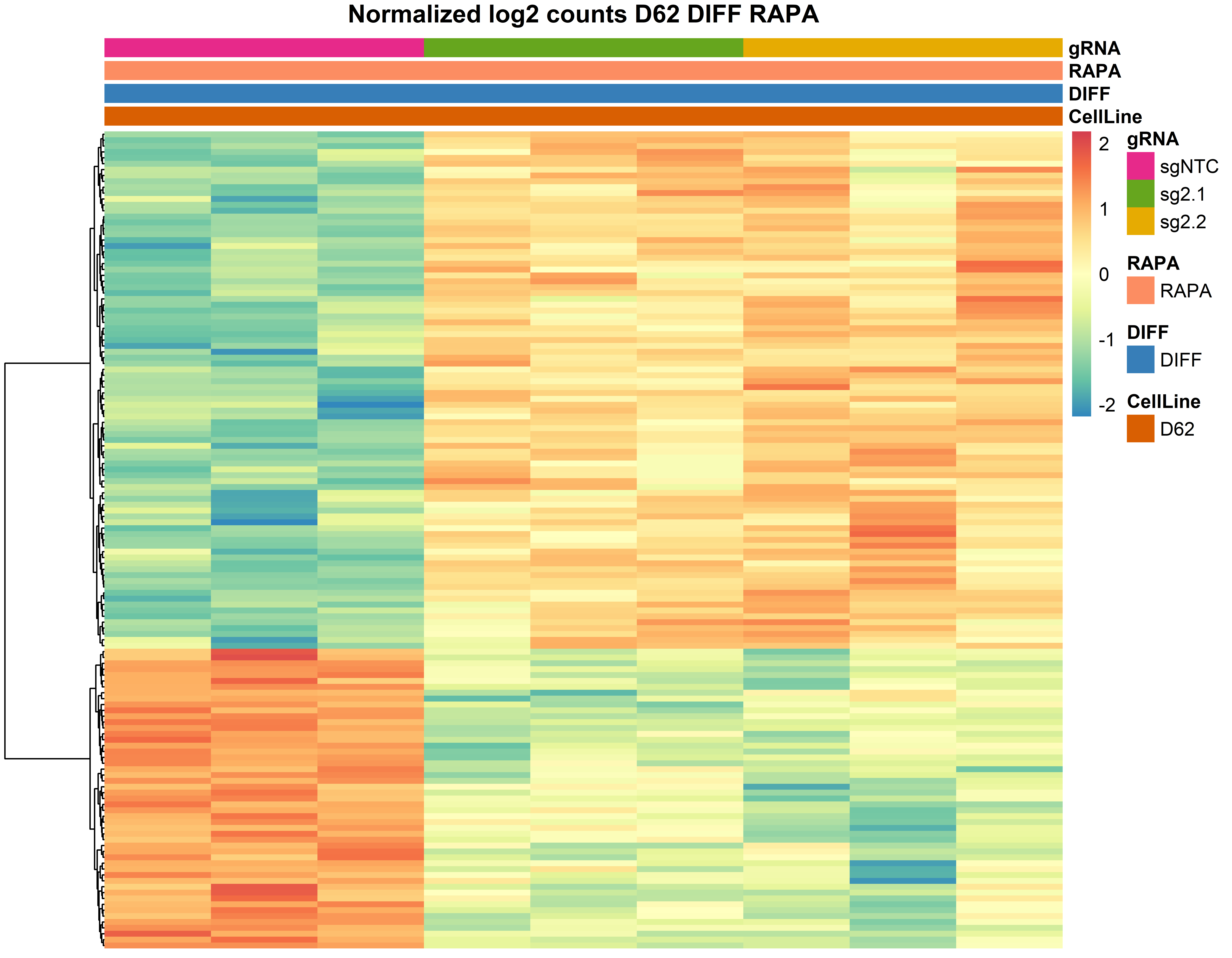
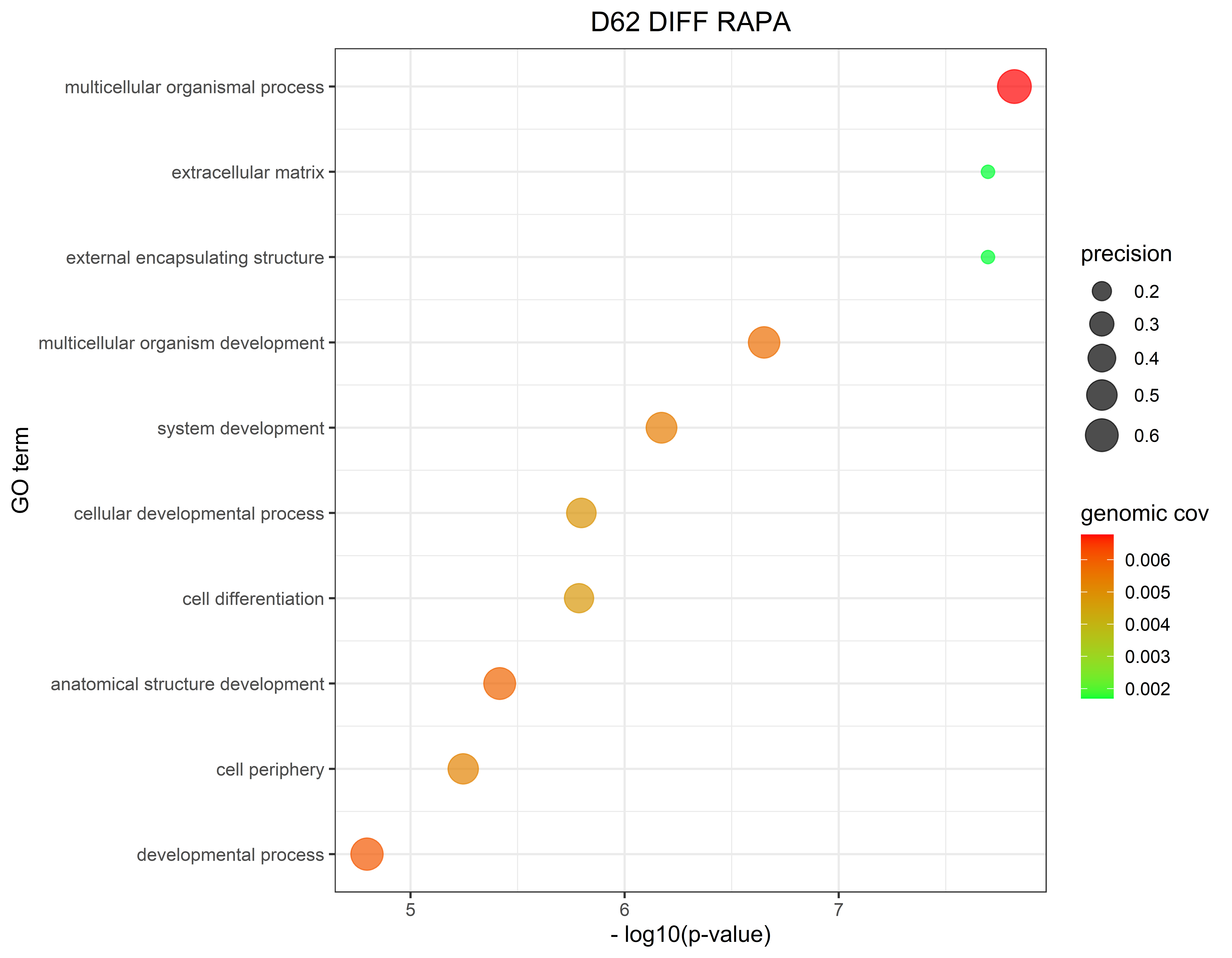
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.3531683 0.56940258 2.763972 3.973374e-01
02_TSC2_KO_Grabole_2016 2.1585572 1.52276295 3.066294 8.934558e-06
03_TSC2_Grabole-2016 1.9664117 1.36679606 2.862101 1.562368e-04
04_TSC2-tuber-Martin2017 4.6371159 1.45237548 11.432969 5.904828e-03
05_TSC2-SEN_SEGA-Martin2017 4.3103219 3.00739593 6.140656 6.426553e-15
06_all_EpilepsyGene 1.3040385 0.41418064 3.147179 4.433955e-01
07_high_conf_EpilepsyGene 2.4223469 0.48662600 7.400112 1.349090e-01
08_ASD_Epilepsy_EpilepsyGene 3.5168838 0.92858316 9.468038 3.146620e-02
09_Epi25_dataset 0.0000000 0.00000000 4.526376 1.000000e+00
10_SFARI_all 3.0516538 1.19006981 6.567324 1.091196e-02
11_SFARI_score4plus 3.5334127 1.25625359 8.057352 9.322558e-03
12_DeRubeis_RDNV 0.0000000 0.00000000 4.768739 1.000000e+00
13_Iossifov_RDNV 0.9613635 0.30580209 2.315034 1.000000e+00
14_FMRP_Darnell 2.0797679 1.12407459 3.587280 1.342012e-02
15_Voineagu_M12 0.8582674 0.17392932 2.583640 1.000000e+00
16_Voineagu_M16 1.4517078 0.46080102 3.506997 4.038906e-01
17_ID_genes_Neel 0.5933426 0.07083178 2.202312 7.762594e-01
18_ID_genes_pinto2014 0.8078992 0.09630282 3.006937 1.000000e+00
19_Fromer_SCZ_FunDN 1.4160882 0.55542785 3.023035 3.555920e-01
20_Cocchi_Pruning 5.8469859 1.82193469 14.520006 2.312936e-03
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.3024488 0.4416410 1.000000e+00
02_TSC2_KO_Grabole_2016 0.7694400 0.1780171 1.786912e-04
03_TSC2_Grabole-2016 0.6762104 0.1855822 3.124735e-03
04_TSC2-tuber-Martin2017 1.5340926 0.5922919 1.180966e-01
05_TSC2-SEN_SEGA-Martin2017 1.4610126 0.1836418 1.285311e-13
06_all_EpilepsyGene 0.2654660 0.5851628 1.000000e+00
07_high_conf_EpilepsyGene 0.8847369 0.8188757 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 1.2575753 0.6794238 6.293241e-01
09_Epi25_dataset -Inf NaN 1.000000e+00
10_SFARI_all 1.1156837 0.4804447 2.182393e-01
11_SFARI_score4plus 1.2622642 0.5276175 1.864512e-01
12_DeRubeis_RDNV -Inf NaN 1.000000e+00
13_Iossifov_RDNV -0.0394027 0.5843951 1.000000e+00
14_FMRP_Darnell 0.7322563 0.3139266 2.684024e-01
15_Voineagu_M12 -0.1528395 0.8144218 1.000000e+00
16_Voineagu_M16 0.3727406 0.5854743 1.000000e+00
17_ID_genes_Neel -0.5219834 1.0844205 1.000000e+00
18_ID_genes_pinto2014 -0.2133179 1.0851734 1.000000e+00
19_Fromer_SCZ_FunDN 0.3478983 0.4775076 1.000000e+00
20_Cocchi_Pruning 1.7659263 0.5949119 4.625873e-02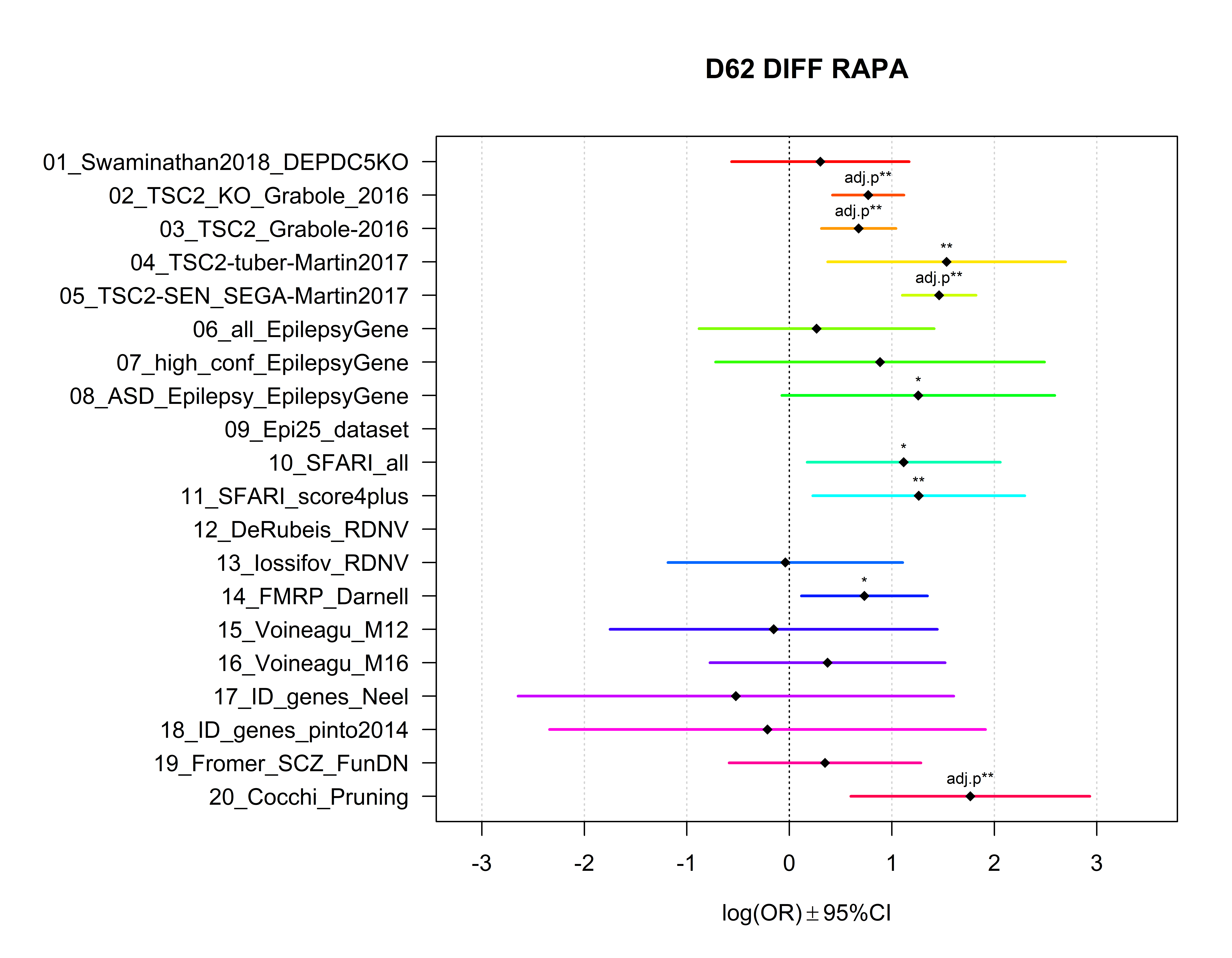
D244
getlinmodoutput(targetline = "D244",targetdiff = "DIFF",targetrapa = "RAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 1969 EPHA2 -0.4156436 1.786505e-03 -0.4614548 1.856825e-02
2 4237 MFAP2 -1.8799463 1.408076e-06 -1.3745964 4.624004e-04
3 644068 <NA> -0.3590070 2.869457e-02 -0.4807850 1.104913e-02
4 3399 ID3 0.5061376 7.729593e-06 0.4283983 3.908169e-04
5 3925 STMN1 0.6965907 7.631803e-04 0.5450212 8.437631e-03
6 1296 COL8A2 -1.2868634 7.261643e-07 -1.2888966 9.397150e-08
7 2899 GRIK3 -2.3312172 6.975035e-10 -0.8969046 4.857303e-02
8 8613 PLPP3 0.9733065 1.474275e-04 0.7153388 4.318621e-02
9 23576 DDAH1 0.7785829 5.192707e-03 0.6091387 4.100636e-02
10 3491 CCN1 0.4678656 2.963636e-06 0.4105692 9.041386e-05
11 648740 <NA> 0.6129398 3.756801e-03 0.5676909 3.742487e-02
12 7482 WNT2B 0.6449332 1.707551e-03 0.6721599 2.189161e-04
13 54879 ST7L 1.3190671 2.143140e-23 0.4878723 4.835702e-03
14 6277 S100A6 -0.6507256 1.266693e-10 -0.6579181 5.467558e-13
15 79957 PAQR6 -1.8264594 4.444419e-10 -1.1683542 7.608800e-04
16 10485 MIR9-1HG -0.9291450 2.838604e-04 -1.2607947 1.034161e-04
17 93183 PIGM -1.3311259 3.672953e-07 -0.6802182 3.895651e-02
18 477 ATP1A2 -1.2574533 1.434561e-16 -0.9292025 2.359299e-06
19 8490 RGS5 0.7061001 9.243401e-03 0.5901435 6.181386e-03
20 9019 MPZL1 0.4981399 5.092077e-03 0.5009347 9.496796e-03
21 5396 PRRX1 1.7234056 9.775055e-14 1.1534507 6.426046e-05
22 9588 PRDX6 0.4707179 5.893131e-04 0.3644539 2.151528e-02
23 55220 KLHDC8A -2.1231857 1.760830e-03 -1.4689768 4.847504e-02
24 50486 G0S2 -2.0420221 9.635264e-145 -2.5996040 2.301167e-217
25 7044 LEFTY2 -1.3192934 3.280549e-18 -2.3110081 5.751049e-64
26 84886 C1orf198 0.9699749 1.586808e-06 0.5353448 3.514587e-02
27 10659 CELF2 1.0623173 5.832457e-08 0.7868175 1.708568e-04
28 3799 KIF5B 0.4577931 5.230602e-04 0.4754410 5.092168e-05
29 220963 SLC16A9 1.5093231 9.717264e-18 1.4702889 6.249604e-20
30 5328 PLAU -0.6860737 4.551671e-05 -0.5197041 9.395697e-03
31 9124 PDLIM1 0.4509083 1.314632e-04 0.6513184 1.993098e-08
32 6319 SCD 0.2511798 4.476925e-02 0.7794106 6.387888e-12
33 114815 SORCS1 -2.1231810 2.307478e-02 -2.3863755 2.202177e-02
34 2674 GFRA1 -0.4799768 1.237850e-03 -0.6378438 1.191763e-05
35 402778 IFITM10 -1.6883116 1.798490e-07 -0.9871409 1.595832e-02
36 283298 OLFML1 1.3133104 5.078602e-10 1.5777211 2.403838e-16
37 6157 RPL27A -0.3857288 3.670998e-07 -0.2303211 1.903700e-02
38 27122 DKK3 -0.5580258 4.985218e-09 -0.5931532 3.691467e-10
39 6207 RPS13 -0.4023854 1.385053e-07 -0.3180084 1.061947e-03
40 9537 TP53I11 -1.9584281 3.300614e-16 -1.3950698 1.344610e-07
41 51524 TMEM138 0.5334741 9.492034e-03 0.5147442 2.156053e-02
42 7351 UCP2 1.1143606 2.025606e-02 1.5299811 7.532644e-04
43 11098 PRSS23 0.4307277 1.073122e-05 0.6784633 1.324860e-11
44 1410 CRYAB -0.7085678 3.425418e-16 -0.6623639 4.194395e-13
45 4684 NCAM1 -0.7849245 6.342683e-09 -0.6951515 8.970563e-08
46 6876 TAGLN 0.3837716 1.888557e-06 0.7098218 5.489084e-18
47 53826 FXYD6 -1.5563297 2.641754e-29 -1.7253622 4.449397e-31
48 63876 PKNOX2 -1.2447871 1.637874e-03 -1.7811272 3.247027e-04
49 50863 NTM -1.2825953 1.370990e-02 -1.2557436 3.308547e-02
50 81029 WNT5B -1.4638818 4.656250e-05 -0.8511374 2.355254e-02
51 894 CCND2 0.5412578 2.056792e-02 0.6213173 1.140256e-02
52 8076 MFAP5 0.8323568 1.312028e-08 1.3427802 1.485392e-17
53 11228 RASSF8 1.5726288 4.434900e-04 1.1483820 4.832576e-02
54 3855 KRT7 1.3224258 2.551487e-05 2.2390807 1.133350e-15
55 3489 IGFBP6 -1.1739264 3.487772e-08 -0.5194090 2.079860e-02
56 4327 MMP19 0.9211195 4.177778e-02 1.2420465 1.677874e-03
57 4637 MYL6 0.3799577 8.194798e-04 0.4319203 2.137579e-04
58 4035 LRP1 -0.3603670 4.450603e-02 -0.3576910 4.028103e-02
59 11247 NXPH4 -2.6634395 2.863258e-03 -2.5454342 1.798882e-02
60 22822 PHLDA1 0.7192724 6.432734e-06 0.5243860 5.347219e-03
61 1848 DUSP6 -1.2864987 6.654183e-07 -1.3246108 5.992254e-06
62 10988 METAP2 -0.4486932 3.733085e-03 -0.4181849 4.795325e-02
63 22877 MLXIP -0.8864682 7.710864e-05 -1.3537995 8.103193e-11
64 114795 TMEM132B -3.1403066 4.570484e-06 -3.0132607 5.957135e-10
65 10129 FRY 3.3215631 7.103118e-03 3.5798206 7.133626e-03
66 1910 EDNRB -0.9341303 8.787660e-05 -1.4018162 2.523953e-09
67 1638 DCT -0.9777289 1.598509e-05 -2.1672010 2.513793e-30
68 1282 COL4A1 0.6959730 6.185122e-07 0.3346814 3.788431e-02
69 2621 GAS6 0.6469828 8.399232e-06 0.4991532 2.771146e-03
70 29091 STXBP6 -1.1957346 5.007498e-03 -1.3175027 5.129299e-03
71 6235 RPS29 -0.2925477 6.939818e-05 -0.2161987 1.523302e-02
72 652 BMP4 0.9172466 4.681102e-09 0.9744670 8.103193e-11
73 23002 DAAM1 0.6938355 2.863809e-03 0.5782007 4.677801e-02
74 9517 SPTLC2 0.8748820 3.023905e-02 0.9166731 4.185679e-02
75 12 SERPINA3 -1.6939009 3.691257e-46 -1.0237219 8.432118e-20
76 70 ACTC1 2.4566660 3.961311e-09 2.3862088 2.641230e-08
77 567 B2M 0.4300572 1.118831e-03 0.4063694 1.601715e-04
78 3671 ISLR -1.5492885 2.020700e-55 -0.6997528 6.387888e-12
79 64220 STRA6 -0.8436189 1.277936e-08 -0.8128532 9.165614e-08
80 9915 ARNT2 0.4654090 2.139752e-04 0.6806762 1.790427e-08
81 57214 CEMIP 0.9918949 8.936810e-09 1.0669818 5.193301e-11
82 30812 SOX8 -1.1040177 1.160839e-03 -0.9873133 1.953103e-03
83 124152 IQCK -0.4233033 9.984072e-03 -0.4341487 9.955852e-03
84 1428 CRYM -5.3477355 1.887871e-06 -2.2455586 2.375695e-02
85 4313 MMP2 -1.1477944 2.187632e-19 -0.8585802 8.659959e-16
86 1728 NQO1 0.6514906 2.602651e-06 0.5555884 2.699670e-03
87 92154 MTSS2 0.9485278 1.521300e-05 0.6963852 2.405820e-02
88 91862 MARVELD3 1.4914019 6.669175e-03 1.5623765 7.427253e-03
89 497190 CLEC18B -1.4112298 1.264664e-11 -0.8408839 3.594117e-04
90 2734 GLG1 -0.7611539 1.728431e-08 -0.4129051 8.723320e-03
91 482 ATP1B2 -0.7442227 8.755923e-07 -0.4070386 3.679534e-02
92 1819 DRG2 1.6549430 4.877721e-02 1.8554310 2.740463e-02
93 9349 RPL23 -0.4447861 1.041512e-09 -0.2199463 2.445119e-02
94 3872 KRT17 0.7072854 2.220292e-02 1.3596930 5.484501e-08
95 284119 CAVIN1 0.3711938 7.598398e-03 0.9338717 7.289930e-17
96 2670 GFAP -1.5458422 1.066141e-81 -0.4623582 5.343221e-06
97 10882 C1QL1 -1.1800573 4.585792e-02 -1.3575829 3.915880e-02
98 4804 NGFR -1.8994842 1.471545e-13 -2.1854385 4.715661e-26
99 1277 COL1A1 -0.6243475 4.152227e-12 -0.5539194 3.691467e-10
100 6909 TBX2 -1.6172291 6.673132e-05 -1.4042385 9.532588e-05
101 3691 ITGB4 -1.5170148 8.222125e-10 -0.7373586 1.101639e-02
102 114897 C1QTNF1 -2.0736184 7.925386e-17 -2.0009867 1.870909e-26
103 84617 TUBB6 0.6784353 6.292697e-10 0.8419118 5.824719e-18
104 79959 CEP76 2.1133577 3.622145e-02 2.3607183 4.289137e-02
105 5771 PTPN2 1.5822769 2.975000e-03 1.4786379 1.570613e-02
106 1630 DCC -1.8781777 3.480412e-02 -5.3854841 4.688455e-07
107 6925 TCF4 -0.3524476 2.137204e-02 -0.8766528 2.523953e-09
108 115701 ALPK2 0.9407645 5.057115e-20 1.2657357 9.796637e-33
109 1153 CIRBP -0.4781032 1.217141e-04 -0.3043657 2.673622e-02
110 1264 CNN1 0.7833203 8.186223e-05 1.2267607 3.757565e-15
111 7171 TPM4 0.7593549 4.432673e-11 0.3268238 4.546490e-02
112 348 APOE 1.1060994 7.703717e-09 1.0078352 5.834122e-07
113 255783 INAFM1 -1.1227281 2.626564e-05 -1.3845743 8.253297e-05
114 284297 SSC5D -1.3402850 2.417846e-04 -0.8886371 3.619953e-02
115 130733 TMEM178A -0.5897743 1.879698e-02 -0.8015248 2.229551e-03
116 53335 BCL11A -1.4709084 2.372813e-02 -2.2288475 4.857303e-02
117 7039 TGFA -1.8402630 3.130862e-20 -2.0866510 1.274152e-27
118 54910 SEMA4C -1.0299388 1.576569e-03 -0.8420389 4.919172e-02
119 6160 RPL31 -0.2593406 1.319576e-03 -0.2546111 2.243391e-02
120 9448 MAP4K4 -0.6868182 5.285781e-13 -0.5734919 9.596336e-10
121 2995 GYPC -2.4190113 1.415998e-15 -2.5897350 5.915615e-21
122 116372 LYPD1 -0.6853058 2.175341e-08 -0.5182769 3.838421e-05
123 390 RND3 1.0879074 2.834714e-09 0.5274762 4.546490e-02
124 253782 CERS6 -1.1675559 4.957898e-03 -1.2028556 6.892510e-03
125 1745 DLX1 -1.2542083 3.178833e-05 -1.4832562 9.618358e-07
126 1123 CHN1 0.7343030 1.000126e-05 0.6061334 1.072571e-03
127 2487 FRZB -2.5860101 3.506698e-05 -2.6775791 3.208487e-05
128 23671 TMEFF2 -1.4239778 4.390502e-02 -1.9950630 5.390865e-03
129 26010 SPATS2L 0.8417012 9.779238e-07 0.5132847 2.278601e-03
130 2335 FN1 0.8618042 5.085179e-08 0.5159728 5.347219e-03
131 3485 IGFBP2 0.4209936 6.052721e-06 0.6294041 7.289930e-17
132 3488 IGFBP5 -1.4790171 1.730247e-60 -1.3238131 6.039310e-53
133 80303 EFHD1 -2.6073677 6.139696e-09 -2.5477600 1.515435e-08
134 10123 ARL4C -0.5040582 9.787143e-05 -0.5198076 3.771654e-05
135 57419 SLC24A3 -1.4139082 3.019072e-02 -1.8615651 4.659175e-03
136 3397 ID1 0.7307632 1.742180e-05 0.9582626 8.103193e-11
137 7052 TGM2 0.6840347 5.918459e-04 0.9553814 3.102921e-08
138 441951 ZFAS1 -0.6031713 1.099503e-09 -0.2692960 4.185679e-02
139 56937 PMEPA1 -0.7010615 8.307140e-19 -0.8845874 3.489978e-32
140 6227 RPS21 -0.3018602 8.204456e-04 -0.3142897 7.115799e-03
141 100192386 <NA> -1.9138055 3.039395e-02 -2.8043265 1.898146e-03
142 6453 ITSN1 0.5955656 1.304253e-03 0.6227147 1.035891e-03
143 80781 COL18A1 -0.8032645 1.294700e-05 -1.0810327 2.084319e-11
144 3985 LIMK2 -1.0339131 2.188103e-05 -0.6257838 5.847607e-03
145 54944 <NA> -1.0270303 3.952230e-02 -1.3586023 3.648476e-02
146 7078 TIMP3 -0.4036202 1.521300e-05 -0.6462015 2.397315e-14
147 4627 MYH9 0.2903156 3.693453e-03 0.3375945 9.493447e-04
148 2192 FBLN1 -1.7626343 8.307140e-19 -1.0111310 2.449077e-06
149 10752 CHL1 -0.8174300 7.247512e-04 -1.6241791 1.656380e-13
150 6161 RPL32 -0.2208801 7.113707e-03 -0.2314840 1.568219e-02
151 2199 FBLN2 -1.4666500 2.617758e-04 -1.6215847 6.685872e-04
152 23180 RFTN1 0.5110522 1.313369e-02 0.6579327 9.918696e-04
153 54756 IL17RD -0.3363103 1.102081e-02 -0.4018532 7.357338e-03
154 55079 FEZF2 -1.5300772 2.853931e-04 -1.2334711 1.442041e-02
155 1295 COL8A1 -2.5667320 1.117183e-51 -2.8067329 2.533751e-78
156 100506621 <NA> 1.8591393 1.896477e-09 1.5419284 1.462876e-05
157 151887 CCDC80 0.8400733 1.550300e-13 1.0909859 8.688693e-17
158 57514 ARHGAP31 -0.6484646 1.506550e-02 -0.8011114 4.835702e-03
159 11167 FSTL1 0.9026565 1.367900e-18 0.6720049 4.779641e-13
160 4710 NDUFB4 -0.5832897 2.332886e-08 -0.3109836 1.396695e-02
161 7018 TF -2.2191138 7.809017e-09 -2.7186226 1.812506e-08
162 51421 AMOTL2 0.5824076 2.870027e-02 0.5524496 2.679794e-02
163 5947 RBP1 -1.0346072 3.851251e-48 -1.1250151 1.013389e-56
164 590 BCHE -3.8950563 2.588810e-03 -4.2863525 4.963999e-03
165 5010 CLDN11 0.7786442 7.416085e-08 1.4385772 3.489978e-32
166 2049 EPHB3 -1.7366493 3.513152e-04 -2.0414255 5.942612e-05
167 1427 CRYGS 0.9015175 4.849154e-03 1.9079385 5.453763e-17
168 3280 HES1 -0.3270127 1.310732e-02 -0.3250811 4.690041e-02
169 131583 FAM43A -0.8770903 3.717157e-03 -0.7031054 6.181386e-03
170 6165 RPL35A -0.2663334 4.613388e-04 -0.2290969 1.474304e-02
171 10417 SPON2 -1.6792183 1.085527e-05 -1.6373186 1.499922e-03
172 285489 DOK7 -1.1007513 6.781052e-03 -1.3615172 1.037528e-03
173 54360 CYTL1 -2.8616496 5.812295e-09 -3.4273988 1.444637e-07
174 1400 CRMP1 -1.2791240 2.432843e-07 -0.7019373 1.428682e-02
175 8532 CPZ 0.9776346 1.719013e-07 1.6903718 6.740650e-28
176 83888 FGFBP2 -2.3020082 9.156676e-12 -4.3108924 4.089891e-24
177 51056 LAP3 0.8816155 7.862749e-03 0.7419129 4.857303e-02
178 7345 UCHL1 -0.5710588 6.820231e-06 -0.5653695 2.663678e-06
179 65997 RASL11B -1.7444023 6.820231e-06 -2.2916507 7.434746e-09
180 25849 PARM1 -2.7565914 7.734866e-06 -1.7833252 1.097964e-02
181 79966 SCD5 0.9156362 7.821225e-06 0.6368364 1.721266e-03
182 6164 RPL34 -0.3914409 9.425583e-07 -0.3131570 3.059568e-03
183 1363 CPE 0.9026028 7.977764e-03 1.3201504 8.164630e-06
184 153478 PLEKHG4B 1.0978381 1.536582e-05 0.9747296 5.207002e-04
185 170690 ADAMTS16 -2.1553969 2.241202e-04 -1.4760633 4.088533e-02
186 100505738 MIR4458HG -0.9318569 7.881478e-06 -0.8737341 9.862142e-04
187 56172 ANKH -1.2372352 2.079010e-05 -0.9869900 3.387664e-02
188 4131 MAP1B 0.4133843 2.202179e-04 0.4158426 6.930263e-05
189 645323 LINC00461 -0.5801673 5.283948e-05 -0.5709236 3.766002e-04
190 9547 CXCL14 -0.5235165 1.148369e-10 -0.4632759 1.335559e-07
191 1495 CTNNA1 0.3228209 1.259825e-02 0.3516482 1.056930e-02
192 1809 DPYSL3 -0.7918848 1.518136e-11 -0.7901843 1.915470e-14
193 972 CD74 0.4593239 4.323614e-02 1.4993242 2.671462e-19
194 11346 SYNPO -1.1937925 8.307140e-19 -0.9245496 6.330591e-09
195 2878 GPX3 -1.0164418 8.801801e-26 -0.7308690 1.484571e-13
196 6678 SPARC -0.4396672 1.953837e-09 -0.3373670 1.267784e-04
197 23286 WWC1 1.5686248 3.671663e-06 1.4815662 3.857516e-05
198 8878 SQSTM1 0.4475931 5.491053e-05 0.5283156 5.957135e-10
199 6659 SOX4 -0.3973998 1.168159e-04 -0.3397728 2.715784e-02
200 10590 SCGN -2.5609670 3.273773e-04 -2.2657997 7.608800e-04
201 4188 MDFI -1.1359481 1.479461e-02 -1.1566529 1.795269e-02
202 10231 RCAN2 -1.4933781 1.004177e-02 -2.1589227 9.375633e-05
203 79627 OGFRL1 1.5182381 1.724549e-05 1.1135665 5.674792e-03
204 167681 PRSS35 4.0435832 1.149050e-02 4.1134468 1.112088e-02
205 387066 <NA> -1.0656946 7.617300e-27 -0.3385440 1.548226e-02
206 7101 NR2E1 1.3256192 4.768159e-03 1.3443631 1.404027e-02
207 3910 LAMA4 1.1638947 3.028116e-05 1.2970794 2.967756e-06
208 2697 GJA1 0.6112536 4.987593e-10 0.5420339 8.545475e-09
209 2173 FABP7 -1.2363700 4.004592e-12 -0.6465519 3.678213e-04
210 26002 MOXD1 -0.8866932 8.755923e-07 -0.6483667 4.421584e-03
211 84947 SERAC1 1.1030080 3.872716e-02 1.1984720 4.269294e-02
212 7430 EZR 0.6208804 6.263804e-04 0.8795920 3.688320e-08
213 6648 SOD2 0.6597792 1.936825e-04 0.5997779 3.396741e-03
214 56975 FAM20C -1.0641255 1.989627e-08 -0.4683416 2.778544e-02
215 6624 FSCN1 0.3671651 2.778250e-03 0.4900222 1.291831e-05
216 10457 GPNMB 1.0447754 7.809017e-09 1.4574443 3.569027e-18
217 4852 NPY -2.7004616 2.746461e-03 -3.4530952 1.028876e-02
218 6424 SFRP4 -1.5686024 2.187938e-85 -1.7488886 8.250920e-100
219 2006 ELN -0.9919962 8.029189e-06 -0.7947487 5.390865e-03
220 10371 SEMA3A -1.7778313 1.487989e-02 -3.7214380 1.615111e-06
221 5244 ABCB4 -1.5833379 4.233616e-08 -2.3270915 1.719399e-10
222 5243 ABCB1 -1.2075793 9.159620e-06 -2.4817056 3.528847e-20
223 1278 COL1A2 -2.3263318 2.250375e-68 -2.9836323 5.280241e-106
224 1750 DLX6 -2.0712636 1.500270e-02 -3.2495957 7.155357e-03
225 1749 DLX5 -2.8085778 4.589048e-16 -2.5851513 8.907072e-10
226 83943 IMMP2L 0.4936623 4.856899e-02 0.5922763 2.150495e-02
227 857 CAV1 0.7578834 4.713328e-04 0.8685401 1.553383e-04
228 57464 STRIP2 -0.8663806 1.596751e-03 -0.8831810 5.004934e-03
229 51200 CPA4 3.2736399 9.784024e-04 2.9246067 2.214509e-02
230 4232 MEST 0.9643726 9.353234e-04 1.1818103 2.801103e-06
231 800 CALD1 0.3000359 6.350977e-04 0.3310221 5.092168e-05
232 28996 HIPK2 -0.5164790 7.729593e-06 -0.6035502 2.410705e-07
233 63898 SH2D4A 1.8424114 1.390738e-02 1.9426630 1.399137e-02
234 526 ATP6V1B2 0.4371004 2.044363e-02 0.5162846 1.111914e-02
235 1191 CLU 0.2778341 2.014947e-02 0.4027919 3.473869e-05
236 5368 PNOC -2.4876664 3.864354e-02 -5.4363244 1.046505e-05
237 3084 NRG1 1.9269670 8.288167e-25 1.6209777 2.309162e-14
238 5327 PLAT -1.0654404 1.426133e-05 -0.7292897 1.778405e-02
239 54212 SNTG1 -3.2339115 3.662222e-08 -1.9941244 5.144108e-04
240 767 CA8 -1.3726361 2.869065e-02 -1.3901392 3.368227e-02
241 286183 NKAIN3 -1.0736917 7.302787e-06 -0.7132043 4.659175e-03
242 7381 UQCRB -0.6671588 1.723706e-13 -0.2602459 2.243391e-02
243 6156 RPL30 -0.4589979 1.833585e-09 -0.2216742 4.235332e-02
244 1345 COX6C -0.5370116 4.106892e-09 -0.3059022 2.052665e-02
245 5168 ENPP2 0.8527160 1.101228e-07 0.7599470 8.177880e-07
246 114907 FBXO32 1.1589137 3.709259e-41 0.8963959 2.214941e-18
247 92949 ADAMTSL1 -1.4495735 2.432980e-06 -1.8584701 2.308692e-06
248 7169 TPM2 0.4519311 2.782340e-03 0.8803835 5.668786e-11
249 7094 TLN1 0.4196639 2.246460e-04 0.4401227 1.721266e-03
250 1515 CTSV 1.4388468 3.349723e-02 1.5941814 3.347452e-02
251 19 ABCA1 -0.3793466 3.165081e-02 -0.5800784 4.172341e-03
252 23446 SLC44A1 -0.5902860 4.635741e-02 -0.8323991 6.911771e-05
253 1620 BRINP1 -1.9895159 1.142425e-07 -2.5070024 1.273119e-11
254 253039 CUTALP -1.1174922 6.224565e-03 -0.9852202 3.584199e-02
255 169611 OLFML2A -1.8952074 2.674510e-06 -1.9555276 6.925892e-06
256 100506119 LINC01503 0.5559469 2.680262e-03 0.9130241 3.579174e-08
257 83543 AIF1L -1.2388526 3.130862e-20 -0.5682422 9.922051e-04
258 4570 <NA> -0.8772274 2.663053e-21 -0.3278338 1.585591e-02
259 2824 GPM6B -0.8074180 2.341468e-06 -0.5686222 1.484076e-02
260 7102 TSPAN7 -1.2510598 9.578798e-04 -0.8291136 3.081950e-02
261 51186 TCEAL9 0.4357520 9.519705e-07 0.3011626 3.423865e-03
262 5354 PLP1 -2.2504674 6.574852e-04 -3.1849628 1.981893e-05
263 5358 PLS3 -0.8969021 3.440901e-03 -1.6719493 3.063947e-10
264 10857 PGRMC1 0.8842565 3.302791e-07 0.5478687 3.603326e-03
265 6170 RPL39 -0.2996484 4.105697e-04 -0.3137189 1.464648e-04
266 633 BGN -2.9029508 4.191791e-25 -1.8005635 3.737921e-11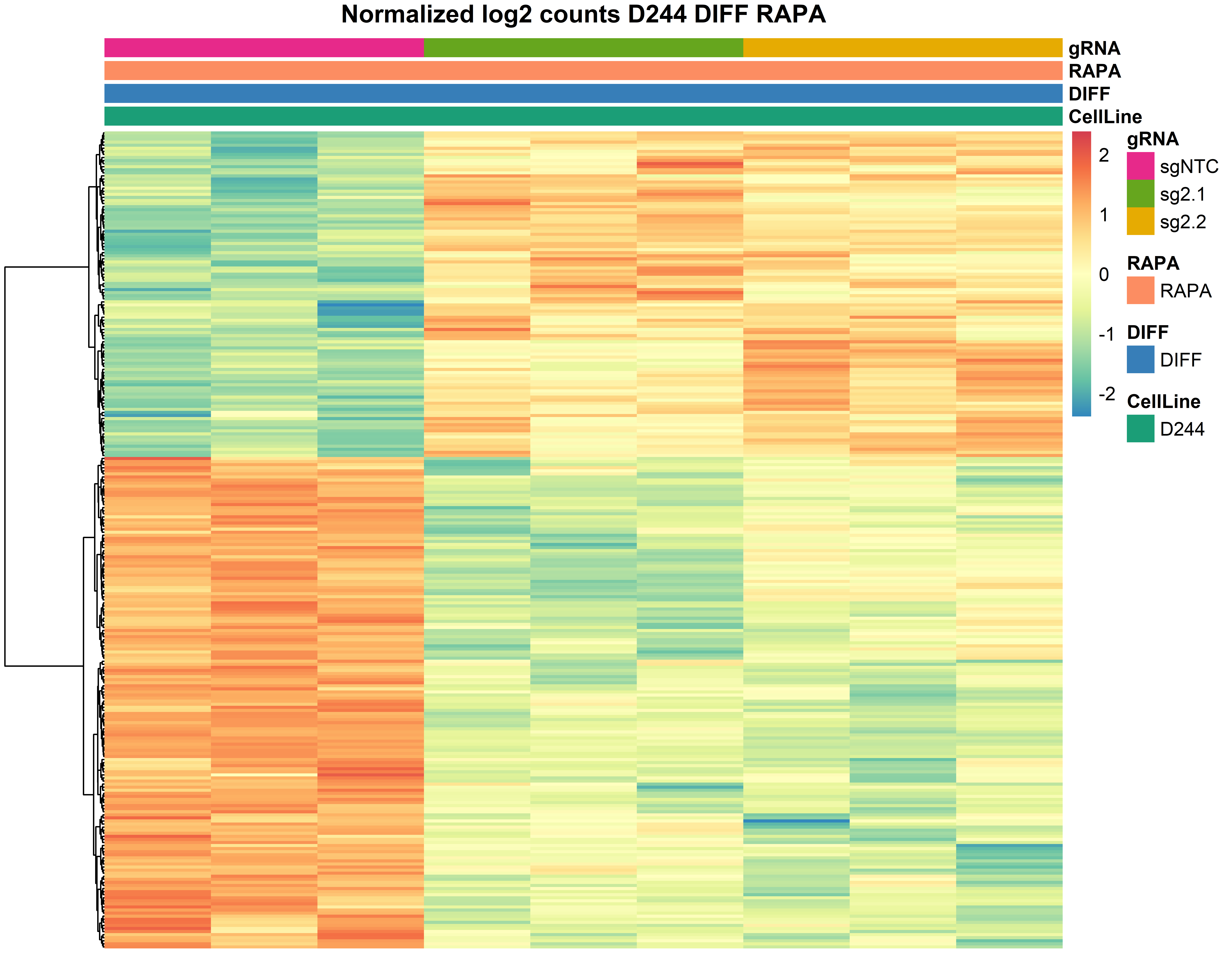
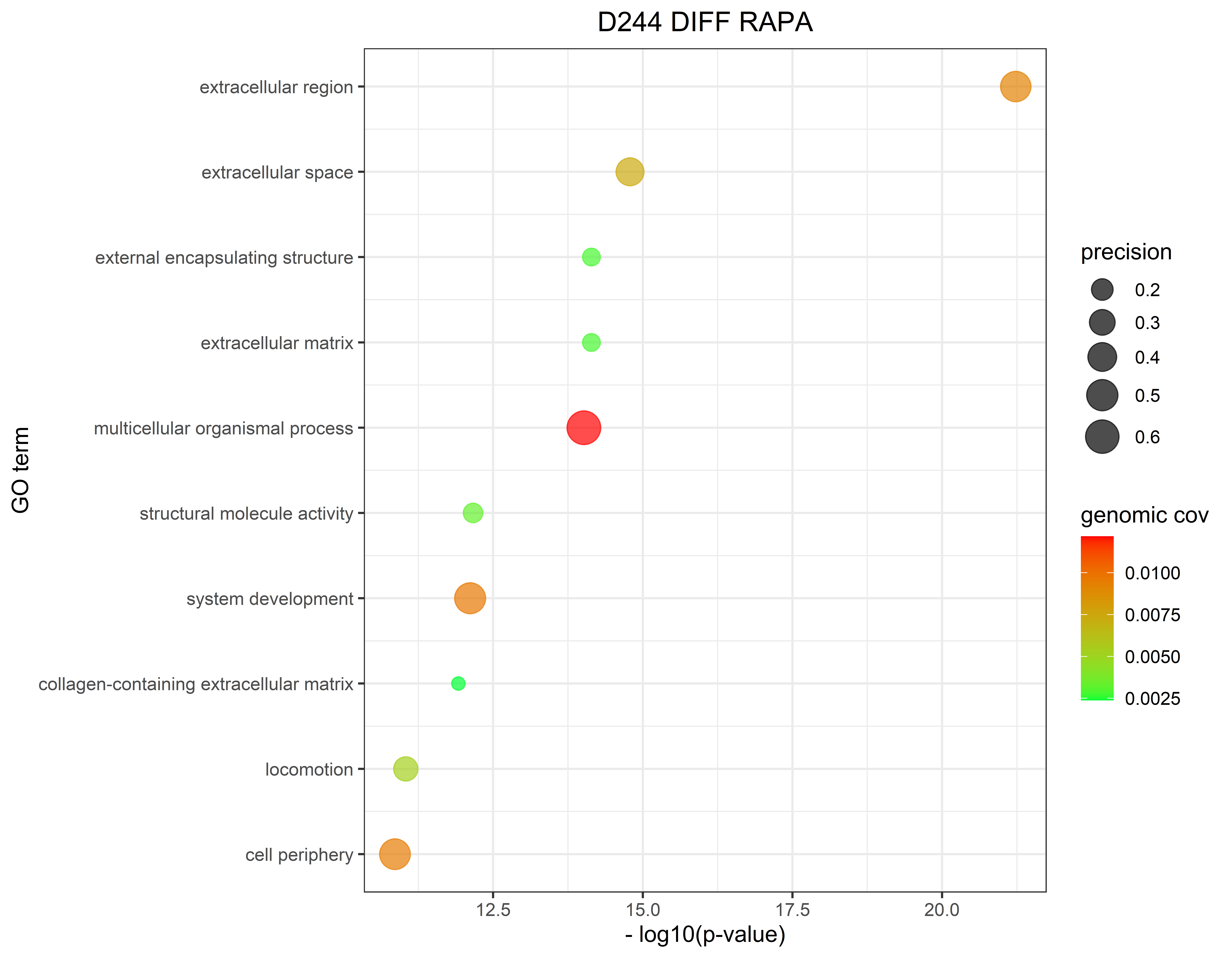
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.0440662 0.52919287 1.870555 8.784975e-01
02_TSC2_KO_Grabole_2016 1.9056788 1.48297727 2.448718 3.122349e-07
03_TSC2_Grabole-2016 1.8786968 1.44836988 2.449813 8.381421e-07
04_TSC2-tuber-Martin2017 6.2255599 3.07274755 11.548446 1.980811e-06
05_TSC2-SEN_SEGA-Martin2017 2.6266379 1.98386578 3.451758 5.114790e-11
06_all_EpilepsyGene 0.6630122 0.21220963 1.581249 4.539375e-01
07_high_conf_EpilepsyGene 0.8171989 0.09734763 3.045438 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 0.4320775 0.01080820 2.476788 7.308581e-01
09_Epi25_dataset 0.0000000 0.00000000 2.328291 4.124445e-01
10_SFARI_all 1.5404546 0.60634143 3.271510 2.347251e-01
11_SFARI_score4plus 1.1705565 0.31300496 3.087989 7.794051e-01
12_DeRubeis_RDNV 0.6557261 0.01632737 3.797089 1.000000e+00
13_Iossifov_RDNV 1.4441535 0.77246671 2.492362 1.885849e-01
14_FMRP_Darnell 1.4716175 0.88915348 2.317771 1.027332e-01
15_Voineagu_M12 0.8973729 0.32395101 2.000942 1.000000e+00
16_Voineagu_M16 3.2821503 1.94389489 5.265616 1.337384e-05
17_ID_genes_Neel 0.6184302 0.16619532 1.618134 5.384706e-01
18_ID_genes_pinto2014 1.5135433 0.59586738 3.213513 2.420997e-01
19_Fromer_SCZ_FunDN 1.2627426 0.63930099 2.265601 4.059027e-01
20_Cocchi_Pruning 2.3423778 0.61946070 6.289339 1.008871e-01
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.04312291 0.3466965 1.000000e+00
02_TSC2_KO_Grabole_2016 0.64483829 0.1279523 6.244698e-06
03_TSC2_Grabole-2016 0.63057836 0.1327243 1.676284e-05
04_TSC2-tuber-Martin2017 1.82866339 0.3602506 3.961622e-05
05_TSC2-SEN_SEGA-Martin2017 0.96570466 0.1431925 1.022958e-09
06_all_EpilepsyGene -0.41096184 0.5812341 1.000000e+00
07_high_conf_EpilepsyGene -0.20187270 1.0855072 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene -0.83915038 1.8817858 1.000000e+00
09_Epi25_dataset -Inf NaN 1.000000e+00
10_SFARI_all 0.43207755 0.4757090 1.000000e+00
11_SFARI_score4plus 0.15747931 0.6729671 1.000000e+00
12_DeRubeis_RDNV -0.42201203 1.8841330 1.000000e+00
13_Iossifov_RDNV 0.36752335 0.3192294 1.000000e+00
14_FMRP_Darnell 0.38636210 0.2570651 1.000000e+00
15_Voineagu_M12 -0.10828382 0.5198363 1.000000e+00
16_Voineagu_M16 1.18849879 0.2672475 2.674769e-04
17_ID_genes_Neel -0.48057094 0.6704187 1.000000e+00
18_ID_genes_pinto2014 0.41445343 0.4756074 1.000000e+00
19_Fromer_SCZ_FunDN 0.23328605 0.3472786 1.000000e+00
20_Cocchi_Pruning 0.85116659 0.6786085 1.000000e+00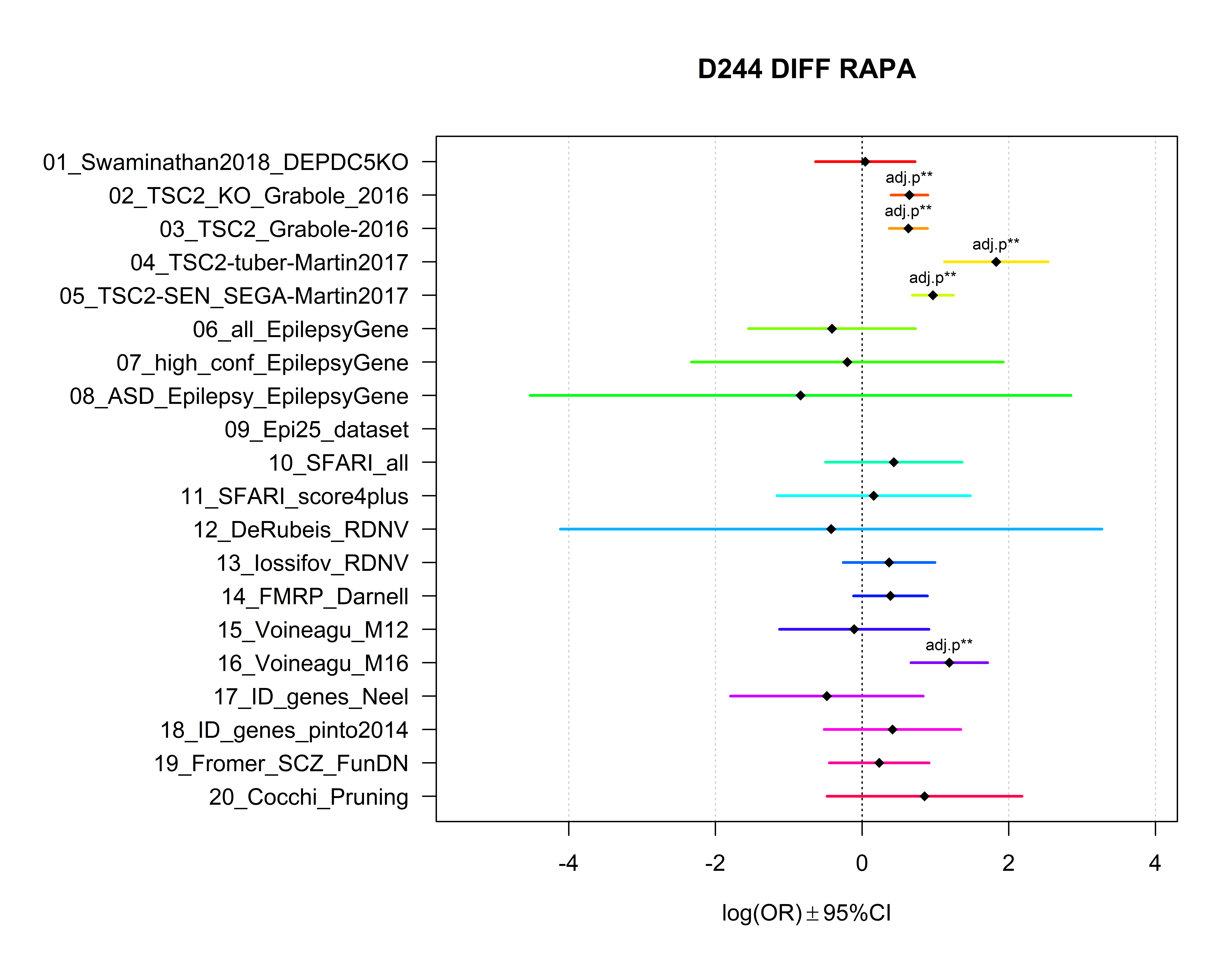
ReN
getlinmodoutput(targetline = "ReN",targetdiff = "DIFF",targetrapa = "RAPA") entrezgene genname log2FC_2.1 fdr_2.1 log2FC_2.2 fdr_2.2
1 25932 CLIC4 -1.3586050 3.608772e-02 -1.8215486 8.231305e-03
2 6281 S100A10 -1.1420986 1.572252e-04 -0.8493016 4.721390e-02
3 10485 MIR9-1HG 3.7161701 3.294323e-30 2.2183209 1.409849e-07
4 63827 BCAN 2.4450337 9.308562e-07 2.1737736 1.359897e-04
5 7044 LEFTY2 -1.1330094 2.120245e-02 -1.3099004 7.931635e-03
6 80760 ITIH5 -2.7415319 3.833229e-02 -4.4070114 1.396392e-03
7 8519 IFITM1 -1.3637673 3.646454e-02 -2.0016739 3.944249e-03
8 10418 SPON1 8.1507287 4.970328e-04 9.7278024 1.256531e-08
9 387763 C11orf96 1.8720845 3.885333e-04 1.8628924 2.863164e-03
10 8076 MFAP5 -2.7489988 8.505600e-07 -2.7429341 2.822997e-05
11 1848 DUSP6 2.2515685 6.816638e-04 2.4201002 3.353992e-04
12 2621 GAS6 -1.9220378 7.170241e-08 -1.6933339 4.166595e-05
13 57447 NDRG2 2.1523934 4.094364e-08 1.8483757 6.910651e-04
14 4053 LTBP2 -2.8637440 1.193540e-03 -3.0589735 4.632015e-03
15 7057 THBS1 -2.3930607 1.151460e-02 -2.7036418 3.229200e-03
16 302 ANXA2 -0.8531947 7.205165e-03 -0.7342226 4.836928e-02
17 728276 CLEC19A -3.5350995 1.157863e-08 -3.0223898 1.867448e-03
18 1277 COL1A1 -3.1711128 4.294311e-17 -3.7120488 3.370409e-18
19 9021 SOCS3 -2.3976598 1.162410e-10 -1.3447580 2.570643e-02
20 114897 C1QTNF1 -2.0752249 3.520626e-05 -1.4310485 3.285384e-02
21 1463 NCAN 1.5037952 6.339942e-12 1.0406722 2.853093e-03
22 1545 CYP1B1 -3.5604784 8.161597e-08 -2.3708176 3.071266e-05
23 72 ACTG2 -2.5773756 7.368089e-17 -2.2067885 1.614029e-08
24 2335 FN1 -3.2251369 2.439609e-09 -4.0660517 1.970600e-15
25 3488 IGFBP5 -1.5452752 8.161597e-08 -1.4546640 3.094511e-06
26 55502 HES6 2.6453784 3.656844e-17 1.7604937 1.138068e-04
27 24141 LAMP5 1.6018892 1.192717e-02 1.5994225 2.425320e-02
28 1291 COL6A1 -1.3459432 3.235914e-05 -1.3121749 4.726705e-05
29 1292 COL6A2 -2.0253143 2.951469e-02 -2.9808726 1.138068e-04
30 10675 CSPG5 2.2364472 4.596981e-11 1.6496521 8.761854e-05
31 11170 FAM107A 2.1586119 2.320013e-05 1.9427053 8.738364e-04
32 56999 ADAMTS9 3.0400577 4.547679e-02 3.3821872 4.836928e-02
33 1295 COL8A1 -3.1075914 1.715765e-06 -2.8215083 6.250435e-08
34 5648 MASP1 7.8413690 2.008683e-03 7.9383795 3.454438e-03
35 683 BST1 -2.8950449 1.610867e-04 -2.5727092 2.357743e-03
36 23231 SEL1L3 1.5780452 2.444935e-02 1.9759758 2.780093e-03
37 4015 LOX -2.2360570 1.501354e-04 -2.0976670 3.229200e-03
38 9547 CXCL14 -2.5651405 1.134080e-06 -2.2279800 5.293257e-04
39 6678 SPARC -1.1273443 7.922301e-05 -0.9770203 3.229200e-03
40 57451 TENM2 -1.9941895 4.787436e-02 -2.7476467 5.935520e-04
41 51617 NSG2 10.2491172 1.312034e-10 8.6800156 5.616583e-05
42 10590 SCGN 3.6842321 6.432714e-25 1.7929381 7.252548e-03
43 2173 FABP7 0.8660790 1.736082e-02 0.9232129 2.223532e-02
44 1490 CCN2 -2.3098104 1.198764e-07 -1.3937364 4.270771e-02
45 9865 TRIL 2.7217711 1.994616e-02 2.8858121 3.285384e-02
46 3486 IGFBP3 -4.6468508 6.357079e-15 -3.3826860 1.256531e-08
47 1278 COL1A2 -3.8996895 2.345305e-07 -3.5277248 2.940563e-06
48 4747 NEFL -2.0357376 2.370999e-05 -2.6024981 4.420266e-06
49 51050 PI15 -1.0151690 2.950165e-02 -1.3134282 3.229200e-03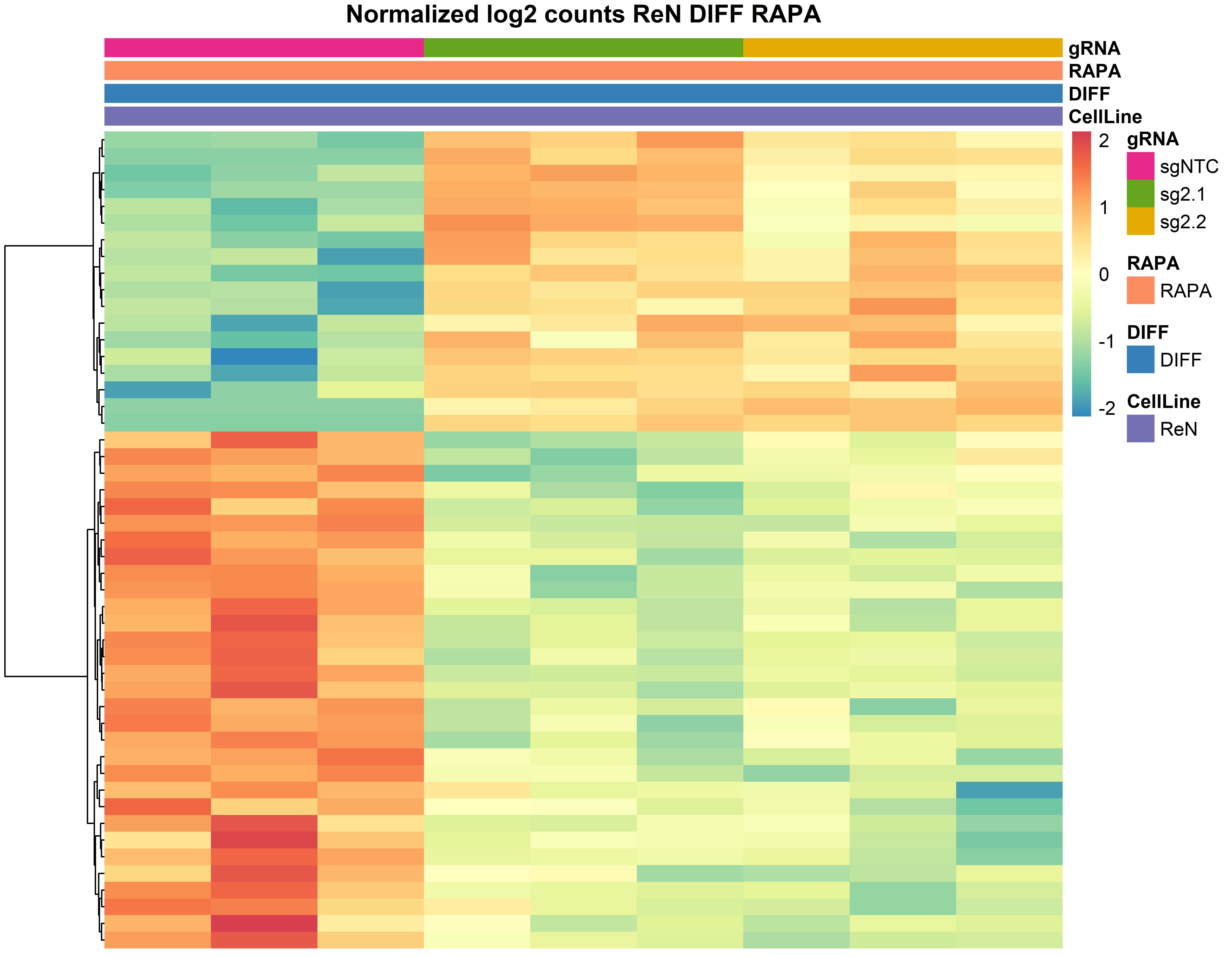
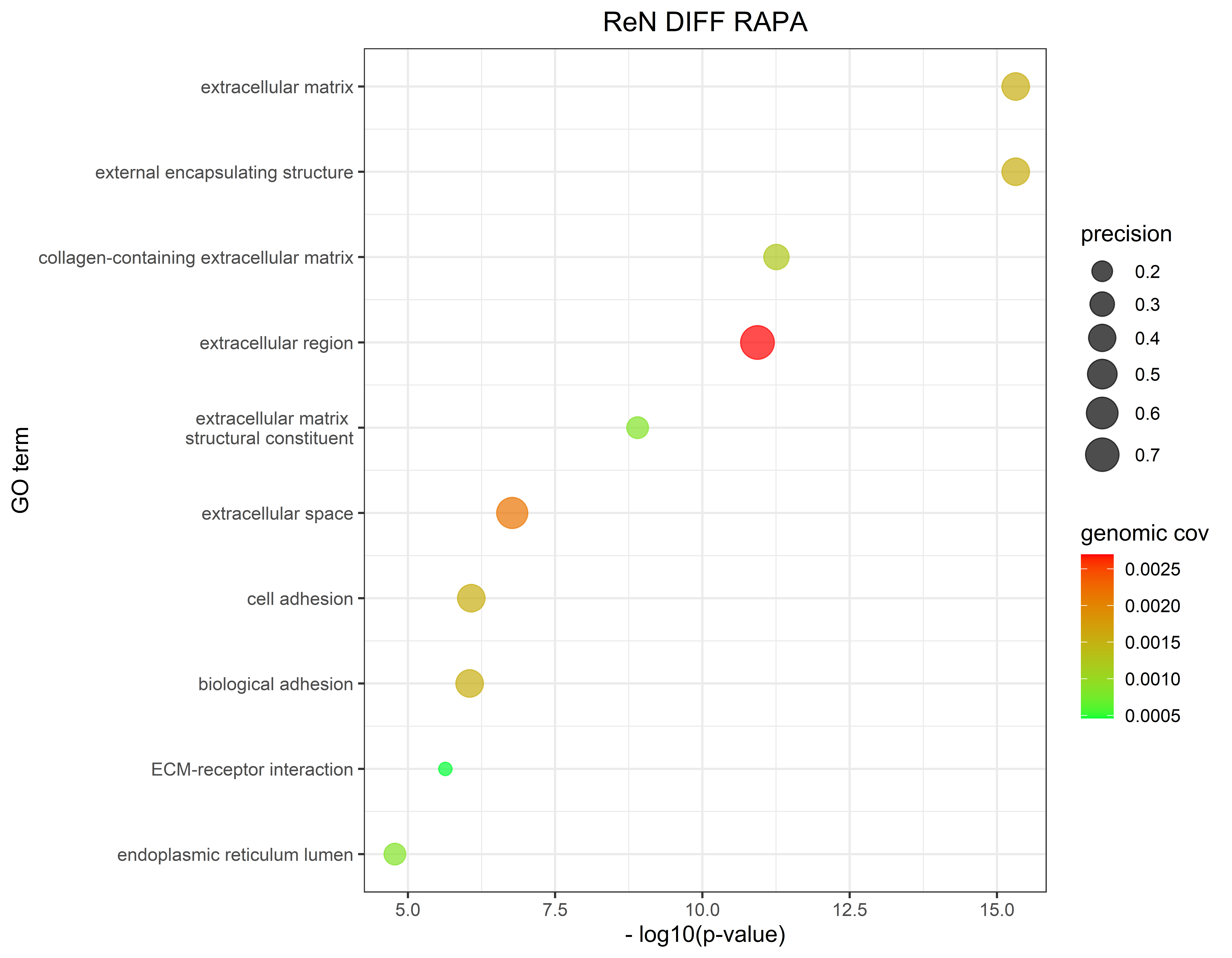
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.9393747 0.11026176 3.602544 1.000000e+00
02_TSC2_KO_Grabole_2016 2.1315004 1.16761531 3.914827 9.893612e-03
03_TSC2_Grabole-2016 2.7728774 1.43445011 5.703361 1.375979e-03
04_TSC2-tuber-Martin2017 8.0077608 1.56951307 25.514772 7.688118e-03
05_TSC2-SEN_SEGA-Martin2017 3.8659302 2.05161168 7.112128 1.819302e-05
06_all_EpilepsyGene 1.4850928 0.17407594 5.708719 3.987910e-01
07_high_conf_EpilepsyGene 0.0000000 0.00000000 8.556545 1.000000e+00
08_ASD_Epilepsy_EpilepsyGene 0.0000000 0.00000000 9.160823 1.000000e+00
09_Epi25_dataset 0.0000000 0.00000000 13.234037 1.000000e+00
10_SFARI_all 1.1758695 0.02904876 6.935834 5.779460e-01
11_SFARI_score4plus 0.0000000 0.00000000 6.013472 1.000000e+00
12_DeRubeis_RDNV 0.0000000 0.00000000 13.942207 1.000000e+00
13_Iossifov_RDNV 0.5360707 0.01327500 3.146750 1.000000e+00
14_FMRP_Darnell 1.9397821 0.59851826 4.899481 1.947059e-01
15_Voineagu_M12 0.0000000 0.00000000 3.051384 6.361856e-01
16_Voineagu_M16 4.4507001 1.36869328 11.296858 7.551960e-03
17_ID_genes_Neel 0.0000000 0.00000000 3.197407 6.327798e-01
18_ID_genes_pinto2014 0.0000000 0.00000000 4.352008 1.000000e+00
19_Fromer_SCZ_FunDN 3.0422422 0.93731156 7.698986 3.155365e-02
20_Cocchi_Pruning 10.0473664 1.96199349 32.213643 4.188780e-03
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -0.06254084 1.0930394 1.0000000000
02_TSC2_KO_Grabole_2016 0.75682617 0.3070728 0.1978722388
03_TSC2_Grabole-2016 1.01988556 0.3362775 0.0275195782
04_TSC2-tuber-Martin2017 2.08041118 0.8314519 0.1537623625
05_TSC2-SEN_SEGA-Martin2017 1.35220232 0.3232534 0.0003638605
06_all_EpilepsyGene 0.39547727 1.0937454 1.0000000000
07_high_conf_EpilepsyGene -Inf NaN 1.0000000000
08_ASD_Epilepsy_EpilepsyGene -Inf NaN 1.0000000000
09_Epi25_dataset -Inf NaN 1.0000000000
10_SFARI_all 0.16200790 1.8881568 1.0000000000
11_SFARI_score4plus -Inf NaN 1.0000000000
12_DeRubeis_RDNV -Inf NaN 1.0000000000
13_Iossifov_RDNV -0.62348920 1.8869303 1.0000000000
14_FMRP_Darnell 0.66257562 0.5999356 1.0000000000
15_Voineagu_M12 -Inf NaN 1.0000000000
16_Voineagu_M16 1.49306141 0.6016352 0.1510391926
17_ID_genes_Neel -Inf NaN 1.0000000000
18_ID_genes_pinto2014 -Inf NaN 1.0000000000
19_Fromer_SCZ_FunDN 1.11259483 0.6006808 0.6310729981
20_Cocchi_Pruning 2.30731055 0.8333416 0.0837756077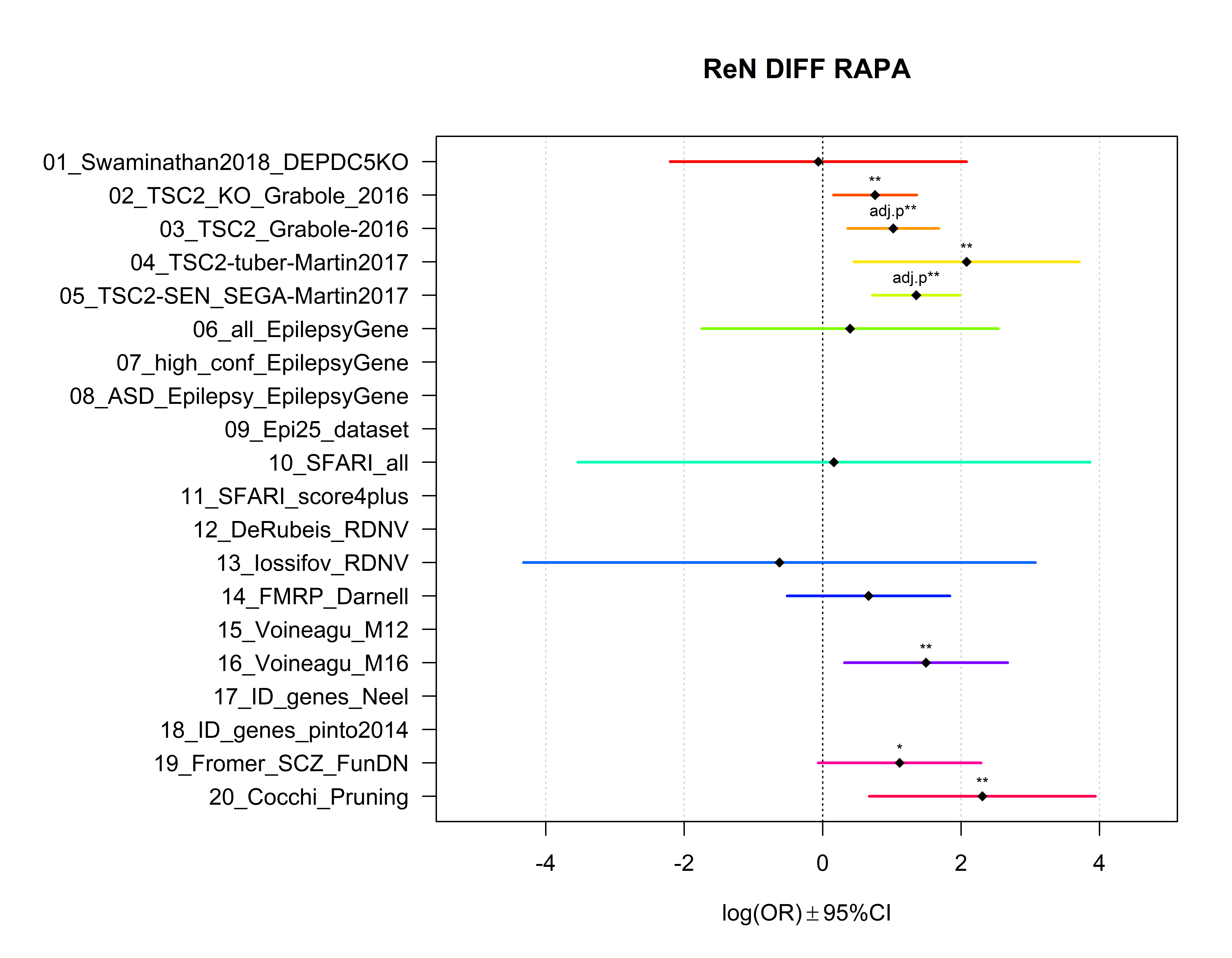
Replicated across cell lines
getlinmodoutput_all = function(targetdiff,
targetrapa){
samplesincl = SampleInfo$DIFF==targetdiff & SampleInfo$RAPA==targetrapa
pvalrep= get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"ReN_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"ReN_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj<mypval
betarep = apply(cbind(get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"ReN_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"ReN_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange),
1,
samesign)
idx=which(betarep & pvalrep)
hits=geneids[idx]
restab=data.frame(
entrezgene = geneids[idx],
genname=rowData(ddsMat)[hits,"hgnc"],
D244_log2FC_2.1 = get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D244_fdr_2.1 = get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj[idx],
D244_log2FC_2.2 = get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D244_fdr_2.2 = get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj[idx],
D62_log2FC_2.1 = get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D62_fdr_2.1 = get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj[idx],
D62_log2FC_2.2 = get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D62_fdr_2.2 = get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj[idx],
ReN_log2FC_2.1 = get(paste0("restab_",
"ReN_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
ReN_fdr_2.1 = get(paste0("restab_",
"ReN_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj[idx],
ReN_log2FC_2.2 = get(paste0("restab_",
"ReN_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
ReN_fdr_2.2 = get(paste0("restab_",
"ReN_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj[idx])
print(restab)
write.xlsx(restab, file=paste0(output, "/Restab_",
"Repl_All_",
targetdiff, "_",
targetrapa, ".xlsx"))
SamplesSet=SampleInfo[samplesincl,]
plotmatrix = log_2cpm[hits,rownames(SamplesSet)]
rownames(SamplesSet)=SamplesSet$label_rep
colnames(plotmatrix)=SamplesSet$label_rep
rownames(plotmatrix)=rowData(ddsMat)[hits,"hgnc"]
cellcol = Dark8[1:nlevels(SampleInfo$CellLine)]
names(cellcol) = levels(SampleInfo$CellLine)
cellcol = cellcol[as.character(unique(SamplesSet$CellLine))]
gRNAcol = Dark8[c(1:nlevels(SampleInfo$gRNA))+nlevels(SampleInfo$CellLine)]
names(gRNAcol) = levels(SampleInfo$gRNA)
gRNAcol = gRNAcol[as.character(unique(SamplesSet$gRNA))]
diffcol = brewer.pal(3,"Set1")[1:nlevels(SampleInfo$DIFF)]
names(diffcol) = levels(SampleInfo$DIFF)
diffcol = diffcol[as.character(unique(SamplesSet$DIFF))]
rapacol = brewer.pal(3,"Set2")[1:nlevels(SampleInfo$RAPA)]
names(rapacol) = levels(SampleInfo$RAPA)
rapacol = rapacol[as.character(unique(SamplesSet$RAPA))]
ann_colors = list(
DIFF = diffcol,
RAPA = rapacol,
gRNA = gRNAcol,
CellLine=cellcol)
collabels = SamplesSet[,c("CellLine","DIFF","RAPA", "gRNA")] %>%
mutate_all(as.character) %>% as.data.frame()
rownames(collabels)=SamplesSet$label_rep
idx=order(SamplesSet$CellLine, SamplesSet$gRNA)
pheatmap(plotmatrix[,idx],
border_color = NA,
annotation_col = collabels[idx,],show_rownames = F, show_colnames = F,
annotation_colors = ann_colors,
clustering_method = "ward.D2",
cluster_cols = F,
col = colors,
scale = "row",
main = "Normalized log2 counts PROLIF noRAPA")
gene_univers = rownames(ddsMat)
gostres = getGOresults(hits, gene_univers)
toptab = gostres$result
write.xlsx2(toptab, file = paste0(output, "/GOres_",targetdiff,targetrapa,".xlsx"), sheetName = "GO_enrichment")
idx = grepl("GO|KEGG", toptab$source)
titlename = paste(targetdiff,targetrapa)
if(!any(idx)){
p = ggplot() + annotate("text", x = 4, y = 25, size=4,
label = "no significant GO term") +
ggtitle(titlename)+theme_void()+
theme(plot.title = element_text(hjust = 0.5))
} else {
p=GOplot(toptab[idx, ], 10, Title = titlename)
}
print(p)
if(length(hits)>=1){
Resall=data.frame()
for (gl in names(genelists)){
tmp=table(signif=geneids %in% hits, targetgene=geneids %in% genelists[[gl]]$ENTREZ_ID)
resfish=fisher.test(tmp)
res = c(resfish$estimate, unlist(resfish$conf.int), resfish$p.value)
Resall = rbind(Resall, res)
}
colnames(Resall)=c("OR", "CI95L", "CI95U", "P")
rownames(Resall)=names(genelists)
Resall$Beta = log(Resall$OR)
Resall$SE = (log(Resall$OR)-log(Resall$CI95L))/1.96
Resall$Padj=p.adjust(Resall$P, method = "bonferroni")
print(Resall)
multiORplot(Resall, Pval = "P", Padj = "Padj", beta="Beta",SE = "SE",
pheno=paste(targetdiff, targetrapa))
}
}noDiff noRapa
getlinmodoutput_all(targetdiff = "noDIFF",targetrapa = "noRAPA") entrezgene genname D244_log2FC_2.1 D244_fdr_2.1 D244_log2FC_2.2 D244_fdr_2.2
1 151887 CCDC80 0.8069456 2.122272e-15 0.7595701 3.271995e-14
2 9568 GABBR2 1.0262131 4.391028e-02 1.8380988 4.032745e-07
D62_log2FC_2.1 D62_fdr_2.1 D62_log2FC_2.2 D62_fdr_2.2 ReN_log2FC_2.1
1 0.7861593 3.884032e-08 0.544454 5.158658e-06 1.3269840
2 1.8836534 3.992926e-19 1.770097 1.993618e-26 0.8591791
ReN_fdr_2.1 ReN_log2FC_2.2 ReN_fdr_2.2
1 1.910378e-09 0.5349808 2.674684e-06
2 1.299630e-04 0.7804877 1.030234e-16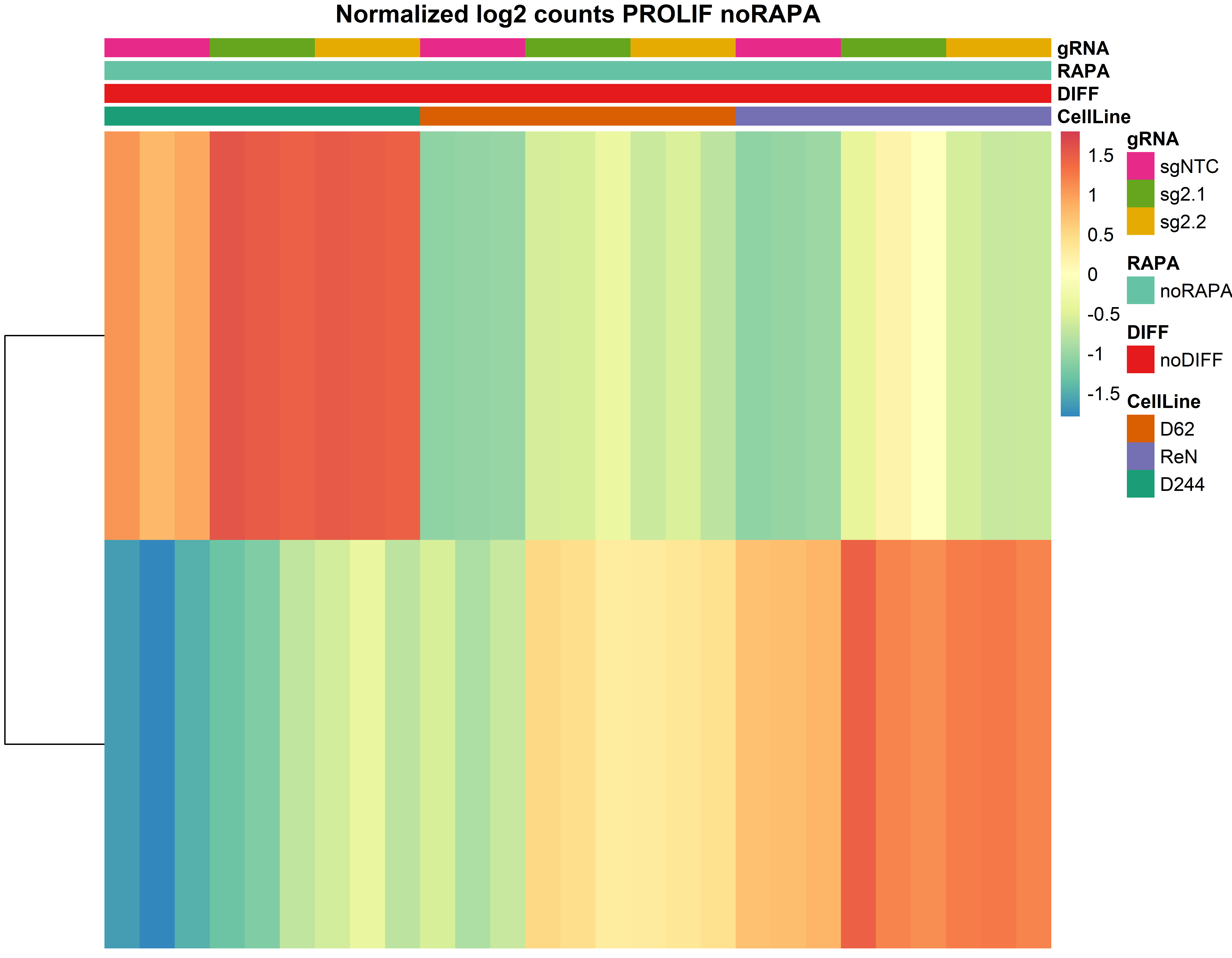
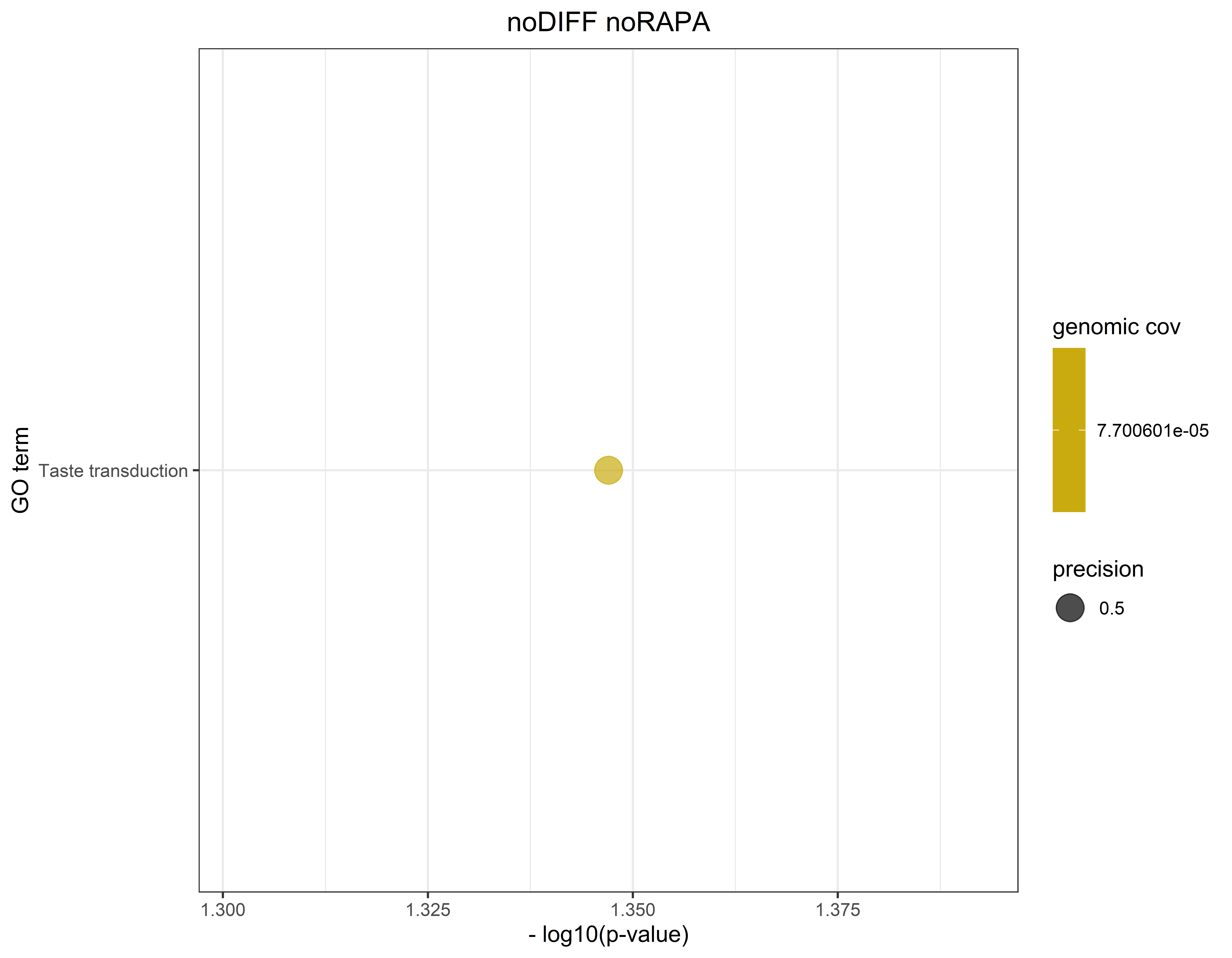
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.0000000 0.00000000 117.67597 1.0000000
02_TSC2_KO_Grabole_2016 1.8802655 0.02394853 147.44493 1.0000000
03_TSC2_Grabole-2016 0.9979352 0.01271854 78.30124 1.0000000
04_TSC2-tuber-Martin2017 0.0000000 0.00000000 638.40262 1.0000000
05_TSC2-SEN_SEGA-Martin2017 6.0612670 0.07719326 474.01693 0.2632349
06_all_EpilepsyGene 0.0000000 0.00000000 185.81492 1.0000000
07_high_conf_EpilepsyGene 0.0000000 0.00000000 574.61522 1.0000000
08_ASD_Epilepsy_EpilepsyGene 0.0000000 0.00000000 615.67354 1.0000000
09_Epi25_dataset 0.0000000 0.00000000 893.42035 1.0000000
10_SFARI_all 0.0000000 0.00000000 301.07655 1.0000000
11_SFARI_score4plus 0.0000000 0.00000000 412.02988 1.0000000
12_DeRubeis_RDNV 0.0000000 0.00000000 938.93111 1.0000000
13_Iossifov_RDNV 0.0000000 0.00000000 137.38433 1.0000000
14_FMRP_Darnell 17.0257674 0.21675750 1321.33550 0.1079356
15_Voineagu_M12 0.0000000 0.00000000 207.85081 1.0000000
16_Voineagu_M16 0.0000000 0.00000000 206.59806 1.0000000
17_ID_genes_Neel 0.0000000 0.00000000 217.73964 1.0000000
18_ID_genes_pinto2014 0.0000000 0.00000000 295.96285 1.0000000
19_Fromer_SCZ_FunDN 0.0000000 0.00000000 141.75165 1.0000000
20_Cocchi_Pruning 0.0000000 0.00000000 806.47010 1.0000000
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -Inf NaN 1
02_TSC2_KO_Grabole_2016 0.631412991 2.226154 1
03_TSC2_Grabole-2016 -0.002066912 2.225830 1
04_TSC2-tuber-Martin2017 -Inf NaN 1
05_TSC2-SEN_SEGA-Martin2017 1.801918851 2.226205 1
06_all_EpilepsyGene -Inf NaN 1
07_high_conf_EpilepsyGene -Inf NaN 1
08_ASD_Epilepsy_EpilepsyGene -Inf NaN 1
09_Epi25_dataset -Inf NaN 1
10_SFARI_all -Inf NaN 1
11_SFARI_score4plus -Inf NaN 1
12_DeRubeis_RDNV -Inf NaN 1
13_Iossifov_RDNV -Inf NaN 1
14_FMRP_Darnell 2.834727924 2.226380 1
15_Voineagu_M12 -Inf NaN 1
16_Voineagu_M16 -Inf NaN 1
17_ID_genes_Neel -Inf NaN 1
18_ID_genes_pinto2014 -Inf NaN 1
19_Fromer_SCZ_FunDN -Inf NaN 1
20_Cocchi_Pruning -Inf NaN 1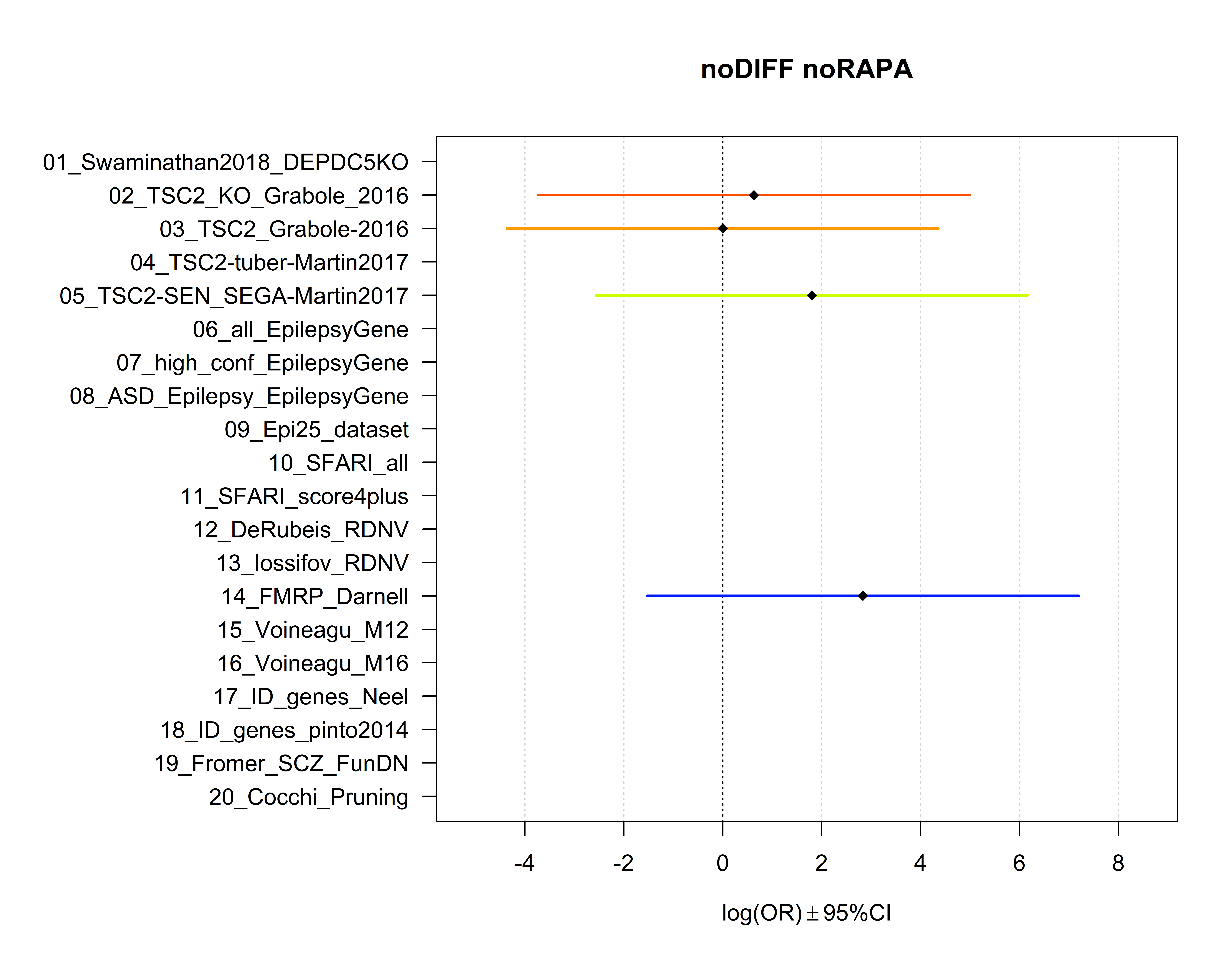
noDiff Rapa
try(getlinmodoutput_all(targetdiff = "noDIFF",targetrapa = "RAPA")) [1] entrezgene genname D244_log2FC_2.1 D244_fdr_2.1
[5] D244_log2FC_2.2 D244_fdr_2.2 D62_log2FC_2.1 D62_fdr_2.1
[9] D62_log2FC_2.2 D62_fdr_2.2 ReN_log2FC_2.1 ReN_fdr_2.1
[13] ReN_log2FC_2.2 ReN_fdr_2.2
<0 Zeilen> (oder row.names mit Länge 0)
Error in mapply(setCellValue, cells[seq_len(nrow(cells)), colIndex[ic]], :
Eingaben mit Länge 0 können nicht mit Eingaben anderer Länge gemischt werdenDiff noRapa
try(getlinmodoutput_all(targetdiff = "DIFF",targetrapa = "noRAPA")) [1] entrezgene genname D244_log2FC_2.1 D244_fdr_2.1
[5] D244_log2FC_2.2 D244_fdr_2.2 D62_log2FC_2.1 D62_fdr_2.1
[9] D62_log2FC_2.2 D62_fdr_2.2 ReN_log2FC_2.1 ReN_fdr_2.1
[13] ReN_log2FC_2.2 ReN_fdr_2.2
<0 Zeilen> (oder row.names mit Länge 0)
Error in mapply(setCellValue, cells[seq_len(nrow(cells)), colIndex[ic]], :
Eingaben mit Länge 0 können nicht mit Eingaben anderer Länge gemischt werdenDiff Rapa
try(getlinmodoutput_all(targetdiff = "DIFF",targetrapa = "RAPA")) entrezgene genname D244_log2FC_2.1 D244_fdr_2.1 D244_log2FC_2.2 D244_fdr_2.2
1 7044 LEFTY2 -1.3192934 3.280549e-18 -2.3110081 5.751049e-64
2 3488 IGFBP5 -1.4790171 1.730247e-60 -1.3238131 6.039310e-53
3 1295 COL8A1 -2.5667320 1.117183e-51 -2.8067329 2.533751e-78
4 9547 CXCL14 -0.5235165 1.148369e-10 -0.4632759 1.335559e-07
D62_log2FC_2.1 D62_fdr_2.1 D62_log2FC_2.2 D62_fdr_2.2 ReN_log2FC_2.1
1 -1.146146 0.0151783820 -2.510127 2.217413e-06 -1.133009
2 -1.204226 0.0014575086 -1.284033 3.354261e-03 -1.545275
3 -4.547185 0.0071613815 -3.737934 2.815452e-02 -3.107591
4 -0.806434 0.0007756694 -1.501844 1.611197e-09 -2.565140
ReN_fdr_2.1 ReN_log2FC_2.2 ReN_fdr_2.2
1 2.120245e-02 -1.309900 7.931635e-03
2 8.161597e-08 -1.454664 3.094511e-06
3 1.715765e-06 -2.821508 6.250435e-08
4 1.134080e-06 -2.227980 5.293257e-04No results to show
Please make sure that the organism is correct or set significant = FALSE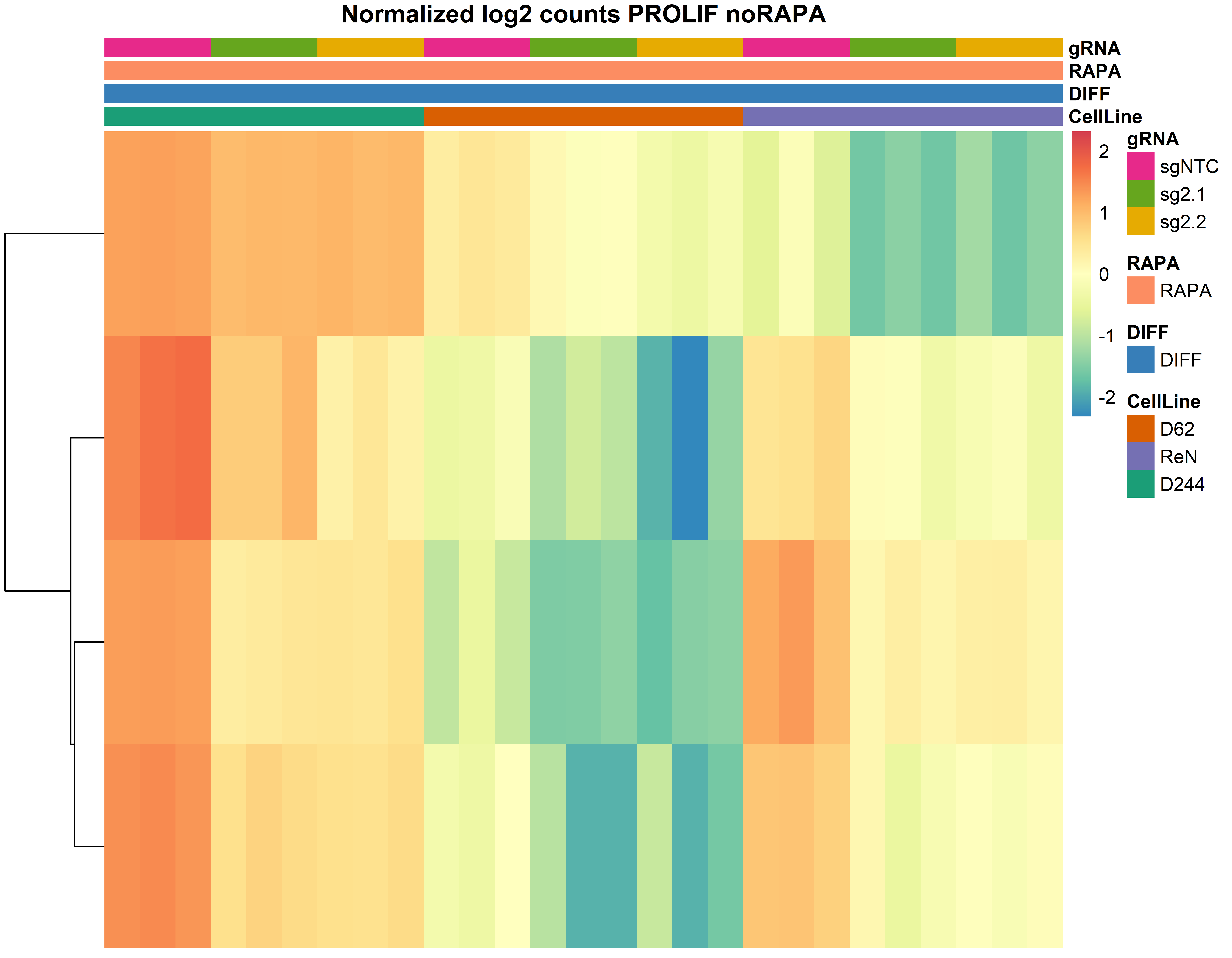
[1] "no significant results"
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.0000000 0.000000000 33.504610 1.00000000
02_TSC2_KO_Grabole_2016 0.6266413 0.011938369 7.807027 1.00000000
03_TSC2_Grabole-2016 0.3325514 0.006338702 4.143006 0.37442864
04_TSC2-tuber-Martin2017 0.0000000 0.000000000 183.665630 1.00000000
05_TSC2-SEN_SEGA-Martin2017 6.0639067 0.439326230 83.495306 0.09882533
06_all_EpilepsyGene 0.0000000 0.000000000 52.915563 1.00000000
07_high_conf_EpilepsyGene 0.0000000 0.000000000 165.829831 1.00000000
08_ASD_Epilepsy_EpilepsyGene 0.0000000 0.000000000 177.312232 1.00000000
09_Epi25_dataset 0.0000000 0.000000000 254.557976 1.00000000
10_SFARI_all 0.0000000 0.000000000 85.956736 1.00000000
11_SFARI_score4plus 0.0000000 0.000000000 116.219641 1.00000000
12_DeRubeis_RDNV 0.0000000 0.000000000 268.001224 1.00000000
13_Iossifov_RDNV 0.0000000 0.000000000 39.101432 1.00000000
14_FMRP_Darnell 0.0000000 0.000000000 25.798317 1.00000000
15_Voineagu_M12 0.0000000 0.000000000 59.261346 1.00000000
16_Voineagu_M16 0.0000000 0.000000000 58.898397 1.00000000
17_ID_genes_Neel 0.0000000 0.000000000 61.903070 1.00000000
18_ID_genes_pinto2014 0.0000000 0.000000000 84.469151 1.00000000
19_Fromer_SCZ_FunDN 0.0000000 0.000000000 40.336332 1.00000000
20_Cocchi_Pruning 0.0000000 0.000000000 228.948566 1.00000000
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -Inf NaN 1
02_TSC2_KO_Grabole_2016 -0.4673809 2.020723 1
03_TSC2_Grabole-2016 -1.1009609 2.020470 1
04_TSC2-tuber-Martin2017 -Inf NaN 1
05_TSC2-SEN_SEGA-Martin2017 1.8023543 1.339218 1
06_all_EpilepsyGene -Inf NaN 1
07_high_conf_EpilepsyGene -Inf NaN 1
08_ASD_Epilepsy_EpilepsyGene -Inf NaN 1
09_Epi25_dataset -Inf NaN 1
10_SFARI_all -Inf NaN 1
11_SFARI_score4plus -Inf NaN 1
12_DeRubeis_RDNV -Inf NaN 1
13_Iossifov_RDNV -Inf NaN 1
14_FMRP_Darnell -Inf NaN 1
15_Voineagu_M12 -Inf NaN 1
16_Voineagu_M16 -Inf NaN 1
17_ID_genes_Neel -Inf NaN 1
18_ID_genes_pinto2014 -Inf NaN 1
19_Fromer_SCZ_FunDN -Inf NaN 1
20_Cocchi_Pruning -Inf NaN 1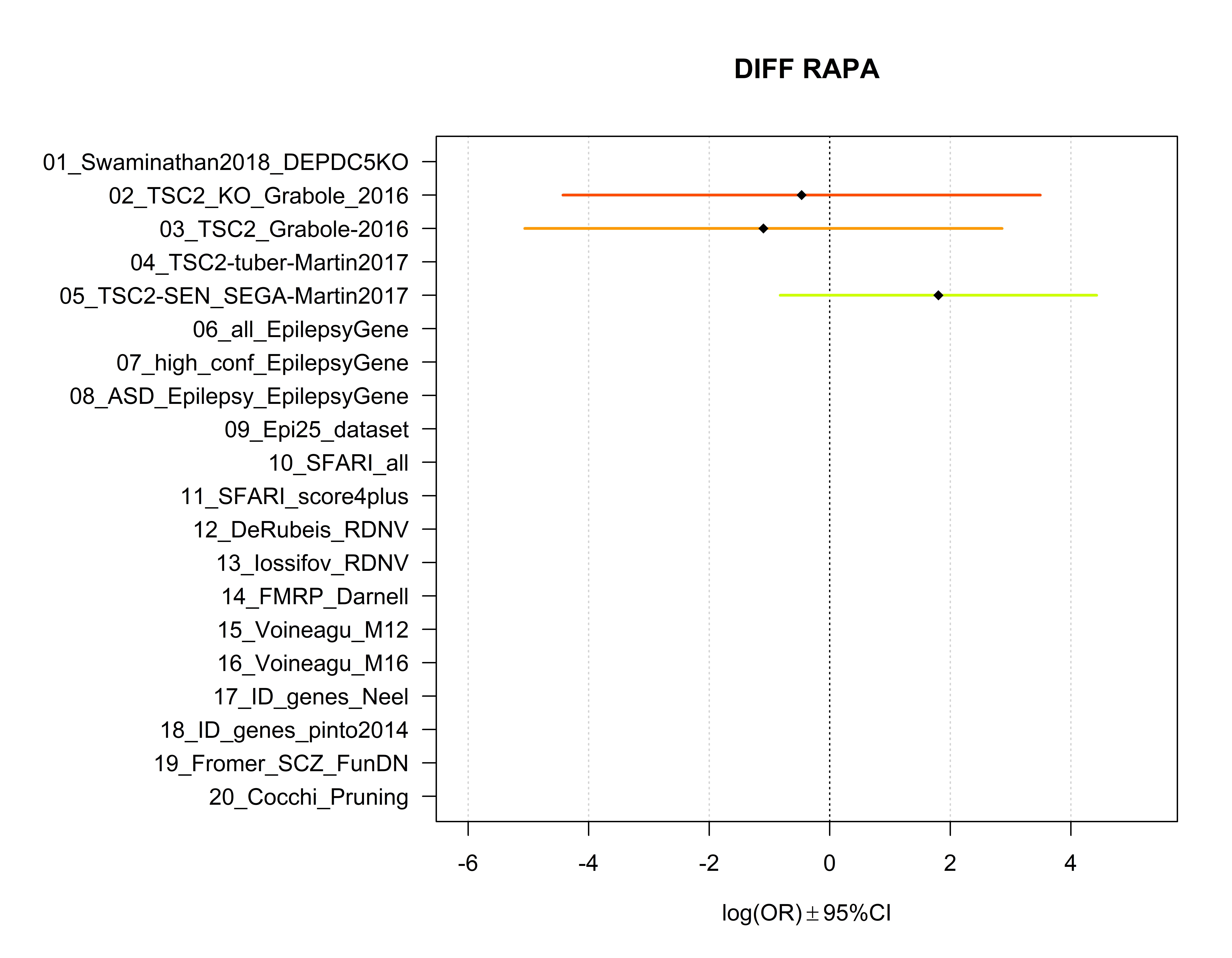
Replicated across primary-HNPC cell lines
getlinmodoutput_D62D224 = function(targetdiff,
targetrapa, targetline){
samplesincl = SampleInfo$DIFF==targetdiff &
SampleInfo$RAPA==targetrapa &
SampleInfo$CellLine %in% targetline
pvalrep= get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj<mypval &
get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj<mypval
betarep = apply(cbind(get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange,
get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange),
1,
samesign)
idx=which(betarep & pvalrep)
hits=geneids[idx]
restab=data.frame(
entrezgene = geneids[idx],
genname=rowData(ddsMat)[hits,"hgnc"],
D244_log2FC_2.1 = get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D244_fdr_2.1 = get(paste0("restab_",
"D244_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj[idx],
D244_log2FC_2.2 = get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D244_fdr_2.2 = get(paste0("restab_",
"D244_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj[idx],
D62_log2FC_2.1 = get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D62_fdr_2.1 = get(paste0("restab_",
"D62_sgNTC_sg2.1_",
targetdiff, "_" ,
targetrapa))$padj[idx],
D62_log2FC_2.2 = get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$log2FoldChange[idx],
D62_fdr_2.2 = get(paste0("restab_",
"D62_sgNTC_sg2.2_",
targetdiff, "_" ,
targetrapa))$padj[idx])
print(restab)
write.xlsx(restab, file=paste0(output, "/Restab_",
"Repl_D62D244_",
targetdiff, "_",
targetrapa, ".xlsx"))
SamplesSet=SampleInfo[samplesincl,]
plotmatrix = log_2cpm[hits,rownames(SamplesSet)]
rownames(SamplesSet)=SamplesSet$label_rep
colnames(plotmatrix)=SamplesSet$label_rep
rownames(plotmatrix)=rowData(ddsMat)[hits,"hgnc"]
cellcol = Dark8[1:nlevels(SampleInfo$CellLine)]
names(cellcol) = levels(SampleInfo$CellLine)
cellcol = cellcol[as.character(unique(SamplesSet$CellLine))]
gRNAcol = Dark8[c(1:nlevels(SampleInfo$gRNA))+nlevels(SampleInfo$CellLine)]
names(gRNAcol) = levels(SampleInfo$gRNA)
gRNAcol = gRNAcol[as.character(unique(SamplesSet$gRNA))]
diffcol = brewer.pal(3,"Set1")[1:nlevels(SampleInfo$DIFF)]
names(diffcol) = levels(SampleInfo$DIFF)
diffcol = diffcol[as.character(unique(SamplesSet$DIFF))]
rapacol = brewer.pal(3,"Set2")[1:nlevels(SampleInfo$RAPA)]
names(rapacol) = levels(SampleInfo$RAPA)
rapacol = rapacol[as.character(unique(SamplesSet$RAPA))]
ann_colors = list(
DIFF = diffcol,
RAPA = rapacol,
gRNA = gRNAcol,
CellLine=cellcol)
collabels = SamplesSet[,c("CellLine","DIFF","RAPA", "gRNA")] %>%
mutate_all(as.character) %>% as.data.frame()
rownames(collabels)=SamplesSet$label_rep
idx=order(SamplesSet$CellLine, SamplesSet$gRNA)
pheatmap(plotmatrix[,idx],
border_color = NA,
annotation_col = collabels[idx,],show_rownames = F, show_colnames = F,
annotation_colors = ann_colors,
clustering_method = "ward.D2",
cluster_cols = F,
col = colors,
scale = "row",
main = "Normalized log2 counts PROLIF noRAPA")
gene_univers = rownames(ddsMat)
gostres = getGOresults(hits, gene_univers)
toptab = gostres$result
write.xlsx2(toptab, file = paste0(output, "/GOres_",targetdiff,targetrapa,".xlsx"), sheetName = "GO_enrichment")
idx = grepl("GO|KEGG", toptab$source)
titlename = paste(targetdiff,targetrapa)
if(!any(idx)){
p = ggplot() + annotate("text", x = 4, y = 25, size=4,
label = "no significant GO term") +
ggtitle(titlename)+theme_void()+
theme(plot.title = element_text(hjust = 0.5))
} else {
p=GOplot(toptab[idx, ], 10, Title = titlename)
}
print(p)
if(length(hits)>=1){
Resall=data.frame()
for (gl in names(genelists)){
tmp=table(signif=geneids %in% hits, targetgene=geneids %in% genelists[[gl]]$ENTREZ_ID)
resfish=fisher.test(tmp)
res = c(resfish$estimate, unlist(resfish$conf.int), resfish$p.value)
Resall = rbind(Resall, res)
}
colnames(Resall)=c("OR", "CI95L", "CI95U", "P")
rownames(Resall)=names(genelists)
Resall$Beta = log(Resall$OR)
Resall$SE = (log(Resall$OR)-log(Resall$CI95L))/1.96
Resall$Padj=p.adjust(Resall$P, method = "bonferroni")
print(Resall)
multiORplot(Resall, Pval = "P", Padj = "Padj", beta="Beta",SE = "SE",
pheno=paste(targetdiff, targetrapa))
}
}noDiff noRapa
try(getlinmodoutput_D62D224(targetdiff = "noDIFF",
targetrapa = "noRAPA",
targetline=c("D62", "D224"))) entrezgene genname D244_log2FC_2.1 D244_fdr_2.1 D244_log2FC_2.2
1 2633 GBP1 2.0297059 2.515639e-02 2.0883616
2 10763 NES 0.3475318 2.516866e-03 0.6871561
3 4811 NID1 -0.7110318 4.728859e-02 -1.2334604
4 6468 FBXW4 0.7390090 2.525435e-02 0.7973168
5 977 CD151 0.4651951 1.184530e-03 0.4485379
6 4193 MDM2 0.5162521 1.420183e-03 0.5722706
7 64093 SMOC1 0.9866785 6.718799e-19 0.6051735
8 9369 NRXN3 -2.0815225 2.496293e-04 -2.9810474
9 2628 GATM 1.0641065 8.803533e-06 0.8997106
10 56963 RGMA 0.6604419 6.084450e-04 0.4291084
11 125144 SNHG29 -0.8703269 1.045102e-07 -0.5452850
12 6426 SRSF1 -0.7835381 1.563324e-03 -0.5706565
13 284184 NDUFAF8 0.5911756 2.083037e-03 0.6861791
14 11031 RAB31 0.5840879 9.078035e-06 0.7492132
15 348 APOE 1.4498686 7.075051e-34 1.2934789
16 27113 BBC3 0.6373515 2.628145e-05 0.6024491
17 3099 HK2 -4.5134193 3.385477e-04 -2.6657547
18 2280 FKBP1A 0.6321454 2.398389e-04 0.7647272
19 3785 KCNQ2 -1.5208703 2.206510e-07 -2.3512731
20 10752 CHL1 -0.4827870 8.236944e-03 -0.5881262
21 151887 CCDC80 0.8069456 2.122272e-15 0.7595701
22 11343 MGLL 1.1144299 9.215220e-17 1.0100009
23 5099 PCDH7 -1.1263769 9.919625e-03 -1.2945977
24 5156 PDGFRA -1.7558863 1.515392e-17 -1.8639525
25 3490 IGFBP7 0.6820844 4.251649e-03 1.0385101
26 3015 H2AZ1 -0.4375541 4.262233e-02 -0.6286706
27 221749 PXDC1 0.9963329 1.487422e-03 1.0337919
28 10590 SCGN -2.3919563 1.044409e-05 -3.9601061
29 221491 SMIM29 0.9451797 9.219084e-05 1.2516034
30 1026 CDKN1A 0.3671832 2.120348e-04 0.5293086
31 3910 LAMA4 1.2757771 6.167079e-05 1.4702063
32 6196 RPS6KA2 0.4482844 1.645518e-02 0.7026317
33 1191 CLU 0.4500957 2.038659e-05 0.3534327
34 9568 GABBR2 1.0262131 4.391028e-02 1.8380988
35 4571 <NA> 3.4044633 2.362771e-119 2.8563893
36 27286 SRPX2 0.9507795 5.249112e-07 0.7486537
37 1641 DCX -0.8457536 7.306344e-03 -0.9610581
D244_fdr_2.2 D62_log2FC_2.1 D62_fdr_2.1 D62_log2FC_2.2 D62_fdr_2.2
1 2.907452e-02 4.3469675 1.478239e-04 4.0111487 9.206477e-05
2 1.662445e-11 0.7597404 9.985796e-14 0.4268953 7.990741e-07
3 3.579051e-03 -1.5148942 1.904365e-03 -1.0105930 2.296366e-02
4 1.758357e-02 1.1765193 1.364798e-03 0.7725283 3.557110e-02
5 5.268446e-03 1.2624849 1.664143e-27 0.3302487 9.754474e-03
6 9.204256e-06 0.6720729 4.176205e-05 0.3624504 1.186010e-02
7 2.779479e-05 0.7195505 1.391368e-07 0.3608222 8.928172e-04
8 1.170221e-06 -1.7172025 2.028184e-05 -1.0867269 7.910845e-04
9 3.609831e-03 0.6518976 3.394949e-02 1.0425253 7.502761e-12
10 2.922212e-02 1.6169697 1.508787e-22 0.7278730 4.458761e-07
11 5.894301e-03 -0.5333552 6.646735e-03 -0.2828075 2.579931e-02
12 1.011060e-02 -0.9535097 1.971245e-06 -0.5774088 6.881405e-04
13 2.647084e-05 0.8491433 5.692768e-12 0.3481626 7.808378e-03
14 4.453646e-09 0.3240090 2.195959e-03 0.2517174 1.169403e-02
15 3.664330e-25 2.0354059 3.708190e-20 1.2386377 5.932062e-11
16 2.306262e-04 1.5019300 2.885492e-09 1.1144906 4.741474e-06
17 4.626470e-07 -1.7911145 3.646131e-03 -1.1136392 3.778682e-03
18 9.299289e-07 0.6962166 2.508668e-05 0.3635890 4.886965e-02
19 1.761040e-10 -0.6122840 2.104394e-03 -0.6171374 3.496026e-05
20 1.121250e-02 -1.0508060 2.599907e-12 -0.4931137 1.022821e-05
21 3.271995e-14 0.7861593 3.884032e-08 0.5444540 5.158658e-06
22 1.558741e-07 0.9202739 4.176205e-05 0.6135539 8.415630e-03
23 1.285023e-03 -2.6030063 5.882622e-03 -1.5811041 1.989404e-02
24 2.069884e-30 -0.9547393 5.308860e-08 -0.4376841 5.750365e-04
25 4.750404e-07 1.4844042 6.295374e-19 1.8282513 1.823991e-53
26 8.081783e-03 -0.7067496 1.619655e-07 -0.4982645 2.218205e-05
27 1.924487e-04 0.5202967 1.478111e-02 0.4275719 3.689260e-02
28 4.182359e-23 -3.2952996 5.370178e-03 -1.0156297 2.414624e-06
29 5.325344e-06 0.9411454 1.543927e-04 0.4827715 4.565178e-02
30 4.230156e-08 1.7451782 3.284453e-77 0.8937057 6.528518e-32
31 2.159632e-06 0.4073207 4.502692e-02 0.4391435 1.254131e-02
32 3.622400e-07 0.8469458 2.974890e-07 0.4754517 4.349091e-03
33 2.737931e-03 0.6341170 6.617167e-18 0.3197075 7.692471e-10
34 4.032745e-07 1.8836534 3.992926e-19 1.7700974 1.993618e-26
35 1.746726e-69 0.9819432 6.551433e-07 0.8385856 2.779733e-06
36 6.254085e-04 6.4021162 4.092541e-03 6.4458707 7.808378e-03
37 5.303766e-03 -0.8995534 1.452689e-07 -0.7023485 1.328081e-05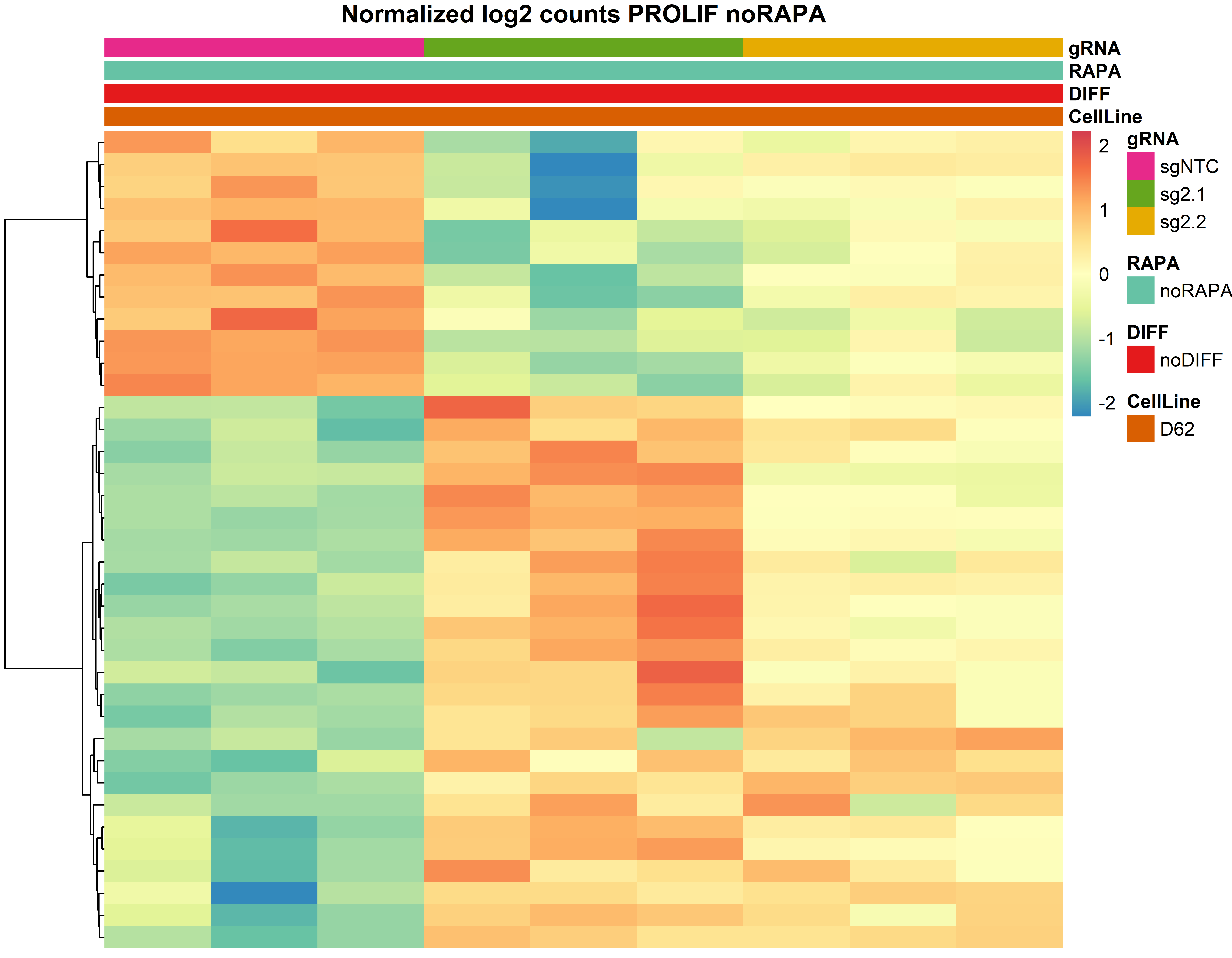
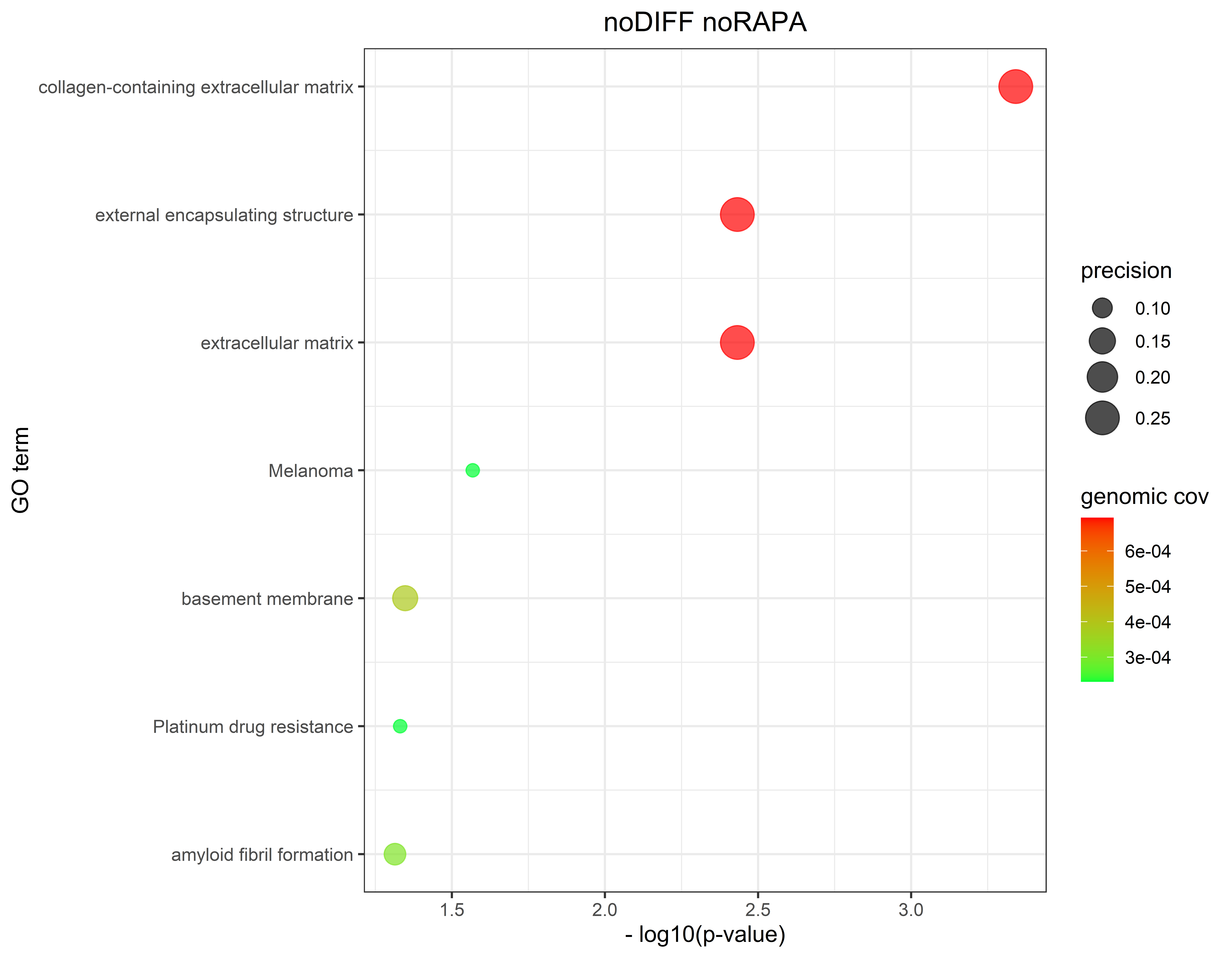
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 1.2625868 0.14679389 4.935157 6.744007e-01
02_TSC2_KO_Grabole_2016 2.7655238 1.36950627 5.738501 2.719201e-03
03_TSC2_Grabole-2016 2.7011590 1.26585169 6.258904 7.552645e-03
04_TSC2-tuber-Martin2017 3.3527641 0.08190452 20.291197 2.647592e-01
05_TSC2-SEN_SEGA-Martin2017 4.6468999 2.26248697 9.365045 1.651705e-05
06_all_EpilepsyGene 4.2567249 1.09002652 12.056600 1.906575e-02
07_high_conf_EpilepsyGene 6.2687831 0.72255483 24.878549 4.494200e-02
08_ASD_Epilepsy_EpilepsyGene 3.2351898 0.07905632 19.566881 2.728255e-01
09_Epi25_dataset 0.0000000 0.00000000 17.761425 1.000000e+00
10_SFARI_all 3.2416941 0.37557781 12.747177 1.355206e-01
11_SFARI_score4plus 4.4045954 0.50929814 17.378479 8.235315e-02
12_DeRubeis_RDNV 0.0000000 0.00000000 18.708435 1.000000e+00
13_Iossifov_RDNV 1.4746408 0.17134239 5.767836 6.486565e-01
14_FMRP_Darnell 2.6693578 0.80967916 6.930734 5.209565e-02
15_Voineagu_M12 0.0000000 0.00000000 4.094992 1.000000e+00
16_Voineagu_M16 6.1247020 1.85146682 15.966187 2.205729e-03
17_ID_genes_Neel 3.6246884 0.70867756 11.619713 5.794069e-02
18_ID_genes_pinto2014 0.0000000 0.00000000 5.838956 1.000000e+00
19_Fromer_SCZ_FunDN 0.7381642 0.01815291 4.403523 1.000000e+00
20_Cocchi_Pruning 8.7116061 0.99992954 34.823945 2.500176e-02
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.2331627 1.0979023 1.000000000
02_TSC2_KO_Grabole_2016 1.0172300 0.3585611 0.054384011
03_TSC2_Grabole-2016 0.9936810 0.3867019 0.151052893
04_TSC2-tuber-Martin2017 1.2097851 1.8938705 1.000000000
05_TSC2-SEN_SEGA-Martin2017 1.5362003 0.3672121 0.000330341
06_all_EpilepsyGene 1.4485001 0.6950500 0.381315031
07_high_conf_EpilepsyGene 1.8355822 1.1023185 0.898840045
08_ASD_Epilepsy_EpilepsyGene 1.1740876 1.8937155 1.000000000
09_Epi25_dataset -Inf NaN 1.000000000
10_SFARI_all 1.1760961 1.0996866 1.000000000
11_SFARI_score4plus 1.4826484 1.1006990 1.000000000
12_DeRubeis_RDNV -Inf NaN 1.000000000
13_Iossifov_RDNV 0.3884144 1.0982173 1.000000000
14_FMRP_Darnell 0.9818379 0.6086506 1.000000000
15_Voineagu_M12 -Inf NaN 1.000000000
16_Voineagu_M16 1.8123301 0.6103836 0.044114575
17_ID_genes_Neel 1.2877683 0.8327158 1.000000000
18_ID_genes_pinto2014 -Inf NaN 1.000000000
19_Fromer_SCZ_FunDN -0.3035889 1.8904773 1.000000000
20_Cocchi_Pruning 2.1646562 1.1044524 0.500035230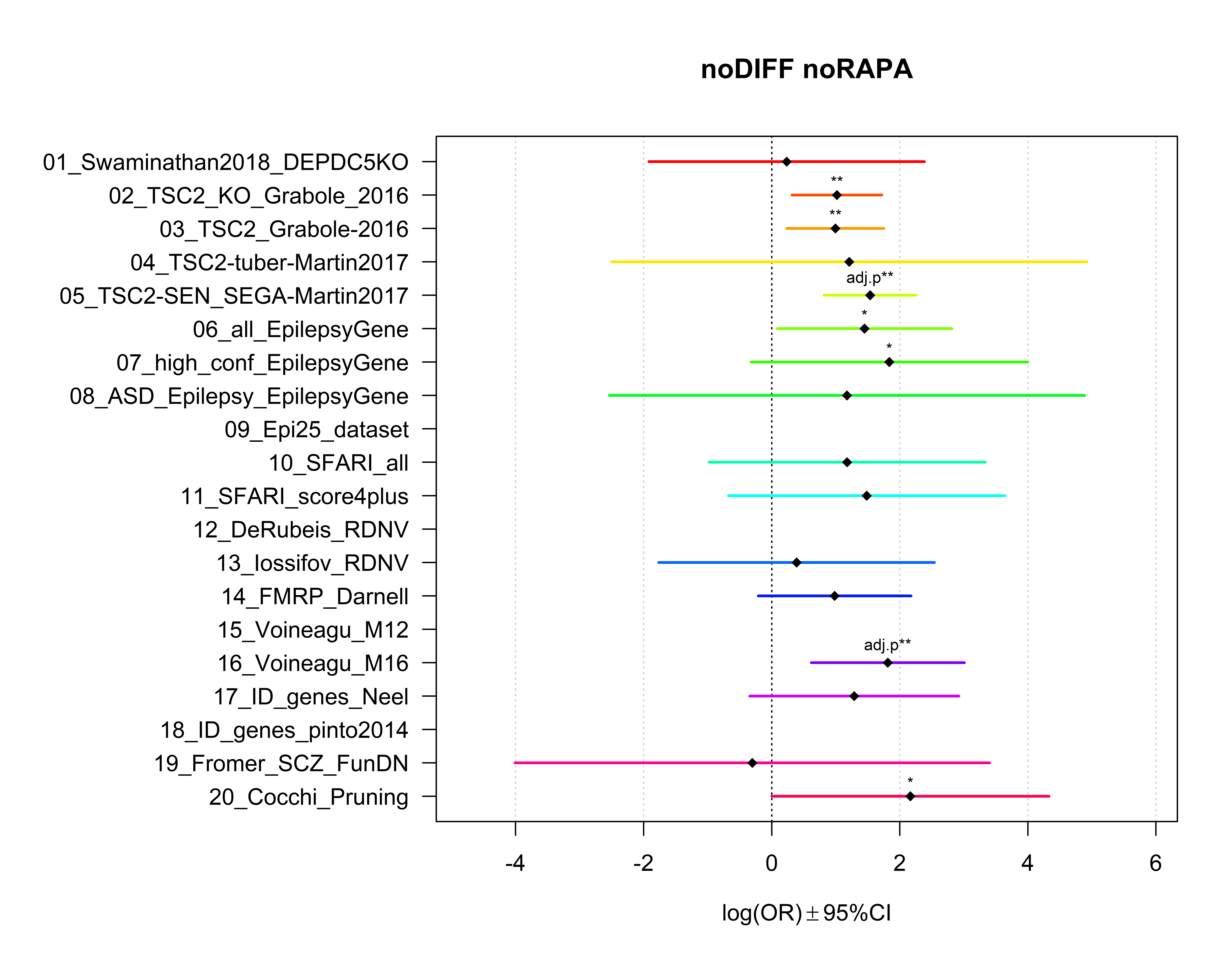
noDiff Rapa
try(getlinmodoutput_D62D224(targetdiff = "noDIFF",targetrapa = "RAPA",
targetline=c("D62", "D224"))) entrezgene genname D244_log2FC_2.1 D244_fdr_2.1 D244_log2FC_2.2 D244_fdr_2.2
1 3709 ITPR2 1.0144843 3.199205e-03 1.0716091 1.786538e-03
2 56963 RGMA 0.7056909 6.604404e-06 0.4876214 1.364992e-02
3 361 AQP4 1.5656176 1.853286e-21 1.4074809 5.825739e-17
4 348 APOE 1.1522867 2.550867e-12 1.1986122 8.038100e-11
5 4852 NPY -2.5012943 4.831459e-03 -4.1586786 6.690797e-06
6 23089 PEG10 1.2813683 2.586992e-39 0.3492587 1.840892e-02
7 1749 DLX5 -1.8207920 5.149658e-03 -9.1287072 6.304782e-10
D62_log2FC_2.1 D62_fdr_2.1 D62_log2FC_2.2 D62_fdr_2.2
1 1.4682329 7.883463e-05 1.4707635 7.064746e-04
2 1.1669346 1.509525e-08 1.0344198 3.851921e-03
3 0.7543128 2.751680e-03 0.8155313 5.746483e-03
4 1.0971097 6.637560e-05 1.0393426 2.965233e-03
5 -1.9241887 7.017330e-08 -1.5819410 2.956766e-06
6 0.7098065 2.053550e-03 0.7282390 8.417096e-03
7 -2.2913973 7.064968e-05 -1.8533389 8.417096e-03No results to show
Please make sure that the organism is correct or set significant = FALSE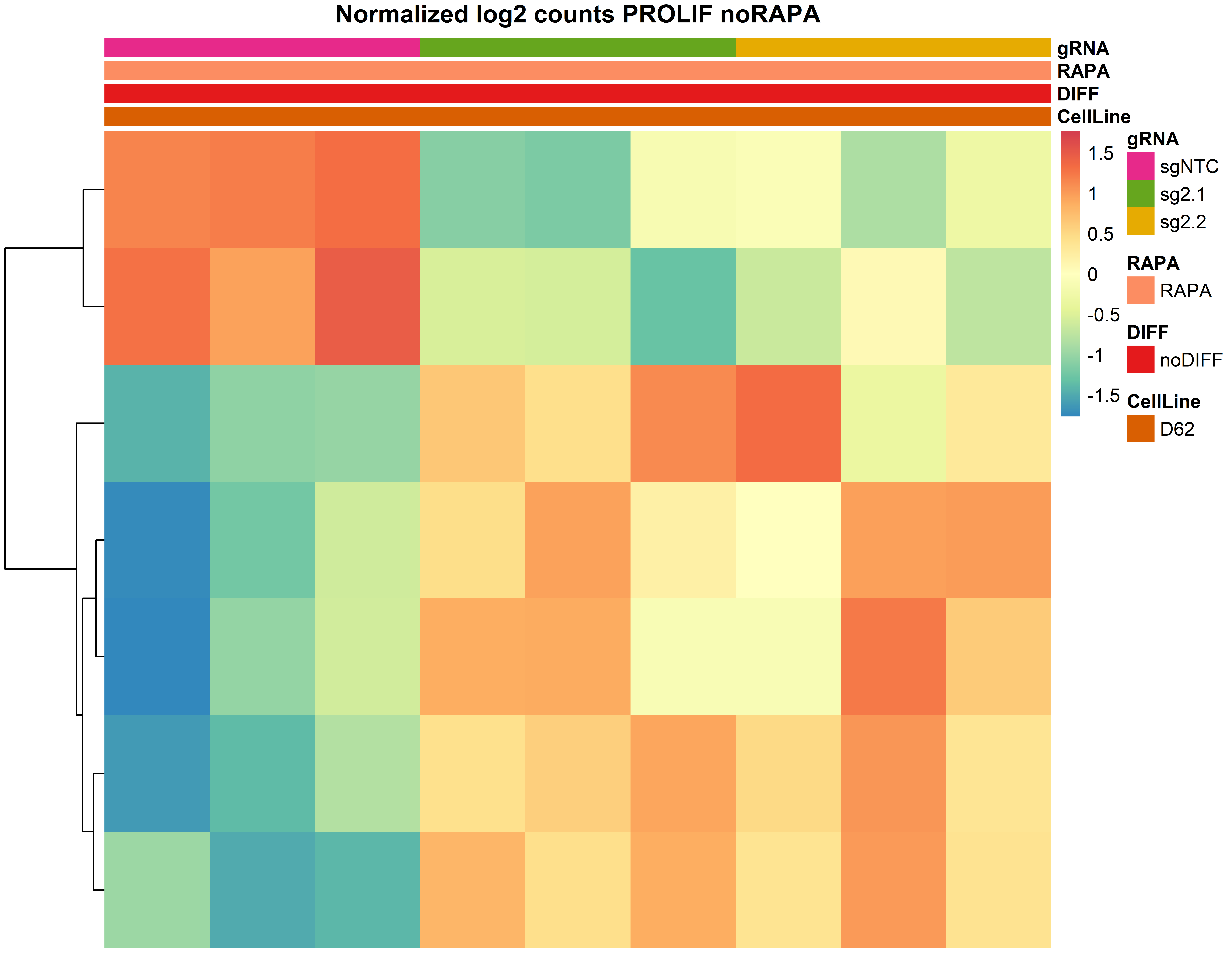
[1] "no significant results"
OR CI95L CI95U P Beta
01_Swaminathan2018_DEPDC5KO 0.000000 0.00000000 15.34525 1.00000000 -Inf
02_TSC2_KO_Grabole_2016 4.703550 0.76969415 49.40436 0.05372569 1.548318
03_TSC2_Grabole-2016 5.991268 0.72659948 275.31245 0.12490452 1.790303
04_TSC2-tuber-Martin2017 20.133355 0.43452425 168.28928 0.05646495 3.002378
05_TSC2-SEN_SEGA-Martin2017 4.549957 0.66594568 26.91631 0.06378594 1.515118
06_all_EpilepsyGene 0.000000 0.00000000 24.23466 1.00000000 -Inf
07_high_conf_EpilepsyGene 0.000000 0.00000000 75.80669 1.00000000 -Inf
08_ASD_Epilepsy_EpilepsyGene 0.000000 0.00000000 81.09474 1.00000000 -Inf
09_Epi25_dataset 0.000000 0.00000000 116.75990 1.00000000 -Inf
10_SFARI_all 0.000000 0.00000000 39.35942 1.00000000 -Inf
11_SFARI_score4plus 0.000000 0.00000000 53.37643 1.00000000 -Inf
12_DeRubeis_RDNV 0.000000 0.00000000 123.67156 1.00000000 -Inf
13_Iossifov_RDNV 4.301383 0.09335061 35.54219 0.23395033 1.458937
14_FMRP_Darnell 2.838083 0.06162727 23.43810 0.32956955 1.043129
15_Voineagu_M12 0.000000 0.00000000 27.12534 1.00000000 -Inf
16_Voineagu_M16 15.560133 1.47728515 95.50797 0.01219980 2.744712
17_ID_genes_Neel 0.000000 0.00000000 28.41846 1.00000000 -Inf
18_ID_genes_pinto2014 0.000000 0.00000000 38.67674 1.00000000 -Inf
19_Fromer_SCZ_FunDN 0.000000 0.00000000 18.49146 1.00000000 -Inf
20_Cocchi_Pruning 25.135627 0.54163841 210.03663 0.04559469 3.224286
SE Padj
01_Swaminathan2018_DEPDC5KO NaN 1.0000000
02_TSC2_KO_Grabole_2016 0.9235100 1.0000000
03_TSC2_Grabole-2016 1.0763689 1.0000000
04_TSC2-tuber-Martin2017 1.9570823 1.0000000
05_TSC2-SEN_SEGA-Martin2017 0.9804413 1.0000000
06_all_EpilepsyGene NaN 1.0000000
07_high_conf_EpilepsyGene NaN 1.0000000
08_ASD_Epilepsy_EpilepsyGene NaN 1.0000000
09_Epi25_dataset NaN 1.0000000
10_SFARI_all NaN 1.0000000
11_SFARI_score4plus NaN 1.0000000
12_DeRubeis_RDNV NaN 1.0000000
13_Iossifov_RDNV 1.9542498 1.0000000
14_FMRP_Darnell 1.9539692 1.0000000
15_Voineagu_M12 NaN 1.0000000
16_Voineagu_M16 1.2012786 0.2439961
17_ID_genes_Neel NaN 1.0000000
18_ID_genes_pinto2014 NaN 1.0000000
19_Fromer_SCZ_FunDN NaN 1.0000000
20_Cocchi_Pruning 1.9578790 0.9118937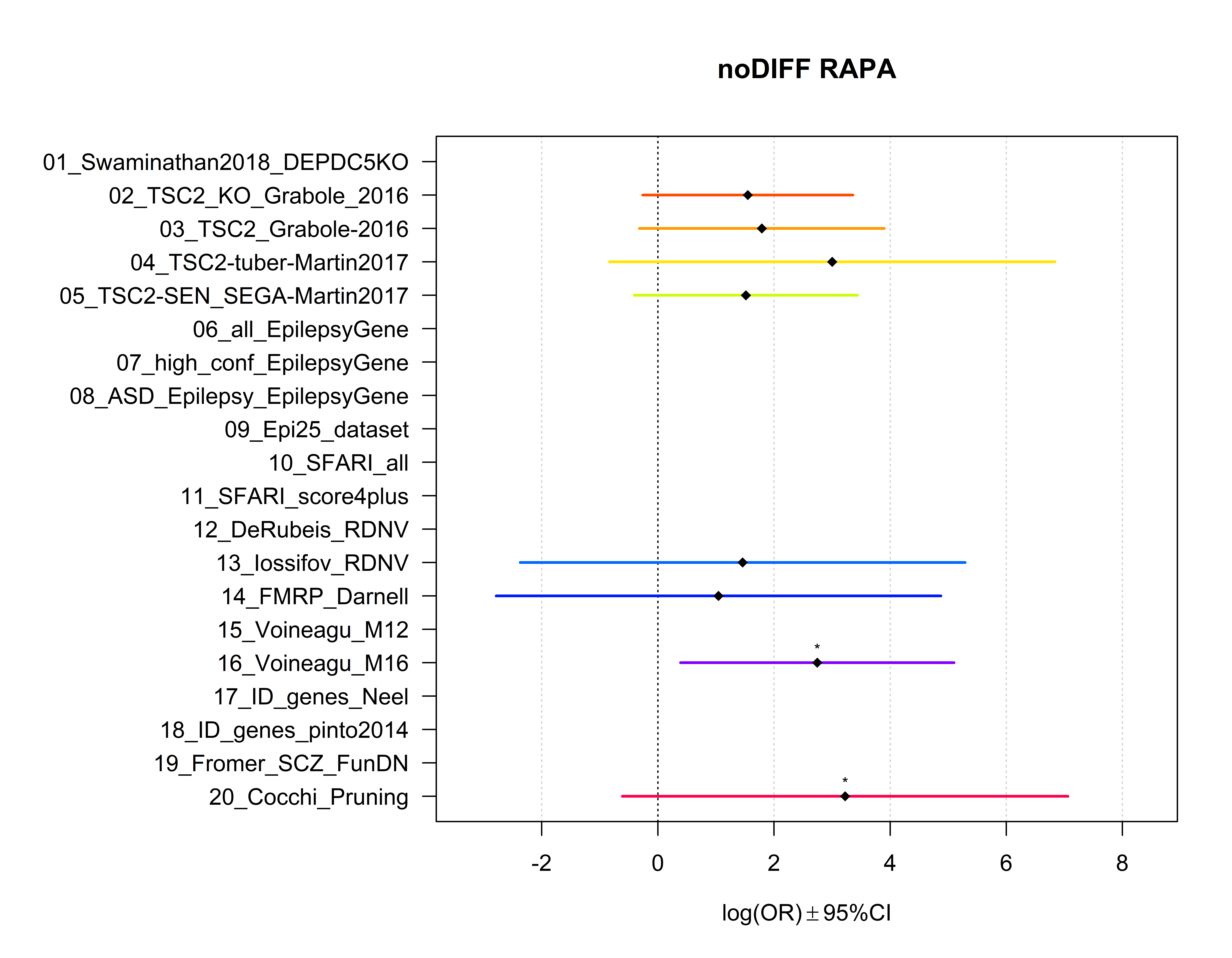
Diff noRapa
try(getlinmodoutput_D62D224(targetdiff = "DIFF",targetrapa = "noRAPA",
targetline=c("D62", "D224"))) entrezgene genname D244_log2FC_2.1 D244_fdr_2.1 D244_log2FC_2.2
1 50486 G0S2 -1.4492217 3.370529e-32 -1.7927431
2 7044 LEFTY2 -1.6863840 1.444934e-19 -2.6934841
3 652 BMP4 1.4247389 3.501929e-09 1.1752283
4 4804 NGFR -2.1767618 3.267739e-23 -2.3203327
5 56937 PMEPA1 -1.1739691 1.378185e-19 -1.1140883
6 80781 COL18A1 -1.0190535 1.418955e-07 -1.0103524
7 7869 SEMA3B -1.3403981 3.060962e-05 -1.2301226
8 7018 TF -1.7913566 9.804860e-08 -2.1881714
9 57537 SORCS2 -1.1432855 4.392572e-07 -1.3209447
10 100505738 MIR4458HG -0.9544000 2.870460e-05 -0.9170490
11 11346 SYNPO -1.0320585 3.175806e-06 -0.8511356
12 1843 DUSP1 1.8428072 9.089214e-06 1.7754384
13 2697 GJA1 0.9786886 1.006861e-11 0.7103970
14 5244 ABCB4 -1.3752686 2.416092e-05 -2.2432327
15 3371 TNC 1.4093005 5.957962e-03 1.4983126
D244_fdr_2.2 D62_log2FC_2.1 D62_fdr_2.1 D62_log2FC_2.2 D62_fdr_2.2
1 4.532054e-17 -1.2769869 3.776809e-02 -3.7478581 2.744552e-36
2 1.215175e-28 -1.8967514 2.076460e-02 -5.4919034 1.299317e-06
3 3.095265e-03 3.4322241 2.358141e-02 3.9413514 1.737453e-02
4 2.270876e-09 -1.8961919 1.638191e-04 -2.0647461 7.699124e-05
5 6.627336e-09 -0.5230080 3.898929e-03 -0.9284119 2.782505e-07
6 6.341399e-05 -0.6429705 4.185984e-03 -1.4012123 9.118276e-11
7 4.896891e-03 -1.6475938 4.752961e-08 -1.4351780 2.091144e-04
8 5.243374e-07 -3.3138202 4.725046e-35 -2.6201244 4.161319e-27
9 1.875633e-06 -0.7871403 4.529303e-02 -1.6736831 1.240910e-04
10 1.585138e-02 -0.6736466 4.960015e-04 -0.9792733 3.598272e-07
11 7.015280e-03 -1.1973194 6.008782e-09 -1.1898142 1.055427e-05
12 2.802331e-03 2.3389517 1.273281e-03 2.5638656 2.811444e-03
13 2.802331e-03 0.7860628 1.505593e-09 0.6923602 6.978354e-08
14 1.296467e-05 -0.7938959 1.121641e-02 -1.7461164 2.042689e-08
15 4.016110e-02 1.4982035 2.413395e-02 2.0004229 2.211218e-03No results to show
Please make sure that the organism is correct or set significant = FALSE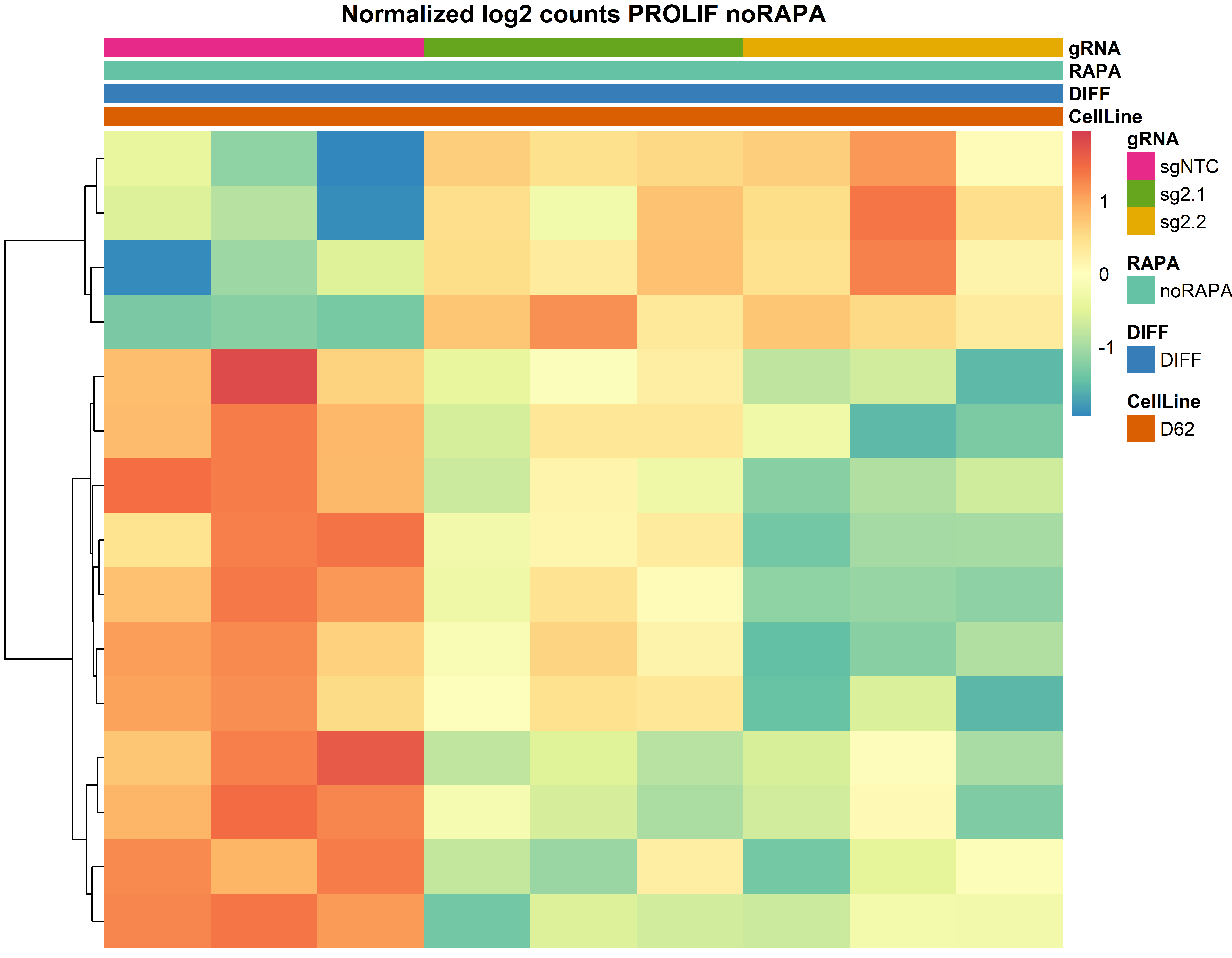
[1] "no significant results"
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 0.0000000 0.00000000 6.167004 1.000000000
02_TSC2_KO_Grabole_2016 1.6460497 0.50777882 5.197577 0.416071164
03_TSC2_Grabole-2016 1.4975117 0.47572921 5.115758 0.607038627
04_TSC2-tuber-Martin2017 18.7449166 2.03117226 84.402907 0.006629492
05_TSC2-SEN_SEGA-Martin2017 0.9322187 0.10205649 4.122830 1.000000000
06_all_EpilepsyGene 0.0000000 0.00000000 9.741362 1.000000000
07_high_conf_EpilepsyGene 0.0000000 0.00000000 30.536195 1.000000000
08_ASD_Epilepsy_EpilepsyGene 0.0000000 0.00000000 32.678358 1.000000000
09_Epi25_dataset 0.0000000 0.00000000 47.215752 1.000000000
10_SFARI_all 0.0000000 0.00000000 15.812671 1.000000000
11_SFARI_score4plus 0.0000000 0.00000000 21.483638 1.000000000
12_DeRubeis_RDNV 0.0000000 0.00000000 49.729830 1.000000000
13_Iossifov_RDNV 0.0000000 0.00000000 7.199790 1.000000000
14_FMRP_Darnell 1.2156917 0.02871371 8.010992 0.575587957
15_Voineagu_M12 0.0000000 0.00000000 10.899210 1.000000000
16_Voineagu_M16 2.7711452 0.06535908 18.311369 0.317900267
17_ID_genes_Neel 0.0000000 0.00000000 11.419272 1.000000000
18_ID_genes_pinto2014 0.0000000 0.00000000 15.542615 1.000000000
19_Fromer_SCZ_FunDN 0.0000000 0.00000000 7.429053 1.000000000
20_Cocchi_Pruning 0.0000000 0.00000000 42.367010 1.000000000
Beta SE Padj
01_Swaminathan2018_DEPDC5KO -Inf NaN 1.0000000
02_TSC2_KO_Grabole_2016 0.49837832 0.6000447 1.0000000
03_TSC2_Grabole-2016 0.40380484 0.5850568 1.0000000
04_TSC2-tuber-Martin2017 2.93092260 1.1338314 0.1325898
05_TSC2-SEN_SEGA-Martin2017 -0.07018781 1.1285923 1.0000000
06_all_EpilepsyGene -Inf NaN 1.0000000
07_high_conf_EpilepsyGene -Inf NaN 1.0000000
08_ASD_Epilepsy_EpilepsyGene -Inf NaN 1.0000000
09_Epi25_dataset -Inf NaN 1.0000000
10_SFARI_all -Inf NaN 1.0000000
11_SFARI_score4plus -Inf NaN 1.0000000
12_DeRubeis_RDNV -Inf NaN 1.0000000
13_Iossifov_RDNV -Inf NaN 1.0000000
14_FMRP_Darnell 0.19531323 1.9110684 1.0000000
15_Voineagu_M12 -Inf NaN 1.0000000
16_Voineagu_M16 1.01926068 1.9117957 1.0000000
17_ID_genes_Neel -Inf NaN 1.0000000
18_ID_genes_pinto2014 -Inf NaN 1.0000000
19_Fromer_SCZ_FunDN -Inf NaN 1.0000000
20_Cocchi_Pruning -Inf NaN 1.0000000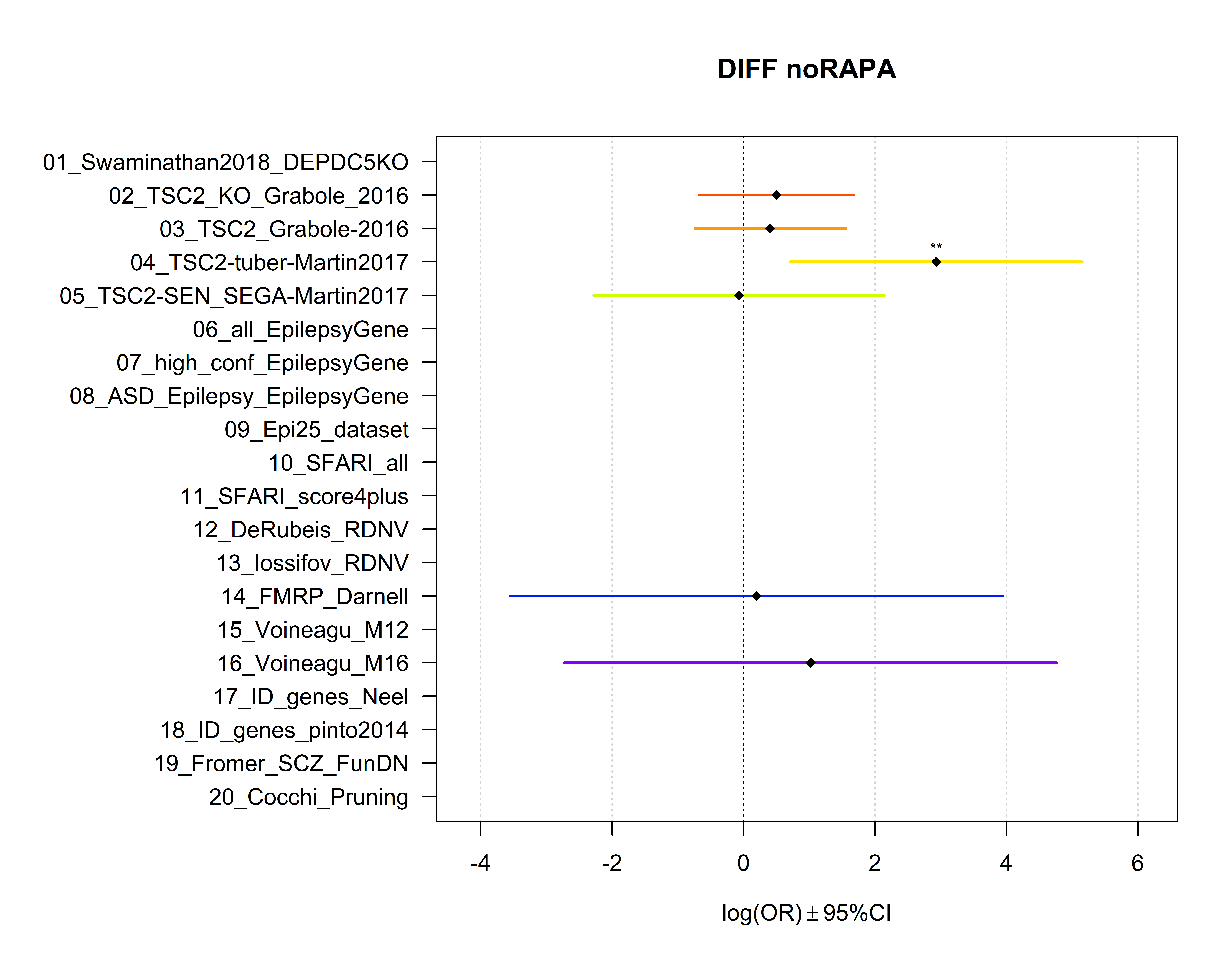
Diff Rapa
try(getlinmodoutput_D62D224(targetdiff = "DIFF",targetrapa = "RAPA",
targetline=c("D62", "D224"))) entrezgene genname D244_log2FC_2.1 D244_fdr_2.1 D244_log2FC_2.2
1 3491 CCN1 0.4678656 2.963636e-06 0.4105692
2 8490 RGS5 0.7061001 9.243401e-03 0.5901435
3 50486 G0S2 -2.0420221 9.635264e-145 -2.5996040
4 7044 LEFTY2 -1.3192934 3.280549e-18 -2.3110081
5 2674 GFRA1 -0.4799768 1.237850e-03 -0.6378438
6 114795 TMEM132B -3.1403066 4.570484e-06 -3.0132607
7 23002 DAAM1 0.6938355 2.863809e-03 0.5782007
8 10882 C1QL1 -1.1800573 4.585792e-02 -1.3575829
9 4804 NGFR -1.8994842 1.471545e-13 -2.1854385
10 2995 GYPC -2.4190113 1.415998e-15 -2.5897350
11 3488 IGFBP5 -1.4790171 1.730247e-60 -1.3238131
12 1295 COL8A1 -2.5667320 1.117183e-51 -2.8067329
13 151887 CCDC80 0.8400733 1.550300e-13 1.0909859
14 7018 TF -2.2191138 7.809017e-09 -2.7186226
15 5947 RBP1 -1.0346072 3.851251e-48 -1.1250151
16 10417 SPON2 -1.6792183 1.085527e-05 -1.6373186
17 83888 FGFBP2 -2.3020082 9.156676e-12 -4.3108924
18 9547 CXCL14 -0.5235165 1.148369e-10 -0.4632759
19 2878 GPX3 -1.0164418 8.801801e-26 -0.7308690
20 23286 WWC1 1.5686248 3.671663e-06 1.4815662
21 6424 SFRP4 -1.5686024 2.187938e-85 -1.7488886
22 83543 AIF1L -1.2388526 3.130862e-20 -0.5682422
D244_fdr_2.2 D62_log2FC_2.1 D62_fdr_2.1 D62_log2FC_2.2 D62_fdr_2.2
1 9.041386e-05 0.9195214 7.581913e-06 0.7819251 2.641835e-02
2 6.181386e-03 0.9423725 2.179796e-03 1.3297066 5.430918e-06
3 2.301167e-217 -2.1437954 2.962521e-18 -2.7798517 3.914474e-23
4 5.751049e-64 -1.1461461 1.517838e-02 -2.5101270 2.217413e-06
5 1.191763e-05 -1.7237479 8.130698e-05 -1.9850405 7.413693e-05
6 5.957135e-10 -2.0420745 9.180844e-15 -1.4932069 6.161726e-06
7 4.677801e-02 0.7554336 2.883318e-02 0.8353463 2.729476e-02
8 3.915880e-02 -1.2462122 5.655205e-06 -1.2834520 1.792262e-03
9 4.715661e-26 -2.2008093 1.158457e-10 -2.1406331 1.813272e-06
10 5.915615e-21 -2.3370979 5.640820e-06 -2.1226178 2.620590e-06
11 6.039310e-53 -1.2042255 1.457509e-03 -1.2840334 3.354261e-03
12 2.533751e-78 -4.5471851 7.161381e-03 -3.7379338 2.815452e-02
13 8.688693e-17 0.7317143 1.740585e-02 0.8723305 1.319498e-03
14 1.812506e-08 -3.7162415 1.267988e-67 -2.8326531 2.204247e-30
15 1.013389e-56 -1.0910735 4.502025e-07 -0.7994095 4.109501e-02
16 1.499922e-03 -2.4640319 1.386561e-02 -2.8331591 2.289093e-03
17 4.089891e-24 -1.9562182 9.567416e-03 -2.3549128 2.393688e-04
18 1.335559e-07 -0.8064340 7.756694e-04 -1.5018443 1.611197e-09
19 1.484571e-13 -1.3013559 3.780855e-08 -2.0926990 6.557503e-11
20 3.857516e-05 1.0141019 4.480737e-03 1.2974451 1.326571e-04
21 8.250920e-100 -0.7815462 4.441654e-04 -1.3426635 1.190509e-07
22 9.922051e-04 -0.9344874 1.208559e-03 -0.9029015 2.547855e-02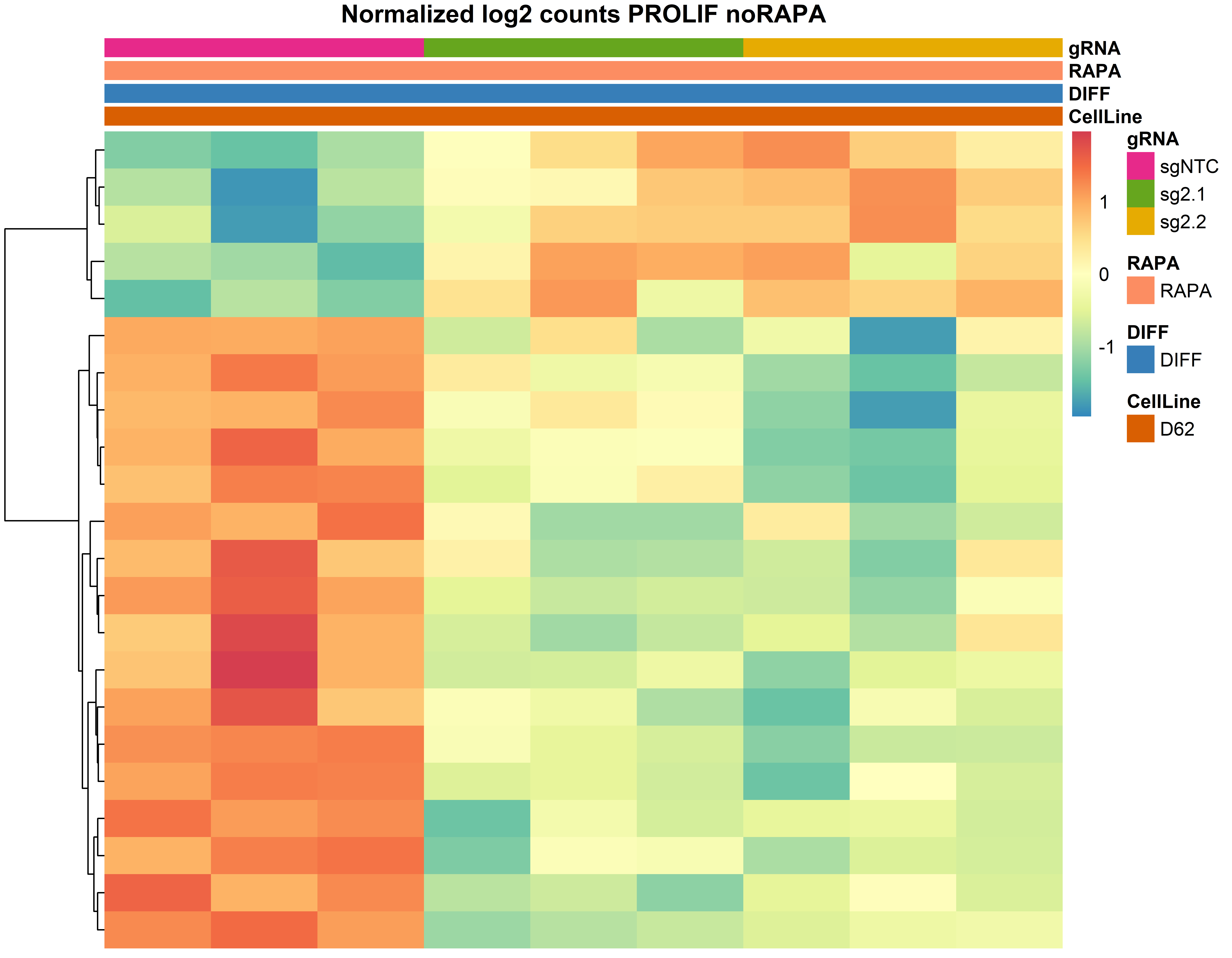
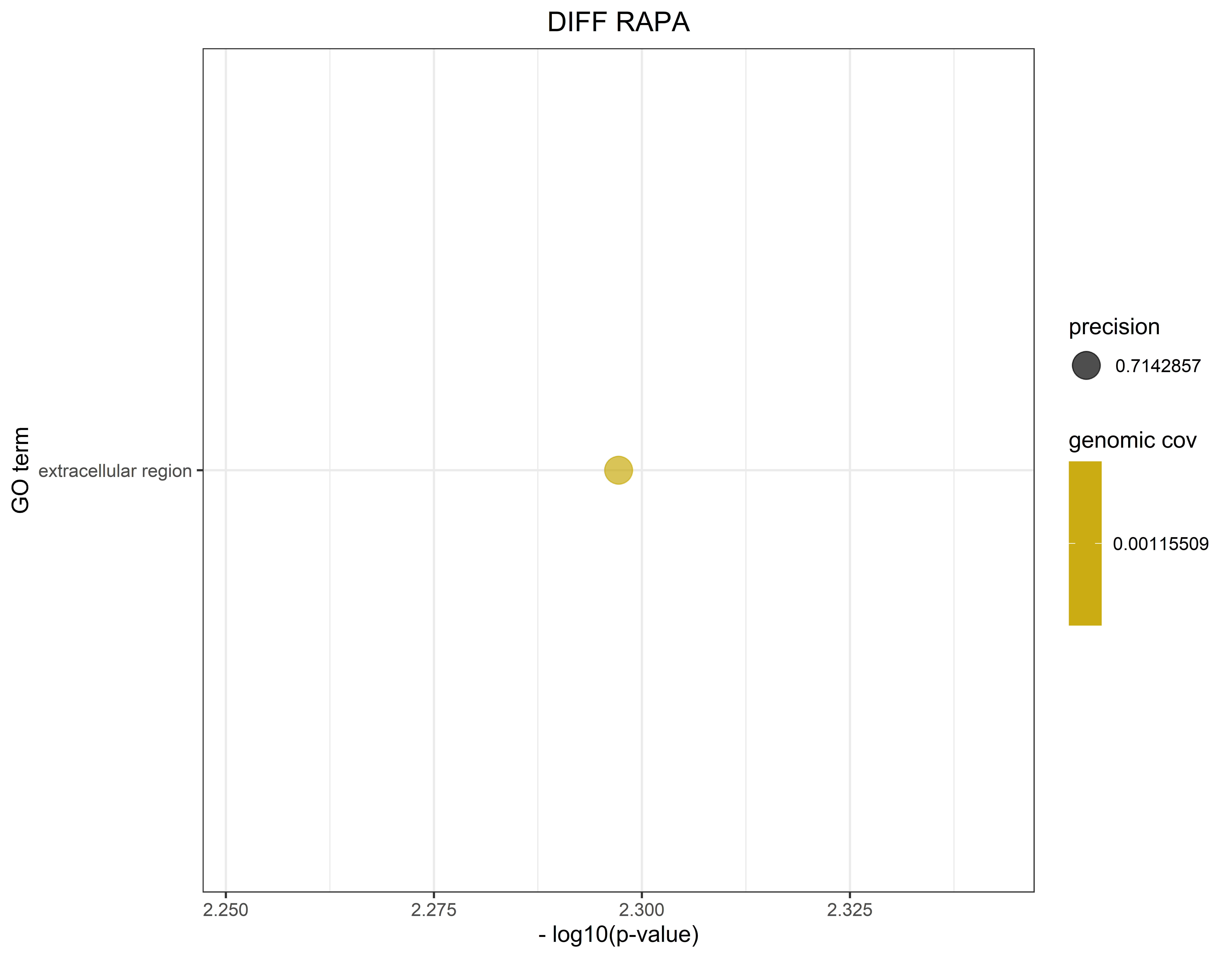
OR CI95L CI95U P
01_Swaminathan2018_DEPDC5KO 2.2120492 0.25009058 9.138635 0.24655625
02_TSC2_KO_Grabole_2016 2.2592188 0.89381168 5.842774 0.07047344
03_TSC2_Grabole-2016 1.4422649 0.56990956 3.825944 0.52313027
04_TSC2-tuber-Martin2017 0.0000000 0.00000000 22.192951 1.00000000
05_TSC2-SEN_SEGA-Martin2017 2.8343541 0.97630724 7.397057 0.02770732
06_all_EpilepsyGene 0.0000000 0.00000000 6.377532 1.00000000
07_high_conf_EpilepsyGene 0.0000000 0.00000000 19.996585 1.00000000
08_ASD_Epilepsy_EpilepsyGene 0.0000000 0.00000000 21.419829 1.00000000
09_Epi25_dataset 0.0000000 0.00000000 30.931119 1.00000000
10_SFARI_all 0.0000000 0.00000000 10.356084 1.00000000
11_SFARI_score4plus 0.0000000 0.00000000 14.057433 1.00000000
12_DeRubeis_RDNV 0.0000000 0.00000000 32.566918 1.00000000
13_Iossifov_RDNV 0.0000000 0.00000000 4.713469 1.00000000
14_FMRP_Darnell 0.8100381 0.01956154 5.050315 1.00000000
15_Voineagu_M12 0.0000000 0.00000000 7.136577 1.00000000
16_Voineagu_M16 0.0000000 0.00000000 7.093410 1.00000000
17_ID_genes_Neel 0.0000000 0.00000000 7.477368 1.00000000
18_ID_genes_pinto2014 0.0000000 0.00000000 10.179156 1.00000000
19_Fromer_SCZ_FunDN 0.0000000 0.00000000 4.863581 1.00000000
20_Cocchi_Pruning 0.0000000 0.00000000 27.766996 1.00000000
Beta SE Padj
01_Swaminathan2018_DEPDC5KO 0.7939193 1.1121691 1.0000000
02_TSC2_KO_Grabole_2016 0.8150191 0.4731017 1.0000000
03_TSC2_Grabole-2016 0.3662147 0.4737206 1.0000000
04_TSC2-tuber-Martin2017 -Inf NaN 1.0000000
05_TSC2-SEN_SEGA-Martin2017 1.0418141 0.5437714 0.5541465
06_all_EpilepsyGene -Inf NaN 1.0000000
07_high_conf_EpilepsyGene -Inf NaN 1.0000000
08_ASD_Epilepsy_EpilepsyGene -Inf NaN 1.0000000
09_Epi25_dataset -Inf NaN 1.0000000
10_SFARI_all -Inf NaN 1.0000000
11_SFARI_score4plus -Inf NaN 1.0000000
12_DeRubeis_RDNV -Inf NaN 1.0000000
13_Iossifov_RDNV -Inf NaN 1.0000000
14_FMRP_Darnell -0.2106739 1.8997531 1.0000000
15_Voineagu_M12 -Inf NaN 1.0000000
16_Voineagu_M16 -Inf NaN 1.0000000
17_ID_genes_Neel -Inf NaN 1.0000000
18_ID_genes_pinto2014 -Inf NaN 1.0000000
19_Fromer_SCZ_FunDN -Inf NaN 1.0000000
20_Cocchi_Pruning -Inf NaN 1.0000000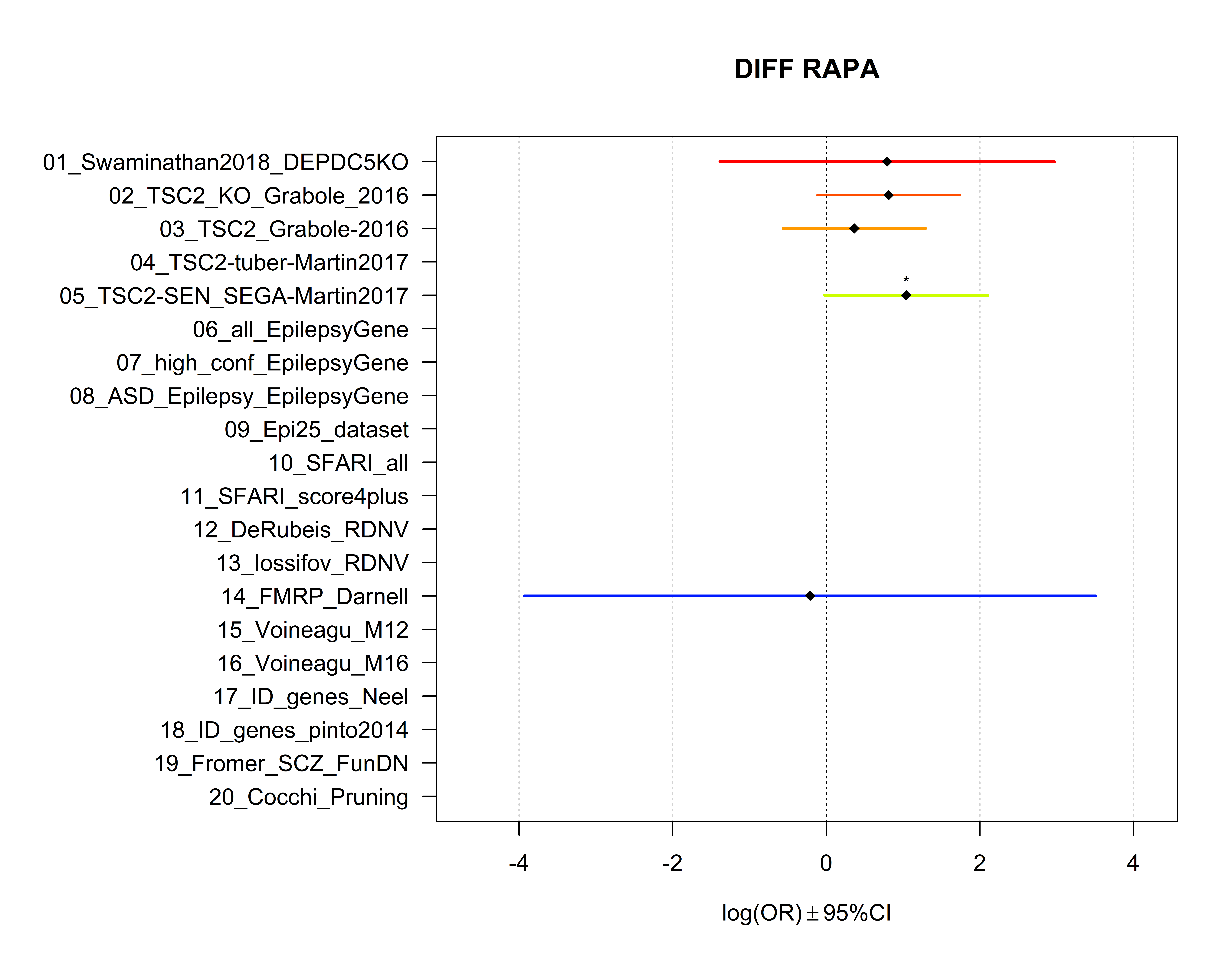
sessionInfo()R version 4.1.2 (2021-11-01)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 18363)
Matrix products: default
locale:
[1] LC_COLLATE=German_Germany.1252 LC_CTYPE=German_Germany.1252
[3] LC_MONETARY=German_Germany.1252 LC_NUMERIC=C
[5] LC_TIME=German_Germany.1252
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] gprofiler2_0.2.1 xlsx_0.6.5
[3] limma_3.50.0 flashClust_1.01-2
[5] WGCNA_1.70-3 fastcluster_1.2.3
[7] dynamicTreeCut_1.63-1 knitr_1.37
[9] DESeq2_1.34.0 SummarizedExperiment_1.24.0
[11] Biobase_2.54.0 MatrixGenerics_1.6.0
[13] matrixStats_0.61.0 GenomicRanges_1.46.1
[15] GenomeInfoDb_1.30.1 IRanges_2.28.0
[17] S4Vectors_0.32.3 BiocGenerics_0.40.0
[19] pheatmap_1.0.12 RColorBrewer_1.1-2
[21] compareGroups_4.5.1 forcats_0.5.1
[23] stringr_1.4.0 dplyr_1.0.7
[25] purrr_0.3.4 readr_2.1.2
[27] tidyr_1.1.4 tibble_3.1.6
[29] ggplot2_3.3.5 tidyverse_1.3.1
[31] kableExtra_1.3.4 workflowr_1.7.0
loaded via a namespace (and not attached):
[1] readxl_1.3.1 uuid_1.0-3 backports_1.4.1
[4] Hmisc_4.6-0 systemfonts_1.0.3 lazyeval_0.2.2
[7] splines_4.1.2 BiocParallel_1.28.3 digest_0.6.29
[10] foreach_1.5.2 htmltools_0.5.2 GO.db_3.14.0
[13] fansi_1.0.2 checkmate_2.0.0 magrittr_2.0.2
[16] Rsolnp_1.16 memoise_2.0.1 cluster_2.1.2
[19] doParallel_1.0.16 tzdb_0.2.0 Biostrings_2.62.0
[22] annotate_1.72.0 modelr_0.1.8 officer_0.4.1
[25] svglite_2.0.0 jpeg_0.1-9 colorspace_2.0-2
[28] blob_1.2.2 rvest_1.0.2 haven_2.4.3
[31] xfun_0.29 callr_3.7.0 crayon_1.4.2
[34] RCurl_1.98-1.5 jsonlite_1.7.3 genefilter_1.76.0
[37] impute_1.68.0 iterators_1.0.13 survival_3.2-13
[40] glue_1.6.1 gtable_0.3.0 zlibbioc_1.40.0
[43] XVector_0.34.0 webshot_0.5.2 DelayedArray_0.20.0
[46] scales_1.1.1 DBI_1.1.2 Rcpp_1.0.8
[49] htmlTable_2.4.0 viridisLite_0.4.0 xtable_1.8-4
[52] foreign_0.8-82 bit_4.0.4 preprocessCore_1.56.0
[55] Formula_1.2-4 truncnorm_1.0-8 htmlwidgets_1.5.4
[58] httr_1.4.2 ellipsis_0.3.2 mice_3.14.0
[61] farver_2.1.0 rJava_1.0-6 pkgconfig_2.0.3
[64] XML_3.99-0.8 nnet_7.3-17 sass_0.4.0
[67] dbplyr_2.1.1 locfit_1.5-9.4 utf8_1.2.2
[70] labeling_0.4.2 tidyselect_1.1.1 rlang_1.0.0
[73] later_1.3.0 AnnotationDbi_1.56.2 munsell_0.5.0
[76] cellranger_1.1.0 tools_4.1.2 cachem_1.0.6
[79] cli_3.1.1 generics_0.1.2 RSQLite_2.2.9
[82] broom_0.7.12 evaluate_0.14 fastmap_1.1.0
[85] yaml_2.2.2 processx_3.5.2 bit64_4.0.5
[88] fs_1.5.2 zip_2.2.0 KEGGREST_1.34.0
[91] whisker_0.4 xml2_1.3.3 compiler_4.1.2
[94] rstudioapi_0.13 plotly_4.10.0 png_0.1-7
[97] reprex_2.0.1 geneplotter_1.72.0 bslib_0.3.1
[100] stringi_1.7.6 HardyWeinberg_1.7.4 highr_0.9
[103] ps_1.6.0 gdtools_0.2.3 lattice_0.20-45
[106] Matrix_1.4-0 vctrs_0.3.8 pillar_1.7.0
[109] lifecycle_1.0.1 jquerylib_0.1.4 data.table_1.14.2
[112] bitops_1.0-7 flextable_0.6.10 httpuv_1.6.5
[115] latticeExtra_0.6-29 R6_2.5.1 promises_1.2.0.1
[118] gridExtra_2.3 writexl_1.4.0 codetools_0.2-18
[121] assertthat_0.2.1 xlsxjars_0.6.1 chron_2.3-56
[124] rprojroot_2.0.2 withr_2.4.3 GenomeInfoDbData_1.2.7
[127] parallel_4.1.2 hms_1.1.1 rpart_4.1.16
[130] grid_4.1.2 rmarkdown_2.11 git2r_0.29.0
[133] getPass_0.2-2 lubridate_1.8.0 base64enc_0.1-3